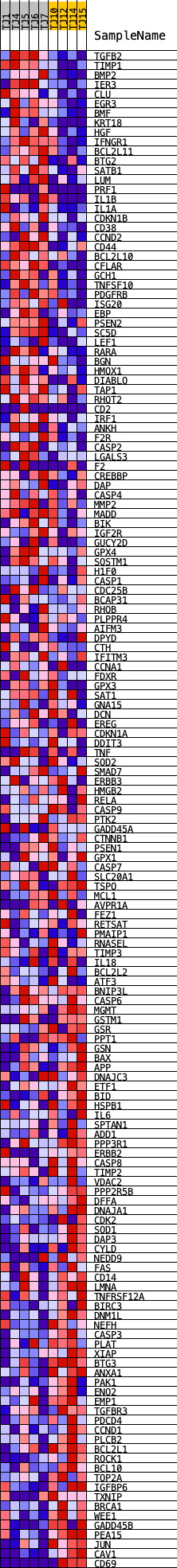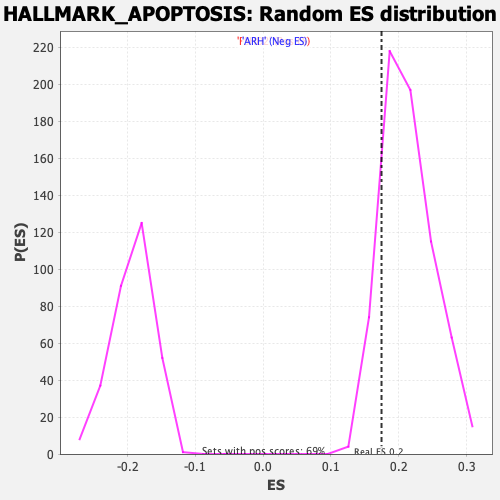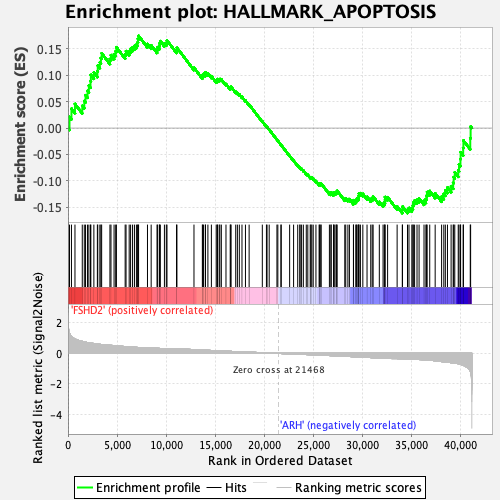
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_APOPTOSIS |
| Enrichment Score (ES) | 0.1745431 |
| Normalized Enrichment Score (NES) | 0.8181992 |
| Nominal p-value | 0.86151606 |
| FDR q-value | 0.8427977 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TGFB2 | ENSG00000092969.12 | 152 | 1.317 | 0.0211 | Yes | ||
| 2 | TIMP1 | ENSG00000102265.12 | 352 | 1.090 | 0.0368 | Yes | ||
| 3 | BMP2 | ENSG00000125845.7 | 710 | 0.936 | 0.0457 | Yes | ||
| 4 | IER3 | ENSG00000137331.12 | 1454 | 0.779 | 0.0422 | Yes | ||
| 5 | CLU | ENSG00000120885.22 | 1659 | 0.752 | 0.0514 | Yes | ||
| 6 | EGR3 | ENSG00000179388.9 | 1788 | 0.735 | 0.0621 | Yes | ||
| 7 | BMF | ENSG00000104081.14 | 1991 | 0.707 | 0.0705 | Yes | ||
| 8 | KRT18 | ENSG00000111057.11 | 2128 | 0.690 | 0.0802 | Yes | ||
| 9 | HGF | ENSG00000019991.18 | 2292 | 0.673 | 0.0889 | Yes | ||
| 10 | IFNGR1 | ENSG00000027697.14 | 2329 | 0.669 | 0.1006 | Yes | ||
| 11 | BCL2L11 | ENSG00000153094.23 | 2637 | 0.640 | 0.1052 | Yes | ||
| 12 | BTG2 | ENSG00000159388.6 | 2993 | 0.606 | 0.1079 | Yes | ||
| 13 | SATB1 | ENSG00000182568.17 | 3034 | 0.604 | 0.1183 | Yes | ||
| 14 | LUM | ENSG00000139329.5 | 3247 | 0.588 | 0.1242 | Yes | ||
| 15 | PRF1 | ENSG00000180644.8 | 3351 | 0.580 | 0.1326 | Yes | ||
| 16 | IL1B | ENSG00000125538.12 | 3430 | 0.572 | 0.1415 | Yes | ||
| 17 | IL1A | ENSG00000115008.5 | 4274 | 0.526 | 0.1309 | Yes | ||
| 18 | CDKN1B | ENSG00000111276.11 | 4380 | 0.521 | 0.1381 | Yes | ||
| 19 | CD38 | ENSG00000004468.13 | 4696 | 0.511 | 0.1400 | Yes | ||
| 20 | CCND2 | ENSG00000118971.8 | 4862 | 0.500 | 0.1454 | Yes | ||
| 21 | CD44 | ENSG00000026508.18 | 4934 | 0.495 | 0.1530 | Yes | ||
| 22 | BCL2L10 | ENSG00000137875.4 | 5825 | 0.442 | 0.1397 | Yes | ||
| 23 | CFLAR | ENSG00000003402.20 | 5916 | 0.438 | 0.1457 | Yes | ||
| 24 | GCH1 | ENSG00000131979.19 | 6236 | 0.421 | 0.1459 | Yes | ||
| 25 | TNFSF10 | ENSG00000121858.11 | 6380 | 0.413 | 0.1502 | Yes | ||
| 26 | PDGFRB | ENSG00000113721.14 | 6580 | 0.405 | 0.1529 | Yes | ||
| 27 | ISG20 | ENSG00000172183.15 | 6772 | 0.397 | 0.1557 | Yes | ||
| 28 | EBP | ENSG00000147155.11 | 6973 | 0.391 | 0.1582 | Yes | ||
