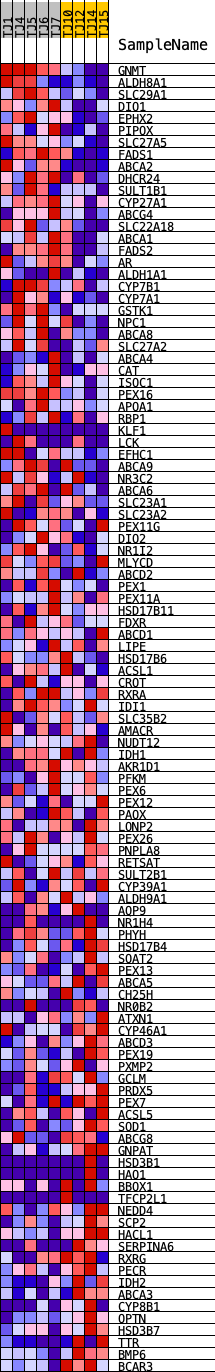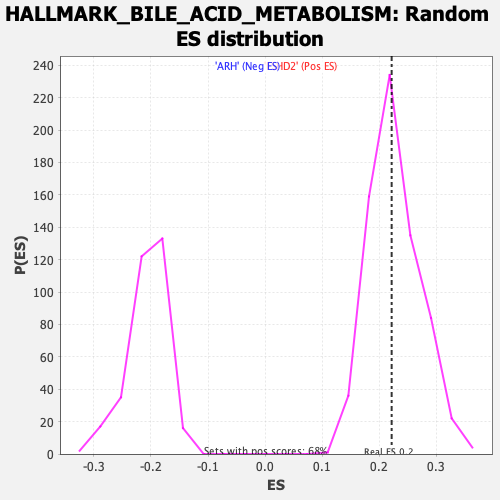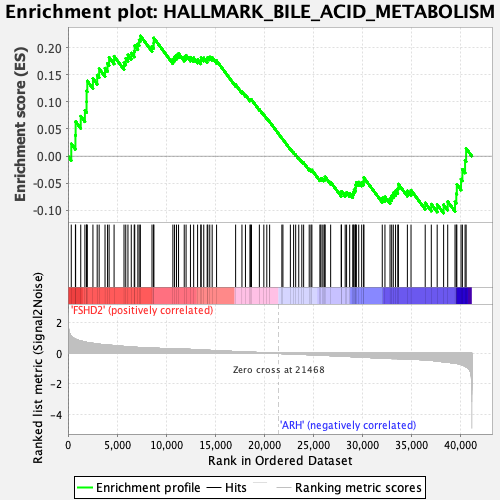
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_BILE_ACID_METABOLISM |
| Enrichment Score (ES) | 0.2215328 |
| Normalized Enrichment Score (NES) | 0.9763365 |
| Nominal p-value | 0.5051852 |
| FDR q-value | 0.56335896 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GNMT | ENSG00000124713.6 | 333 | 1.100 | 0.0228 | Yes | ||
| 2 | ALDH8A1 | ENSG00000118514.14 | 756 | 0.924 | 0.0385 | Yes | ||
| 3 | SLC29A1 | ENSG00000112759.18 | 778 | 0.917 | 0.0637 | Yes | ||
| 4 | DIO1 | ENSG00000211452.10 | 1299 | 0.803 | 0.0736 | Yes | ||
| 5 | EPHX2 | ENSG00000120915.14 | 1723 | 0.744 | 0.0842 | Yes | ||
| 6 | PIPOX | ENSG00000179761.12 | 1891 | 0.718 | 0.1003 | Yes | ||
| 7 | SLC27A5 | ENSG00000083807.10 | 1897 | 0.717 | 0.1203 | Yes | ||
| 8 | FADS1 | ENSG00000149485.19 | 1968 | 0.709 | 0.1385 | Yes | ||
| 9 | ABCA2 | ENSG00000107331.17 | 2541 | 0.650 | 0.1428 | Yes | ||
| 10 | DHCR24 | ENSG00000116133.13 | 2975 | 0.608 | 0.1493 | Yes | ||
| 11 | SULT1B1 | ENSG00000173597.9 | 3175 | 0.593 | 0.1611 | Yes | ||
| 12 | CYP27A1 | ENSG00000135929.9 | 3774 | 0.552 | 0.1621 | Yes | ||
| 13 | ABCG4 | ENSG00000172350.10 | 4031 | 0.539 | 0.1710 | Yes | ||
| 14 | SLC22A18 | ENSG00000110628.15 | 4190 | 0.531 | 0.1820 | Yes | ||
| 15 | ABCA1 | ENSG00000165029.16 | 4697 | 0.511 | 0.1840 | Yes | ||
| 16 | FADS2 | ENSG00000134824.14 | 5709 | 0.447 | 0.1720 | Yes | ||
| 17 | AR | ENSG00000169083.17 | 5886 | 0.439 | 0.1800 | Yes | ||
| 18 | ALDH1A1 | ENSG00000165092.13 | 6111 | 0.428 | 0.1866 | Yes | ||
| 19 | CYP7B1 | ENSG00000172817.4 | 6449 | 0.410 | 0.1899 | Yes | ||
| 20 | CYP7A1 | ENSG00000167910.4 | 6750 | 0.398 | 0.1938 | Yes | ||
| 21 | GSTK1 | ENSG00000197448.14 | 6802 | 0.396 | 0.2036 | Yes | ||
| 22 | NPC1 | ENSG00000141458.13 | 7120 | 0.384 | 0.2067 | Yes | ||
| 23 | ABCA8 | ENSG00000141338.14 | 7255 | 0.377 | 0.2140 | Yes | ||
