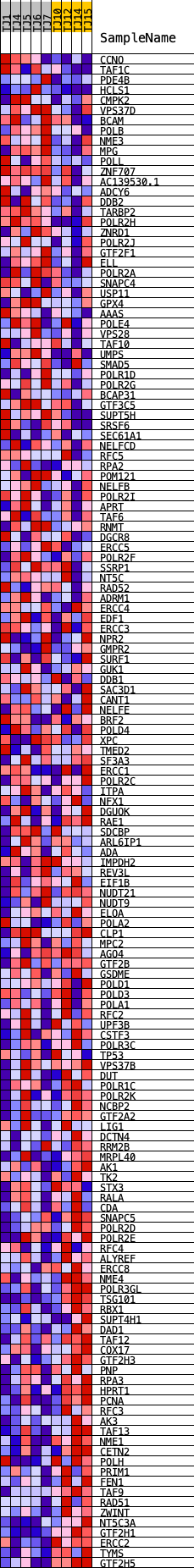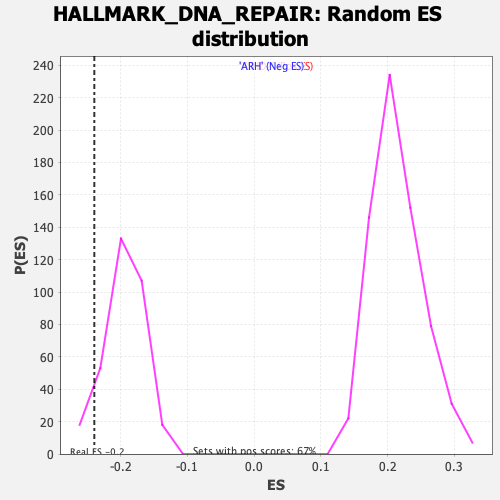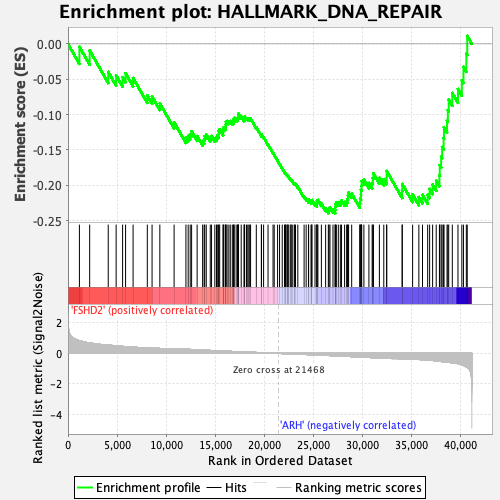
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_DNA_REPAIR |
| Enrichment Score (ES) | -0.2394628 |
| Normalized Enrichment Score (NES) | -1.2323029 |
| Nominal p-value | 0.075987846 |
| FDR q-value | 0.13616782 |
| FWER p-Value | 0.777 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CCNO | ENSG00000152669.9 | 1163 | 0.824 | -0.0040 | No | ||
| 2 | TAF1C | ENSG00000103168.16 | 2209 | 0.683 | -0.0093 | No | ||
| 3 | PDE4B | ENSG00000184588.18 | 4101 | 0.537 | -0.0395 | No | ||
| 4 | HCLS1 | ENSG00000180353.11 | 4907 | 0.497 | -0.0445 | No | ||
| 5 | CMPK2 | ENSG00000134326.11 | 5565 | 0.456 | -0.0470 | No | ||
| 6 | VPS37D | ENSG00000176428.6 | 5870 | 0.440 | -0.0415 | No | ||
| 7 | BCAM | ENSG00000187244.11 | 6634 | 0.402 | -0.0482 | No | ||
| 8 | POLB | ENSG00000070501.12 | 8084 | 0.358 | -0.0729 | No | ||
| 9 | NME3 | ENSG00000103024.7 | 8569 | 0.348 | -0.0744 | No | ||
| 10 | MPG | ENSG00000103152.11 | 9361 | 0.320 | -0.0843 | No | ||
| 11 | POLL | ENSG00000166169.17 | 10815 | 0.298 | -0.1109 | No | ||
| 12 | ZNF707 | ENSG00000181135.16 | 12020 | 0.270 | -0.1323 | No | ||
| 13 | AC139530.1 | ENSG00000262049.1 | 12250 | 0.263 | -0.1301 | No | ||
| 14 | ADCY6 | ENSG00000174233.11 | 12456 | 0.256 | -0.1275 | No | ||
| 15 | DDB2 | ENSG00000134574.11 | 12599 | 0.251 | -0.1236 | No | ||
| 16 | TARBP2 | ENSG00000139546.11 | 13162 | 0.233 | -0.1304 | No | ||
| 17 | POLR2H | ENSG00000163882.9 | 13722 | 0.215 | -0.1376 | No | ||
| 18 | ZNRD1 | ENSG00000066379.15 | 13876 | 0.210 | -0.1352 | No | ||
| 19 | POLR2J | ENSG00000005075.15 | 13937 | 0.208 | -0.1305 | No | ||
| 20 | GTF2F1 | ENSG00000125651.14 | 14092 | 0.203 | -0.1283 | No | ||
| 21 | ELL | ENSG00000105656.13 | 14500 | 0.191 | -0.1325 | No | ||
| 22 | POLR2A | ENSG00000181222.16 | 14631 | 0.186 | -0.1302 | No | ||
| 23 | SNAPC4 | ENSG00000165684.4 | 14967 | 0.175 | -0.1332 | No | ||
| 24 | USP11 | ENSG00000102226.9 | 15137 | 0.170 | -0.1323 | No | ||
| 25 | GPX4 | ENSG00000167468.17 | 15215 | 0.168 | -0.1292 | No | ||
| 26 | AAAS | ENSG00000094914.14 | 15362 | 0.163 | -0.1280 | No | ||
| 27 | POLE4 | ENSG00000115350.12 | 15368 | 0.163 | -0.1233 | No | ||
| 28 | VPS28 | ENSG00000160948.14 | 15461 | 0.160 | -0.1208 | No | ||
| 29 | TAF10 | ENSG00000166337.10 | 15814 | 0.150 | -0.1250 | No | ||