| 29 | PSEN2 | ENSG00000143801.17 | 7101 | 0.385 | 0.1624 | Yes | ||
| 30 | SC5D | ENSG00000109929.10 | 7132 | 0.384 | 0.1689 | Yes | ||
| 31 | LEF1 | ENSG00000138795.10 | 7194 | 0.381 | 0.1745 | Yes | ||
| 32 | RARA | ENSG00000131759.18 | 8101 | 0.358 | 0.1592 | No | ||
| 33 | BGN | ENSG00000182492.16 | 8477 | 0.352 | 0.1567 | No | ||
| 34 | HMOX1 | ENSG00000100292.17 | 9075 | 0.332 | 0.1484 | No | ||
| 35 | DIABLO | ENSG00000184047.19 | 9135 | 0.329 | 0.1531 | No | ||
| 36 | TAP1 | ENSG00000168394.11 | 9314 | 0.323 | 0.1549 | No | ||
| 37 | RHOT2 | ENSG00000140983.14 | 9334 | 0.322 | 0.1605 | No | ||
| 38 | CD2 | ENSG00000116824.5 | 9412 | 0.319 | 0.1646 | No | ||
| 39 | IRF1 | ENSG00000125347.14 | 9828 | 0.318 | 0.1605 | No | ||
| 40 | ANKH | ENSG00000154122.14 | 10019 | 0.311 | 0.1617 | No | ||
| 41 | F2R | ENSG00000181104.7 | 10089 | 0.309 | 0.1658 | No | ||
| 42 | CASP2 | ENSG00000106144.20 | 11060 | 0.289 | 0.1476 | No | ||
| 43 | LGALS3 | ENSG00000131981.16 | 11103 | 0.287 | 0.1520 | No | ||
| 44 | F2 | ENSG00000180210.14 | 12841 | 0.243 | 0.1142 | No | ||
| 45 | CREBBP | ENSG00000005339.14 | 13694 | 0.216 | 0.0975 | No | ||
| 46 | DAP | ENSG00000112977.15 | 13727 | 0.215 | 0.1007 | No | ||
| 47 | CASP4 | ENSG00000196954.14 | 13818 | 0.213 | 0.1025 | No | ||
| 48 | MMP2 | ENSG00000087245.13 | 14001 | 0.206 | 0.1020 | No | ||
| 49 | MADD | ENSG00000110514.19 | 14013 | 0.205 | 0.1056 | No | ||
| 50 | BIK | ENSG00000100290.3 | 14260 | 0.198 | 0.1033 | No | ||
| 51 | IGF2R | ENSG00000197081.14 | 14615 | 0.187 | 0.0982 | No | ||
| 52 | GUCY2D | ENSG00000132518.6 | 15106 | 0.171 | 0.0895 | No | ||
| 53 | GPX4 | ENSG00000167468.17 | 15215 | 0.168 | 0.0900 | No | ||
| 54 | SQSTM1 | ENSG00000161011.20 | 15233 | 0.167 | 0.0927 | No | ||
| 55 | H1F0 | ENSG00000189060.5 | 15408 | 0.161 | 0.0915 | No | ||
| 56 | CASP1 | ENSG00000137752.24 | 15477 | 0.160 | 0.0929 | No | ||
| 57 | CDC25B | ENSG00000101224.17 | 15629 | 0.156 | 0.0921 | No | ||
| 58 | BCAP31 | ENSG00000185825.17 | 16103 | 0.142 | 0.0832 | No | ||
| 59 | RHOB | ENSG00000143878.10 | 16535 | 0.130 | 0.0752 | No | ||
| 60 | PLPPR4 | ENSG00000117600.12 | 16593 | 0.128 | 0.0762 | No | ||
| 61 | AIFM3 | ENSG00000183773.15 | 16607 | 0.128 | 0.0783 | No | ||
| 62 | DPYD | ENSG00000188641.13 | 17096 | 0.115 | 0.0686 | No | ||