| 24 | SLC27A2 | ENSG00000140284.11 | 7378 | 0.373 | 0.2215 | Yes | ||
| 25 | ABCA4 | ENSG00000198691.13 | 8563 | 0.349 | 0.2025 | No | ||
| 26 | CAT | ENSG00000121691.7 | 8736 | 0.346 | 0.2080 | No | ||
| 27 | ISOC1 | ENSG00000066583.12 | 8741 | 0.346 | 0.2176 | No | ||
| 28 | PEX16 | ENSG00000121680.16 | 10690 | 0.303 | 0.1787 | No | ||
| 29 | APOA1 | ENSG00000118137.9 | 10862 | 0.297 | 0.1829 | No | ||
| 30 | RBP1 | ENSG00000114115.10 | 11061 | 0.289 | 0.1862 | No | ||
| 31 | KLF1 | ENSG00000105610.5 | 11268 | 0.285 | 0.1891 | No | ||
| 32 | LCK | ENSG00000182866.17 | 11848 | 0.276 | 0.1828 | No | ||
| 33 | EFHC1 | ENSG00000096093.16 | 12036 | 0.269 | 0.1858 | No | ||
| 34 | ABCA9 | ENSG00000154258.17 | 12481 | 0.255 | 0.1822 | No | ||
| 35 | NR3C2 | ENSG00000151623.15 | 12809 | 0.244 | 0.1810 | No | ||
| 36 | ABCA6 | ENSG00000154262.13 | 13209 | 0.232 | 0.1778 | No | ||
| 37 | SLC23A1 | ENSG00000170482.17 | 13538 | 0.221 | 0.1761 | No | ||
| 38 | SLC23A2 | ENSG00000089057.15 | 13569 | 0.220 | 0.1815 | No | ||
| 39 | PEX11G | ENSG00000104883.8 | 13838 | 0.211 | 0.1809 | No | ||
| 40 | DIO2 | ENSG00000211448.12 | 14188 | 0.200 | 0.1780 | No | ||
| 41 | NR1I2 | ENSG00000144852.17 | 14251 | 0.198 | 0.1821 | No | ||
| 42 | MLYCD | ENSG00000103150.7 | 14450 | 0.192 | 0.1826 | No | ||
| 43 | ABCD2 | ENSG00000173208.4 | 14695 | 0.184 | 0.1819 | No | ||
| 44 | PEX1 | ENSG00000127980.16 | 15144 | 0.170 | 0.1757 | No | ||
| 45 | PEX11A | ENSG00000166821.9 | 17086 | 0.115 | 0.1317 | No | ||
| 46 | HSD17B11 | ENSG00000198189.11 | 17731 | 0.098 | 0.1187 | No | ||
| 47 | FDXR | ENSG00000161513.12 | 18088 | 0.088 | 0.1126 | No | ||
| 48 | ABCD1 | ENSG00000101986.12 | 18545 | 0.078 | 0.1036 | No | ||
| 49 | LIPE | ENSG00000079435.10 | 18614 | 0.076 | 0.1041 | No | ||
| 50 | HSD17B6 | ENSG00000025423.11 | 18703 | 0.073 | 0.1040 | No | ||
| 51 | ACSL1 | ENSG00000151726.14 | 19506 | 0.051 | 0.0859 | No | ||
| 52 | CROT | ENSG00000005469.11 | 19961 | 0.039 | 0.0760 | No | ||
