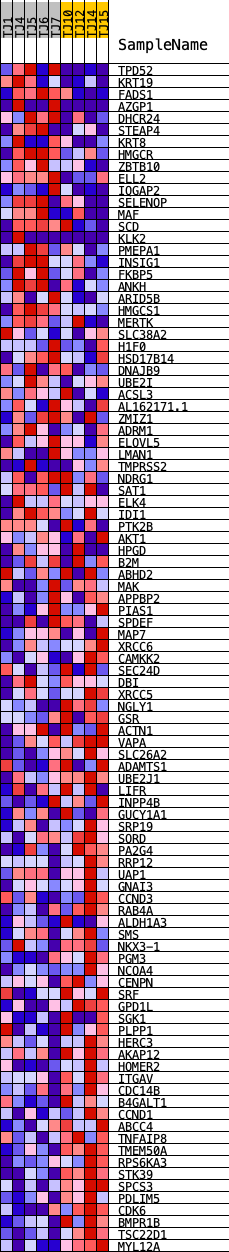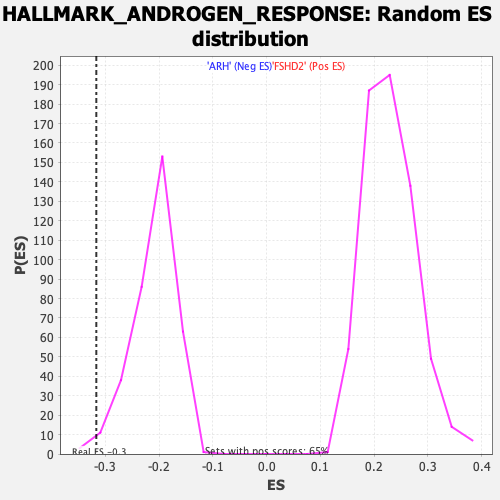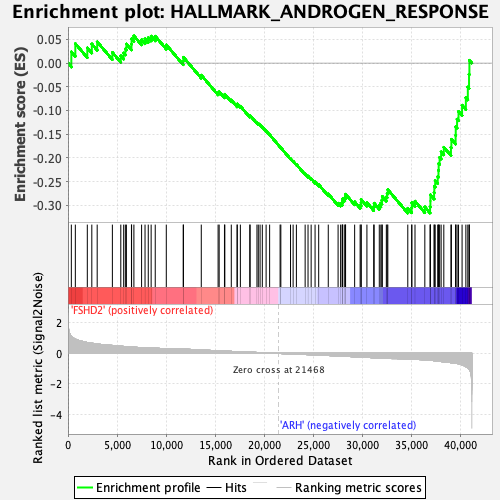
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_ANDROGEN_RESPONSE |
| Enrichment Score (ES) | -0.3166921 |
| Normalized Enrichment Score (NES) | -1.5185899 |
| Nominal p-value | 0.011267605 |
| FDR q-value | 0.021686843 |
| FWER p-Value | 0.128 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TPD52 | ENSG00000076554.15 | 340 | 1.096 | 0.0238 | No | ||
| 2 | KRT19 | ENSG00000171345.13 | 747 | 0.927 | 0.0411 | No | ||
| 3 | FADS1 | ENSG00000149485.19 | 1968 | 0.709 | 0.0322 | No | ||
| 4 | AZGP1 | ENSG00000160862.13 | 2427 | 0.660 | 0.0404 | No | ||
| 5 | DHCR24 | ENSG00000116133.13 | 2975 | 0.608 | 0.0449 | No | ||
| 6 | STEAP4 | ENSG00000127954.12 | 4523 | 0.518 | 0.0224 | No | ||
| 7 | KRT8 | ENSG00000170421.12 | 5381 | 0.466 | 0.0152 | No | ||
| 8 | HMGCR | ENSG00000113161.16 | 5676 | 0.449 | 0.0212 | No | ||
| 9 | ZBTB10 | ENSG00000205189.12 | 5845 | 0.441 | 0.0301 | No | ||
| 10 | ELL2 | ENSG00000118985.16 | 5952 | 0.436 | 0.0403 | No | ||
| 11 | IQGAP2 | ENSG00000145703.16 | 6469 | 0.409 | 0.0397 | No | ||
| 12 | SELENOP | ENSG00000250722.6 | 6481 | 0.409 | 0.0514 | No | ||
| 13 | MAF | ENSG00000178573.7 | 6717 | 0.400 | 0.0574 | No | ||
| 14 | SCD | ENSG00000099194.6 | 7501 | 0.371 | 0.0492 | No | ||
| 15 | KLK2 | ENSG00000167751.13 | 7852 | 0.362 | 0.0513 | No | ||
| 16 | PMEPA1 | ENSG00000124225.16 | 8181 | 0.354 | 0.0537 | No | ||
| 17 | INSIG1 | ENSG00000186480.13 | 8475 | 0.352 | 0.0569 | No | ||
| 18 | FKBP5 | ENSG00000096060.14 | 8892 | 0.340 | 0.0567 | No | ||
| 19 | ANKH | ENSG00000154122.14 | 10019 | 0.311 | 0.0384 | No | ||
| 20 | ARID5B | ENSG00000150347.16 | 11745 | 0.281 | 0.0047 | No | ||
| 21 | HMGCS1 | ENSG00000112972.15 | 11768 | 0.279 | 0.0123 | No | ||
| 22 | MERTK | ENSG00000153208.17 | 13585 | 0.219 | -0.0255 | No | ||
| 23 | SLC38A2 | ENSG00000134294.14 | 15300 | 0.165 | -0.0624 | No | ||
| 24 | H1F0 | ENSG00000189060.5 | 15408 | 0.161 | -0.0603 | No | ||
| 25 | HSD17B14 | ENSG00000087076.9 | 15949 | 0.146 | -0.0691 | No | ||
| 26 | DNAJB9 | ENSG00000128590.5 | 15986 | 0.146 | -0.0657 | No | ||
| 27 | UBE2I | ENSG00000103275.20 | 16644 | 0.127 | -0.0780 | No | ||
| 28 | ACSL3 | ENSG00000123983.14 | 17241 | 0.111 | -0.0893 | No | ||
| 29 | AL162171.1 | ENSG00000258789.1 | 17248 | 0.111 | -0.0862 | No | ||
| 30 | ZMIZ1 | ENSG00000108175.17 | 17570 | 0.102 | -0.0910 | No | ||
