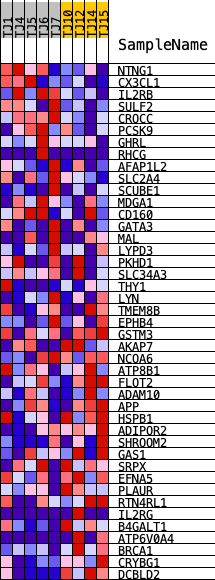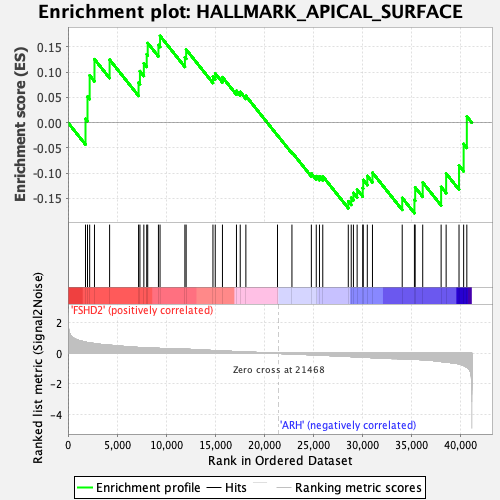
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_APICAL_SURFACE |
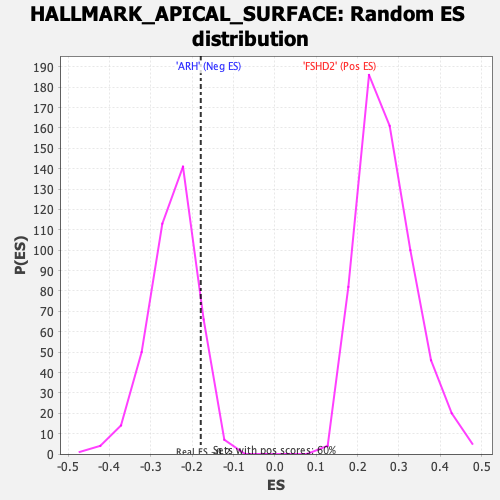
| Enrichment Score (ES) | -0.17921737 |
| Normalized Enrichment Score (NES) | -0.72730726 |
| Nominal p-value | 0.9040404 |
| FDR q-value | 0.9666066 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NTNG1 | ENSG00000162631.18 | 1784 | 0.736 | 0.0076 | Yes | ||
| 2 | CX3CL1 | ENSG00000006210.7 | 1988 | 0.707 | 0.0517 | Yes | ||
| 3 | IL2RB | ENSG00000100385.14 | 2208 | 0.683 | 0.0937 | Yes | ||
| 4 | SULF2 | ENSG00000196562.14 | 2699 | 0.634 | 0.1257 | Yes | ||
| 5 | CROCC | ENSG00000058453.17 | 4241 | 0.528 | 0.1248 | Yes | ||
| 6 | PCSK9 | ENSG00000169174.10 | 7193 | 0.381 | 0.0794 | Yes | ||
| 7 | GHRL | ENSG00000157017.16 | 7332 | 0.375 | 0.1021 | Yes | ||
| 8 | RHCG | ENSG00000140519.14 | 7731 | 0.363 | 0.1176 | Yes | ||
| 9 | AFAP1L2 | ENSG00000169129.15 | 8024 | 0.360 | 0.1354 | Yes | ||
| 10 | SLC2A4 | ENSG00000181856.15 | 8126 | 0.356 | 0.1577 | Yes | ||
| 11 | SCUBE1 | ENSG00000159307.19 | 9218 | 0.326 | 0.1538 | Yes | ||
| 12 | MDGA1 | ENSG00000112139.16 | 9382 | 0.320 | 0.1720 | Yes | ||
| 13 | CD160 | ENSG00000117281.16 | 11915 | 0.274 | 0.1294 | No | ||
| 14 | GATA3 | ENSG00000107485.18 | 12041 | 0.269 | 0.1450 | No | ||
| 15 | MAL | ENSG00000172005.11 | 14775 | 0.181 | 0.0911 | No | ||
| 16 | LYPD3 | ENSG00000124466.9 | 15009 | 0.173 | 0.0975 | No | ||
| 17 | PKHD1 | ENSG00000170927.15 | 15744 | 0.152 | 0.0901 | No | ||
| 18 | SLC34A3 | ENSG00000198569.9 | 17172 | 0.113 | 0.0632 | No | ||
| 19 | THY1 | ENSG00000154096.13 | 17556 | 0.102 | 0.0610 | No | ||
| 20 | LYN | ENSG00000254087.8 | 18130 | 0.087 | 0.0531 | No | ||
| 21 | TMEM8B | ENSG00000137103.20 | 21354 | 0.002 | -0.0251 | No | ||
| 22 | EPHB4 | ENSG00000196411.10 | 22826 | -0.034 | -0.0585 | No | ||
| 23 | GSTM3 | ENSG00000134202.11 | 24806 | -0.086 | -0.1007 | No | ||
| 24 | AKAP7 | ENSG00000118507.17 | 25306 | -0.099 | -0.1060 | No | ||
| 25 | NCOA6 | ENSG00000198646.14 | 25634 | -0.109 | -0.1063 | No | ||
| 26 | ATP8B1 | ENSG00000081923.14 | 25969 | -0.119 | -0.1062 | No | ||
| 27 | FLOT2 | ENSG00000132589.16 | 28570 | -0.198 | -0.1557 | No | ||
| 28 | ADAM10 | ENSG00000137845.15 | 28870 | -0.207 | -0.1486 | No | ||
| 29 | APP | ENSG00000142192.21 | 29094 | -0.215 | -0.1391 | No | ||
| 30 | HSPB1 | ENSG00000106211.9 | 29472 | -0.226 | -0.1326 | No | ||
| 31 | ADIPOR2 | ENSG00000006831.10 | 30043 | -0.245 | -0.1294 | No | ||
| 32 | SHROOM2 | ENSG00000146950.13 | 30092 | -0.247 | -0.1135 | No | ||
| 33 | GAS1 | ENSG00000180447.7 | 30508 | -0.260 | -0.1056 | No | ||
| 34 | SRPX | ENSG00000101955.15 | 31035 | -0.278 | -0.0991 | No | ||
| 35 | EFNA5 | ENSG00000184349.13 | 34067 | -0.343 | -0.1490 | No | ||
| 36 | PLAUR | ENSG00000011422.12 | 35312 | -0.379 | -0.1529 | No | ||
| 37 | RTN4RL1 | ENSG00000185924.7 | 35388 | -0.383 | -0.1282 | No | ||
| 38 | IL2RG | ENSG00000147168.12 | 36157 | -0.412 | -0.1184 | No | ||
| 39 | B4GALT1 | ENSG00000086062.12 | 38033 | -0.530 | -0.1272 | No | ||
| 40 | ATP6V0A4 | ENSG00000105929.16 | 38544 | -0.561 | -0.1008 | No | ||
| 41 | BRCA1 | ENSG00000012048.22 | 39857 | -0.687 | -0.0851 | No | ||
| 42 | CRYBG1 | ENSG00000112297.15 | 40329 | -0.784 | -0.0422 | No | ||
| 43 | DCBLD2 | ENSG00000057019.16 | 40653 | -0.900 | 0.0124 | No |