| 63 | CTH | ENSG00000116761.12 | 17265 | 0.110 | 0.0665 | No | ||
| 64 | IFITM3 | ENSG00000142089.16 | 17463 | 0.105 | 0.0637 | No | ||
| 65 | CCNA1 | ENSG00000133101.10 | 17734 | 0.098 | 0.0590 | No | ||
| 66 | FDXR | ENSG00000161513.12 | 18088 | 0.088 | 0.0520 | No | ||
| 67 | GPX3 | ENSG00000211445.12 | 18462 | 0.080 | 0.0444 | No | ||
| 68 | SAT1 | ENSG00000130066.16 | 19807 | 0.043 | 0.0125 | No | ||
| 69 | GNA15 | ENSG00000060558.4 | 20226 | 0.033 | 0.0029 | No | ||
| 70 | DCN | ENSG00000011465.17 | 20288 | 0.032 | 0.0020 | No | ||
| 71 | EREG | ENSG00000124882.4 | 20508 | 0.024 | -0.0029 | No | ||
| 72 | CDKN1A | ENSG00000124762.13 | 21297 | 0.004 | -0.0220 | No | ||
| 73 | DDIT3 | ENSG00000175197.12 | 21390 | 0.001 | -0.0242 | No | ||
| 74 | TNF | ENSG00000232810.4 | 21705 | -0.006 | -0.0318 | No | ||
| 75 | SOD2 | ENSG00000112096.18 | 21731 | -0.007 | -0.0322 | No | ||
| 76 | SMAD7 | ENSG00000101665.9 | 22592 | -0.028 | -0.0527 | No | ||
| 77 | ERBB3 | ENSG00000065361.15 | 22995 | -0.039 | -0.0617 | No | ||
| 78 | HMGB2 | ENSG00000164104.12 | 23415 | -0.050 | -0.0710 | No | ||
| 79 | RELA | ENSG00000173039.19 | 23632 | -0.056 | -0.0752 | No | ||
| 80 | CASP9 | ENSG00000132906.18 | 23652 | -0.057 | -0.0746 | No | ||
| 81 | PTK2 | ENSG00000169398.19 | 23810 | -0.061 | -0.0773 | No | ||
| 82 | GADD45A | ENSG00000116717.13 | 23989 | -0.064 | -0.0804 | No | ||
| 83 | CTNNB1 | ENSG00000168036.18 | 24288 | -0.073 | -0.0863 | No | ||
| 84 | PSEN1 | ENSG00000080815.19 | 24443 | -0.076 | -0.0887 | No | ||
| 85 | GPX1 | ENSG00000233276.5 | 24689 | -0.082 | -0.0931 | No | ||
| 86 | CASP7 | ENSG00000165806.20 | 24757 | -0.084 | -0.0932 | No | ||
| 87 | SLC20A1 | ENSG00000144136.11 | 24853 | -0.087 | -0.0938 | No | ||
| 88 | TSPO | ENSG00000100300.18 | 24994 | -0.091 | -0.0955 | No | ||
| 89 | MCL1 | ENSG00000143384.13 | 25273 | -0.098 | -0.1005 | No | ||
| 90 | AVPR1A | ENSG00000166148.3 | 25596 | -0.108 | -0.1063 | No | ||
| 91 | FEZ1 | ENSG00000149557.14 | 25646 | -0.110 | -0.1054 | No | ||
| 92 | RETSAT | ENSG00000042445.14 | 25766 | -0.113 | -0.1062 | No | ||
| 93 | PMAIP1 | ENSG00000141682.11 | 25799 | -0.113 | -0.1048 | No | ||
| 94 | RNASEL | ENSG00000135828.11 | 26645 | -0.139 | -0.1228 | No | ||
| 95 | TIMP3 | ENSG00000100234.11 | 26802 | -0.144 | -0.1239 | No | ||