| 31 | ADRM1 | ENSG00000130706.13 | 18532 | 0.078 | -0.1121 | No | ||
| 32 | ELOVL5 | ENSG00000012660.14 | 18561 | 0.077 | -0.1105 | No | ||
| 33 | LMAN1 | ENSG00000074695.6 | 19261 | 0.058 | -0.1259 | No | ||
| 34 | TMPRSS2 | ENSG00000184012.12 | 19396 | 0.054 | -0.1275 | No | ||
| 35 | NDRG1 | ENSG00000104419.14 | 19578 | 0.049 | -0.1305 | No | ||
| 36 | SAT1 | ENSG00000130066.16 | 19807 | 0.043 | -0.1348 | No | ||
| 37 | ELK4 | ENSG00000158711.13 | 20194 | 0.034 | -0.1432 | No | ||
| 38 | IDI1 | ENSG00000067064.11 | 20552 | 0.023 | -0.1512 | No | ||
| 39 | PTK2B | ENSG00000120899.18 | 21620 | -0.004 | -0.1771 | No | ||
| 40 | AKT1 | ENSG00000142208.16 | 21695 | -0.006 | -0.1787 | No | ||
| 41 | HPGD | ENSG00000164120.14 | 22682 | -0.030 | -0.2018 | No | ||
| 42 | B2M | ENSG00000166710.20 | 22947 | -0.038 | -0.2071 | No | ||
| 43 | ABHD2 | ENSG00000140526.18 | 23280 | -0.046 | -0.2139 | No | ||
| 44 | MAK | ENSG00000111837.11 | 24172 | -0.069 | -0.2335 | No | ||
| 45 | APPBP2 | ENSG00000062725.10 | 24467 | -0.076 | -0.2385 | No | ||
| 46 | PIAS1 | ENSG00000033800.13 | 24784 | -0.085 | -0.2437 | No | ||
| 47 | SPDEF | ENSG00000124664.11 | 25184 | -0.096 | -0.2506 | No | ||
| 48 | MAP7 | ENSG00000135525.18 | 25553 | -0.107 | -0.2564 | No | ||
| 49 | XRCC6 | ENSG00000196419.12 | 26529 | -0.136 | -0.2762 | No | ||
| 50 | CAMKK2 | ENSG00000110931.19 | 27531 | -0.166 | -0.2957 | No | ||
| 51 | SEC24D | ENSG00000150961.15 | 27788 | -0.173 | -0.2969 | No | ||
| 52 | DBI | ENSG00000155368.16 | 27942 | -0.178 | -0.2954 | No | ||
| 53 | XRCC5 | ENSG00000079246.16 | 27984 | -0.180 | -0.2911 | No | ||
| 54 | NGLY1 | ENSG00000151092.17 | 28006 | -0.181 | -0.2863 | No | ||
| 55 | GSR | ENSG00000104687.14 | 28212 | -0.187 | -0.2858 | No | ||
| 56 | ACTN1 | ENSG00000072110.13 | 28265 | -0.188 | -0.2816 | No | ||
| 57 | VAPA | ENSG00000101558.13 | 28294 | -0.189 | -0.2767 | No | ||
| 58 | SLC26A2 | ENSG00000155850.7 | 29218 | -0.218 | -0.2928 | No | ||
| 59 | ADAMTS1 | ENSG00000154734.15 | 29777 | -0.237 | -0.2994 | No | ||
| 60 | UBE2J1 | ENSG00000198833.7 | 29873 | -0.240 | -0.2947 | No | ||
| 61 | LIFR | ENSG00000113594.10 | 29881 | -0.241 | -0.2878 | No | ||
| 62 | INPP4B | ENSG00000109452.12 | 30469 | -0.259 | -0.2945 | No | ||
| 63 | GUCY1A1 | ENSG00000164116.16 | 31161 | -0.283 | -0.3030 | No | ||
| 64 | SRP19 | ENSG00000153037.15 | 31205 | -0.285 | -0.2957 | No | ||
| 65 | SORD | ENSG00000140263.14 | 31730 | -0.287 | -0.3001 | No | ||
