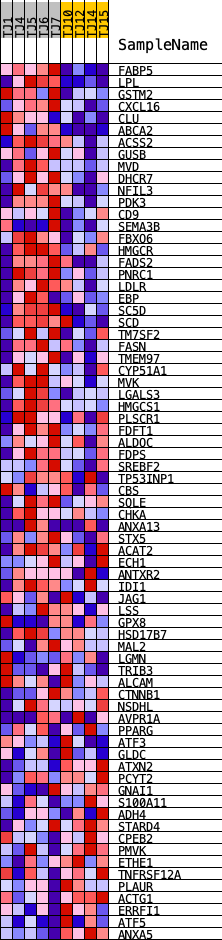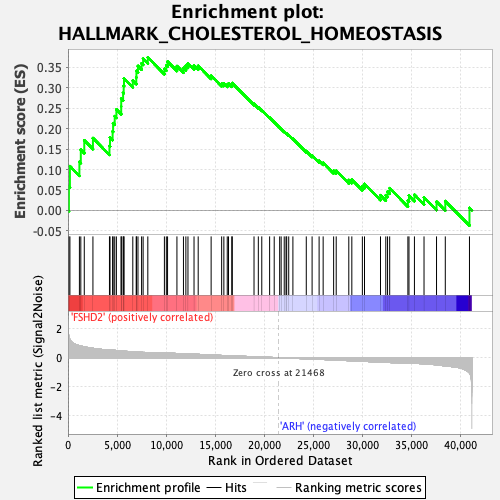
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_CHOLESTEROL_HOMEOSTASIS |
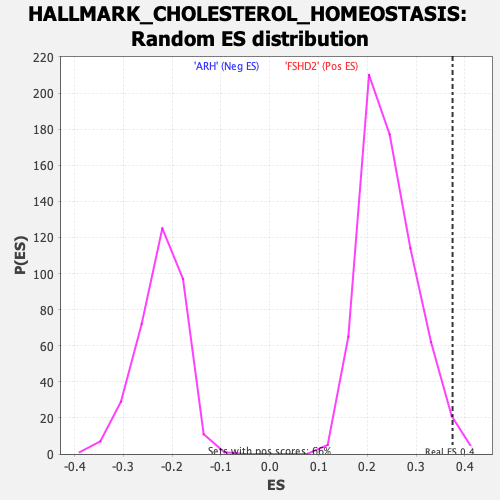
| Enrichment Score (ES) | 0.3746192 |
| Normalized Enrichment Score (NES) | 1.5384724 |
| Nominal p-value | 0.00913242 |
| FDR q-value | 0.04900585 |
| FWER p-Value | 0.219 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FABP5 | ENSG00000164687.11 | 92 | 1.467 | 0.0587 | Yes | ||
| 2 | LPL | ENSG00000175445.17 | 194 | 1.247 | 0.1081 | Yes | ||
| 3 | GSTM2 | ENSG00000213366.13 | 1162 | 0.824 | 0.1189 | Yes | ||
| 4 | CXCL16 | ENSG00000161921.15 | 1306 | 0.801 | 0.1487 | Yes | ||
| 5 | CLU | ENSG00000120885.22 | 1659 | 0.752 | 0.1714 | Yes | ||
| 6 | ABCA2 | ENSG00000107331.17 | 2541 | 0.650 | 0.1769 | Yes | ||
| 7 | ACSS2 | ENSG00000131069.20 | 4226 | 0.529 | 0.1579 | Yes | ||
| 8 | GUSB | ENSG00000169919.17 | 4282 | 0.525 | 0.1784 | Yes | ||
| 9 | MVD | ENSG00000167508.12 | 4547 | 0.516 | 0.1934 | Yes | ||
| 10 | DHCR7 | ENSG00000172893.15 | 4598 | 0.514 | 0.2135 | Yes | ||
| 11 | NFIL3 | ENSG00000165030.4 | 4763 | 0.507 | 0.2306 | Yes | ||
| 12 | PDK3 | ENSG00000067992.15 | 4932 | 0.495 | 0.2471 | Yes | ||
| 13 | CD9 | ENSG00000010278.14 | 5422 | 0.464 | 0.2545 | Yes | ||
| 14 | SEMA3B | ENSG00000012171.20 | 5425 | 0.463 | 0.2737 | Yes | ||
| 15 | FBXO6 | ENSG00000116663.11 | 5616 | 0.452 | 0.2879 | Yes | ||
| 16 | HMGCR | ENSG00000113161.16 | 5676 | 0.449 | 0.3051 | Yes | ||
| 17 | FADS2 | ENSG00000134824.14 | 5709 | 0.447 | 0.3229 | Yes | ||
| 18 | PNRC1 | ENSG00000146278.11 | 6605 | 0.403 | 0.3179 | Yes | ||
| 19 | LDLR | ENSG00000130164.13 | 6959 | 0.391 | 0.3256 | Yes | ||
| 20 | EBP | ENSG00000147155.11 | 6973 | 0.391 | 0.3415 | Yes | ||
| 21 | SC5D | ENSG00000109929.10 | 7132 | 0.384 | 0.3536 | Yes | ||
| 22 | SCD | ENSG00000099194.6 | 7501 | 0.371 | 0.3601 | Yes | ||
| 23 | TM7SF2 | ENSG00000149809.15 | 7658 | 0.364 | 0.3714 | Yes | ||
| 24 | FASN | ENSG00000169710.9 | 8136 | 0.356 | 0.3746 | Yes | ||
| 25 | TMEM97 | ENSG00000109084.14 | 9826 | 0.318 | 0.3467 | No | ||
| 26 | CYP51A1 | ENSG00000001630.17 | 10022 | 0.310 | 0.3549 | No | ||
| 27 | MVK | ENSG00000110921.14 | 10148 | 0.305 | 0.3646 | No | ||
| 28 | LGALS3 | ENSG00000131981.16 | 11103 | 0.287 | 0.3533 | No | ||
| 29 | HMGCS1 | ENSG00000112972.15 | 11768 | 0.279 | 0.3487 | No | ||
| 30 | PLSCR1 | ENSG00000188313.13 | 12014 | 0.270 | 0.3540 | No | ||
| 31 | FDFT1 | ENSG00000079459.13 | 12228 | 0.263 | 0.3598 | No | ||
| 32 | ALDOC | ENSG00000109107.14 | 12846 | 0.243 | 0.3548 | No | ||
| 33 | FDPS | ENSG00000160752.14 | 13271 | 0.229 | 0.3541 | No | ||
| 34 | SREBF2 | ENSG00000198911.12 | 14589 | 0.187 | 0.3298 | No | ||