| 96 | IL18 | ENSG00000150782.12 | 26822 | -0.144 | -0.1217 | No | ||
| 97 | BCL2L2 | ENSG00000129473.9 | 27071 | -0.151 | -0.1249 | No | ||
| 98 | ATF3 | ENSG00000162772.17 | 27076 | -0.151 | -0.1221 | No | ||
| 99 | BNIP3L | ENSG00000104765.16 | 27191 | -0.155 | -0.1220 | No | ||
| 100 | CASP6 | ENSG00000138794.10 | 27327 | -0.159 | -0.1223 | No | ||
| 101 | MGMT | ENSG00000170430.10 | 27385 | -0.160 | -0.1206 | No | ||
| 102 | GSTM1 | ENSG00000134184.13 | 27430 | -0.162 | -0.1187 | No | ||
| 103 | GSR | ENSG00000104687.14 | 28212 | -0.187 | -0.1342 | No | ||
| 104 | PPT1 | ENSG00000131238.17 | 28310 | -0.190 | -0.1330 | No | ||
| 105 | GSN | ENSG00000148180.19 | 28530 | -0.197 | -0.1346 | No | ||
| 106 | BAX | ENSG00000087088.20 | 28685 | -0.202 | -0.1346 | No | ||
| 107 | APP | ENSG00000142192.21 | 29094 | -0.215 | -0.1405 | No | ||
| 108 | DNAJC3 | ENSG00000102580.15 | 29103 | -0.215 | -0.1366 | No | ||
| 109 | ETF1 | ENSG00000120705.13 | 29337 | -0.222 | -0.1381 | No | ||
| 110 | BID | ENSG00000015475.18 | 29396 | -0.225 | -0.1353 | No | ||
| 111 | HSPB1 | ENSG00000106211.9 | 29472 | -0.226 | -0.1329 | No | ||
| 112 | IL6 | ENSG00000136244.12 | 29624 | -0.231 | -0.1322 | No | ||
| 113 | SPTAN1 | ENSG00000197694.16 | 29628 | -0.232 | -0.1279 | No | ||
| 114 | ADD1 | ENSG00000087274.17 | 29654 | -0.233 | -0.1241 | No | ||
| 115 | PPP3R1 | ENSG00000221823.11 | 29805 | -0.238 | -0.1233 | No | ||
| 116 | ERBB2 | ENSG00000141736.13 | 30053 | -0.246 | -0.1247 | No | ||
| 117 | CASP8 | ENSG00000064012.21 | 30481 | -0.259 | -0.1302 | No | ||
| 118 | TIMP2 | ENSG00000035862.12 | 30859 | -0.272 | -0.1343 | No | ||
| 119 | VDAC2 | ENSG00000165637.13 | 31005 | -0.277 | -0.1326 | No | ||
| 120 | PPP2R5B | ENSG00000068971.14 | 31101 | -0.281 | -0.1297 | No | ||
| 121 | DFFA | ENSG00000160049.12 | 31733 | -0.287 | -0.1396 | No | ||
| 122 | DNAJA1 | ENSG00000086061.16 | 32115 | -0.301 | -0.1433 | No | ||
| 123 | CDK2 | ENSG00000123374.11 | 32246 | -0.307 | -0.1407 | No | ||
| 124 | SOD1 | ENSG00000142168.14 | 32307 | -0.310 | -0.1363 | No | ||
| 125 | DAP3 | ENSG00000132676.16 | 32312 | -0.310 | -0.1306 | No | ||
| 126 | CYLD | ENSG00000083799.17 | 32578 | -0.319 | -0.1310 | No | ||
| 127 | NEDD9 | ENSG00000111859.17 | 33548 | -0.338 | -0.1483 | No | ||