| 53 | RXRA | ENSG00000186350.12 | 20264 | 0.032 | 0.0695 | No | ||
| 54 | IDI1 | ENSG00000067064.11 | 20552 | 0.023 | 0.0632 | No | ||
| 55 | SLC35B2 | ENSG00000157593.19 | 21800 | -0.009 | 0.0331 | No | ||
| 56 | AMACR | ENSG00000242110.8 | 21907 | -0.012 | 0.0308 | No | ||
| 57 | NUDT12 | ENSG00000112874.10 | 22666 | -0.030 | 0.0132 | No | ||
| 58 | IDH1 | ENSG00000138413.13 | 22996 | -0.039 | 0.0063 | No | ||
| 59 | AKR1D1 | ENSG00000122787.15 | 23207 | -0.045 | 0.0024 | No | ||
| 60 | PFKM | ENSG00000152556.17 | 23521 | -0.053 | -0.0037 | No | ||
| 61 | PEX6 | ENSG00000124587.14 | 23832 | -0.061 | -0.0095 | No | ||
| 62 | PEX12 | ENSG00000108733.10 | 24019 | -0.065 | -0.0122 | No | ||
| 63 | PAOX | ENSG00000148832.16 | 24587 | -0.080 | -0.0238 | No | ||
| 64 | LONP2 | ENSG00000102910.14 | 24767 | -0.085 | -0.0258 | No | ||
| 65 | PEX26 | ENSG00000215193.12 | 24859 | -0.087 | -0.0256 | No | ||
| 66 | PNPLA8 | ENSG00000135241.17 | 25677 | -0.110 | -0.0424 | No | ||
| 67 | RETSAT | ENSG00000042445.14 | 25766 | -0.113 | -0.0414 | No | ||
| 68 | SULT2B1 | ENSG00000088002.11 | 25907 | -0.117 | -0.0415 | No | ||
| 69 | CYP39A1 | ENSG00000146233.8 | 26087 | -0.123 | -0.0424 | No | ||
| 70 | ALDH9A1 | ENSG00000143149.12 | 26176 | -0.125 | -0.0410 | No | ||
| 71 | AQP9 | ENSG00000103569.10 | 26201 | -0.126 | -0.0381 | No | ||
| 72 | NR1H4 | ENSG00000012504.15 | 26772 | -0.143 | -0.0479 | No | ||
| 73 | PHYH | ENSG00000107537.14 | 27848 | -0.175 | -0.0692 | No | ||
| 74 | HSD17B4 | ENSG00000133835.16 | 27875 | -0.175 | -0.0649 | No | ||
| 75 | SOAT2 | ENSG00000167780.12 | 28254 | -0.188 | -0.0688 | No | ||
| 76 | PEX13 | ENSG00000162928.9 | 28396 | -0.193 | -0.0669 | No | ||
| 77 | ABCA5 | ENSG00000154265.16 | 28711 | -0.202 | -0.0688 | No | ||
| 78 | CH25H | ENSG00000138135.7 | 29028 | -0.213 | -0.0705 | No | ||
| 79 | NR0B2 | ENSG00000131910.5 | 29084 | -0.215 | -0.0659 | No | ||
| 80 | ATXN1 | ENSG00000124788.18 | 29169 | -0.217 | -0.0618 | No | ||
| 81 | CYP46A1 | ENSG00000036530.9 | 29305 | -0.221 | -0.0589 | No | ||