| 35 | TP53INP1 | ENSG00000164938.14 | 15662 | 0.154 | 0.3101 | No | ||
| 36 | CBS | ENSG00000160200.17 | 15887 | 0.148 | 0.3108 | No | ||
| 37 | SQLE | ENSG00000104549.12 | 16232 | 0.139 | 0.3082 | No | ||
| 38 | CHKA | ENSG00000110721.12 | 16361 | 0.135 | 0.3107 | No | ||
| 39 | ANXA13 | ENSG00000104537.17 | 16693 | 0.125 | 0.3079 | No | ||
| 40 | STX5 | ENSG00000162236.12 | 16747 | 0.124 | 0.3117 | No | ||
| 41 | ACAT2 | ENSG00000120437.9 | 18961 | 0.066 | 0.2606 | No | ||
| 42 | ECH1 | ENSG00000104823.9 | 19400 | 0.054 | 0.2522 | No | ||
| 43 | ANTXR2 | ENSG00000163297.17 | 19752 | 0.045 | 0.2455 | No | ||
| 44 | IDI1 | ENSG00000067064.11 | 20552 | 0.023 | 0.2270 | No | ||
| 45 | JAG1 | ENSG00000101384.12 | 21019 | 0.012 | 0.2162 | No | ||
| 46 | LSS | ENSG00000160285.15 | 21576 | -0.003 | 0.2028 | No | ||
| 47 | GPX8 | ENSG00000164294.14 | 21726 | -0.007 | 0.1994 | No | ||
| 48 | HSD17B7 | ENSG00000132196.14 | 22040 | -0.015 | 0.1924 | No | ||
| 49 | MAL2 | ENSG00000147676.14 | 22223 | -0.019 | 0.1888 | No | ||
| 50 | LGMN | ENSG00000100600.15 | 22291 | -0.020 | 0.1880 | No | ||
| 51 | TRIB3 | ENSG00000101255.11 | 22498 | -0.027 | 0.1841 | No | ||
| 52 | ALCAM | ENSG00000170017.12 | 22924 | -0.037 | 0.1753 | No | ||
| 53 | CTNNB1 | ENSG00000168036.18 | 24288 | -0.073 | 0.1451 | No | ||
| 54 | NSDHL | ENSG00000147383.11 | 24881 | -0.088 | 0.1344 | No | ||
| 55 | AVPR1A | ENSG00000166148.3 | 25596 | -0.108 | 0.1215 | No | ||
| 56 | PPARG | ENSG00000132170.21 | 26008 | -0.121 | 0.1165 | No | ||
| 57 | ATF3 | ENSG00000162772.17 | 27076 | -0.151 | 0.0968 | No | ||
| 58 | GLDC | ENSG00000178445.9 | 27339 | -0.159 | 0.0971 | No | ||
| 59 | ATXN2 | ENSG00000204842.17 | 28622 | -0.200 | 0.0742 | No | ||
| 60 | PCYT2 | ENSG00000185813.10 | 28931 | -0.210 | 0.0754 | No | ||
| 61 | GNAI1 | ENSG00000127955.17 | 30005 | -0.244 | 0.0594 | No | ||
| 62 | S100A11 | ENSG00000163191.6 | 30223 | -0.251 | 0.0646 | No | ||
| 63 | ADH4 | ENSG00000198099.8 | 31853 | -0.292 | 0.0370 | No | ||
| 64 | STARD4 | ENSG00000164211.13 | 32393 | -0.313 | 0.0370 | No | ||
| 65 | CPEB2 | ENSG00000137449.16 | 32572 | -0.319 | 0.0459 | No | ||
| 66 | PMVK | ENSG00000163344.6 | 32790 | -0.327 | 0.0542 | No | ||
| 67 | ETHE1 | ENSG00000105755.8 | 34645 | -0.353 | 0.0238 | No | ||
| 68 | TNFRSF12A | ENSG00000006327.13 | 34756 | -0.359 | 0.0360 | No | ||
| 69 | PLAUR | ENSG00000011422.12 | 35312 | -0.379 | 0.0383 | No | ||
| 70 | ACTG1 | ENSG00000184009.12 | 36290 | -0.416 | 0.0318 | No | ||
| 71 | ERRFI1 | ENSG00000116285.13 | 37565 | -0.490 | 0.0211 | No | ||
| 72 | ATF5 | ENSG00000169136.11 | 38458 | -0.558 | 0.0226 | No | ||
| 73 | ANXA5 | ENSG00000164111.15 | 40922 | -1.038 | 0.0058 | No |