| 128 | FAS | ENSG00000026103.22 | 34075 | -0.344 | -0.1546 | No | ||
| 129 | CD14 | ENSG00000170458.14 | 34097 | -0.345 | -0.1487 | No | ||
| 130 | LMNA | ENSG00000160789.20 | 34605 | -0.352 | -0.1544 | No | ||
| 131 | TNFRSF12A | ENSG00000006327.13 | 34756 | -0.359 | -0.1513 | No | ||
| 132 | BIRC3 | ENSG00000023445.14 | 35047 | -0.367 | -0.1515 | No | ||
| 133 | DNM1L | ENSG00000087470.18 | 35155 | -0.372 | -0.1471 | No | ||
| 134 | NEFH | ENSG00000100285.10 | 35194 | -0.374 | -0.1410 | No | ||
| 135 | CASP3 | ENSG00000164305.19 | 35329 | -0.380 | -0.1371 | No | ||
| 136 | PLAT | ENSG00000104368.18 | 35576 | -0.392 | -0.1357 | No | ||
| 137 | XIAP | ENSG00000101966.12 | 35786 | -0.401 | -0.1332 | No | ||
| 138 | BTG3 | ENSG00000154640.14 | 36289 | -0.416 | -0.1376 | No | ||
| 139 | ANXA1 | ENSG00000135046.14 | 36480 | -0.425 | -0.1343 | No | ||
| 140 | PAK1 | ENSG00000149269.9 | 36549 | -0.430 | -0.1278 | No | ||
| 141 | ENO2 | ENSG00000111674.9 | 36606 | -0.431 | -0.1211 | No | ||
| 142 | EMP1 | ENSG00000134531.10 | 36872 | -0.444 | -0.1192 | No | ||
| 143 | TGFBR3 | ENSG00000069702.11 | 37426 | -0.479 | -0.1236 | No | ||
| 144 | PDCD4 | ENSG00000150593.18 | 38083 | -0.533 | -0.1296 | No | ||
| 145 | CCND1 | ENSG00000110092.3 | 38307 | -0.552 | -0.1246 | No | ||
| 146 | PLCB2 | ENSG00000137841.12 | 38462 | -0.558 | -0.1179 | No | ||
| 147 | BCL2L1 | ENSG00000171552.13 | 38679 | -0.568 | -0.1124 | No | ||
| 148 | ROCK1 | ENSG00000067900.8 | 39050 | -0.606 | -0.1101 | No | ||
| 149 | BCL10 | ENSG00000142867.14 | 39261 | -0.619 | -0.1035 | No | ||
| 150 | TOP2A | ENSG00000131747.15 | 39313 | -0.624 | -0.0930 | No | ||
| 151 | IGFBP6 | ENSG00000167779.9 | 39424 | -0.628 | -0.0839 | No | ||
| 152 | TXNIP | ENSG00000265972.6 | 39777 | -0.670 | -0.0798 | No | ||
| 153 | BRCA1 | ENSG00000012048.22 | 39857 | -0.687 | -0.0688 | No | ||
| 154 | WEE1 | ENSG00000166483.11 | 40002 | -0.714 | -0.0589 | No | ||
| 155 | GADD45B | ENSG00000099860.9 | 40019 | -0.716 | -0.0458 | No | ||
| 156 | PEA15 | ENSG00000162734.12 | 40282 | -0.772 | -0.0376 | No | ||
| 157 | JUN | ENSG00000177606.7 | 40305 | -0.779 | -0.0235 | No | ||
| 158 | CAV1 | ENSG00000105974.12 | 41003 | -1.149 | -0.0189 | No | ||
| 159 | CD69 | ENSG00000110848.8 | 41040 | -1.206 | 0.0030 | No |