| 66 | PA2G4 | ENSG00000170515.14 | 31890 | -0.293 | -0.2953 | No | ||
| 67 | RRP12 | ENSG00000052749.13 | 31997 | -0.296 | -0.2892 | No | ||
| 68 | UAP1 | ENSG00000117143.13 | 32033 | -0.297 | -0.2814 | No | ||
| 69 | GNAI3 | ENSG00000065135.11 | 32448 | -0.315 | -0.2822 | No | ||
| 70 | CCND3 | ENSG00000112576.12 | 32532 | -0.317 | -0.2749 | No | ||
| 71 | RAB4A | ENSG00000168118.12 | 32600 | -0.320 | -0.2672 | No | ||
| 72 | ALDH1A3 | ENSG00000184254.17 | 34635 | -0.353 | -0.3063 | Yes | ||
| 73 | SMS | ENSG00000102172.16 | 35034 | -0.366 | -0.3053 | Yes | ||
| 74 | NKX3-1 | ENSG00000167034.10 | 35049 | -0.367 | -0.2949 | Yes | ||
| 75 | PGM3 | ENSG00000013375.16 | 35370 | -0.382 | -0.2914 | Yes | ||
| 76 | NCOA4 | ENSG00000266412.5 | 36372 | -0.420 | -0.3035 | Yes | ||
| 77 | CENPN | ENSG00000166451.13 | 36906 | -0.447 | -0.3034 | Yes | ||
| 78 | SRF | ENSG00000112658.8 | 36941 | -0.449 | -0.2911 | Yes | ||
| 79 | GPD1L | ENSG00000152642.10 | 36942 | -0.449 | -0.2779 | Yes | ||
| 80 | SGK1 | ENSG00000118515.11 | 37317 | -0.472 | -0.2732 | Yes | ||
| 81 | PLPP1 | ENSG00000067113.17 | 37340 | -0.474 | -0.2598 | Yes | ||
| 82 | HERC3 | ENSG00000138641.18 | 37427 | -0.479 | -0.2478 | Yes | ||
| 83 | AKAP12 | ENSG00000131016.17 | 37685 | -0.501 | -0.2394 | Yes | ||
| 84 | HOMER2 | ENSG00000103942.13 | 37764 | -0.508 | -0.2264 | Yes | ||
| 85 | ITGAV | ENSG00000138448.12 | 37784 | -0.511 | -0.2119 | Yes | ||
| 86 | CDC14B | ENSG00000081377.17 | 37879 | -0.518 | -0.1990 | Yes | ||
| 87 | B4GALT1 | ENSG00000086062.12 | 38033 | -0.530 | -0.1872 | Yes | ||
| 88 | CCND1 | ENSG00000110092.3 | 38307 | -0.552 | -0.1776 | Yes | ||
| 89 | ABCC4 | ENSG00000125257.16 | 39035 | -0.605 | -0.1776 | Yes | ||
| 90 | TNFAIP8 | ENSG00000145779.8 | 39080 | -0.611 | -0.1607 | Yes | ||
| 91 | TMEM50A | ENSG00000183726.11 | 39510 | -0.636 | -0.1525 | Yes | ||
| 92 | RPS6KA3 | ENSG00000177189.14 | 39524 | -0.639 | -0.1341 | Yes | ||
| 93 | STK39 | ENSG00000198648.11 | 39659 | -0.656 | -0.1182 | Yes | ||
| 94 | SPCS3 | ENSG00000129128.13 | 39807 | -0.676 | -0.1019 | Yes | ||
| 95 | PDLIM5 | ENSG00000163110.15 | 40169 | -0.746 | -0.0888 | Yes | ||
| 96 | CDK6 | ENSG00000105810.9 | 40555 | -0.858 | -0.0731 | Yes | ||
| 97 | BMPR1B | ENSG00000138696.11 | 40755 | -0.937 | -0.0504 | Yes | ||
| 98 | TSC22D1 | ENSG00000102804.15 | 40879 | -1.021 | -0.0235 | Yes | ||
| 99 | MYL12A | ENSG00000101608.12 | 40916 | -1.034 | 0.0060 | Yes |