| 82 | ABCD3 | ENSG00000117528.14 | 29322 | -0.222 | -0.0530 | No | ||
| 83 | PEX19 | ENSG00000162735.18 | 29392 | -0.224 | -0.0484 | No | ||
| 84 | PXMP2 | ENSG00000176894.10 | 29634 | -0.232 | -0.0478 | No | ||
| 85 | GCLM | ENSG00000023909.10 | 29935 | -0.242 | -0.0483 | No | ||
| 86 | PRDX5 | ENSG00000126432.14 | 30144 | -0.249 | -0.0464 | No | ||
| 87 | PEX7 | ENSG00000112357.13 | 30151 | -0.249 | -0.0395 | No | ||
| 88 | ACSL5 | ENSG00000197142.10 | 32034 | -0.298 | -0.0770 | No | ||
| 89 | SOD1 | ENSG00000142168.14 | 32307 | -0.310 | -0.0749 | No | ||
| 90 | ABCG8 | ENSG00000143921.9 | 32826 | -0.329 | -0.0783 | No | ||
| 91 | GNPAT | ENSG00000116906.13 | 32995 | -0.336 | -0.0730 | No | ||
| 92 | HSD3B1 | ENSG00000203857.10 | 33158 | -0.336 | -0.0675 | No | ||
| 93 | HAO1 | ENSG00000101323.5 | 33389 | -0.336 | -0.0636 | No | ||
| 94 | BBOX1 | ENSG00000129151.9 | 33619 | -0.341 | -0.0596 | No | ||
| 95 | TFCP2L1 | ENSG00000115112.8 | 33677 | -0.342 | -0.0514 | No | ||
| 96 | NEDD4 | ENSG00000069869.16 | 34593 | -0.351 | -0.0639 | No | ||
| 97 | SCP2 | ENSG00000116171.18 | 34965 | -0.364 | -0.0627 | No | ||
| 98 | HACL1 | ENSG00000131373.15 | 36411 | -0.421 | -0.0861 | No | ||
| 99 | SERPINA6 | ENSG00000170099.6 | 37032 | -0.454 | -0.0884 | No | ||
| 100 | RXRG | ENSG00000143171.13 | 37631 | -0.496 | -0.0890 | No | ||
| 101 | PECR | ENSG00000115425.14 | 38287 | -0.550 | -0.0896 | No | ||
| 102 | IDH2 | ENSG00000182054.9 | 38704 | -0.571 | -0.0836 | No | ||
| 103 | ABCA3 | ENSG00000167972.14 | 39450 | -0.631 | -0.0841 | No | ||
| 104 | CYP8B1 | ENSG00000180432.6 | 39579 | -0.645 | -0.0691 | No | ||
| 105 | OPTN | ENSG00000123240.17 | 39634 | -0.651 | -0.0521 | No | ||
| 106 | HSD3B7 | ENSG00000099377.14 | 40063 | -0.724 | -0.0422 | No | ||
| 107 | TTR | ENSG00000118271.10 | 40201 | -0.753 | -0.0244 | No | ||
| 108 | BMP6 | ENSG00000153162.9 | 40475 | -0.830 | -0.0077 | No | ||
| 109 | BCAR3 | ENSG00000137936.18 | 40581 | -0.870 | 0.0141 | No |