| 30 | UMPS | ENSG00000114491.14 | 15817 | 0.150 | -0.1206 | No | ||
| 31 | SMAD5 | ENSG00000113658.18 | 15888 | 0.148 | -0.1179 | No | ||
| 32 | POLR1D | ENSG00000186184.18 | 16056 | 0.144 | -0.1178 | No | ||
| 33 | POLR2G | ENSG00000168002.12 | 16079 | 0.143 | -0.1141 | No | ||
| 34 | BCAP31 | ENSG00000185825.17 | 16103 | 0.142 | -0.1104 | No | ||
| 35 | GTF3C5 | ENSG00000148308.17 | 16214 | 0.139 | -0.1090 | No | ||
| 36 | SUPT5H | ENSG00000196235.14 | 16395 | 0.134 | -0.1094 | No | ||
| 37 | SRSF6 | ENSG00000124193.16 | 16541 | 0.130 | -0.1091 | No | ||
| 38 | SEC61A1 | ENSG00000058262.10 | 16746 | 0.124 | -0.1104 | No | ||
| 39 | NELFCD | ENSG00000101158.15 | 16860 | 0.121 | -0.1096 | No | ||
| 40 | RFC5 | ENSG00000111445.14 | 16883 | 0.120 | -0.1066 | No | ||
| 41 | RPA2 | ENSG00000117748.10 | 16945 | 0.118 | -0.1046 | No | ||
| 42 | POM121 | ENSG00000196313.11 | 17139 | 0.114 | -0.1059 | No | ||
| 43 | NELFB | ENSG00000188986.7 | 17280 | 0.110 | -0.1061 | No | ||
| 44 | POLR2I | ENSG00000105258.9 | 17330 | 0.109 | -0.1040 | No | ||
| 45 | APRT | ENSG00000198931.11 | 17367 | 0.108 | -0.1017 | No | ||
| 46 | TAF6 | ENSG00000106290.15 | 17371 | 0.108 | -0.0986 | No | ||
| 47 | RNMT | ENSG00000101654.17 | 17655 | 0.100 | -0.1026 | No | ||
| 48 | DGCR8 | ENSG00000128191.16 | 17938 | 0.092 | -0.1067 | No | ||
| 49 | ERCC5 | ENSG00000134899.21 | 17990 | 0.091 | -0.1053 | No | ||
| 50 | POLR2F | ENSG00000100142.15 | 18001 | 0.091 | -0.1028 | No | ||
| 51 | SSRP1 | ENSG00000149136.9 | 18191 | 0.086 | -0.1049 | No | ||
| 52 | NT5C | ENSG00000125458.7 | 18324 | 0.083 | -0.1057 | No | ||
| 53 | RAD52 | ENSG00000002016.18 | 18437 | 0.081 | -0.1060 | No | ||
| 54 | ADRM1 | ENSG00000130706.13 | 18532 | 0.078 | -0.1060 | No | ||
| 55 | ERCC4 | ENSG00000175595.14 | 18610 | 0.076 | -0.1056 | No | ||
| 56 | EDF1 | ENSG00000107223.13 | 19194 | 0.060 | -0.1181 | No | ||
| 57 | ERCC3 | ENSG00000163161.13 | 19714 | 0.046 | -0.1294 | No | ||
| 58 | NPR2 | ENSG00000159899.14 | 19718 | 0.045 | -0.1281 | No | ||
| 59 | GMPR2 | ENSG00000100938.17 | 19938 | 0.040 | -0.1323 | No | ||
| 60 | SURF1 | ENSG00000148290.10 | 20388 | 0.028 | -0.1424 | No | ||
| 61 | GUK1 | ENSG00000143774.16 | 20900 | 0.015 | -0.1544 | No | ||
| 62 | DDB1 | ENSG00000167986.13 | 21018 | 0.012 | -0.1569 | No | ||
| 63 | SAC3D1 | ENSG00000168061.15 | 21362 | 0.002 | -0.1652 | No | ||
| 64 | CANT1 | ENSG00000171302.17 | 21563 | -0.003 | -0.1700 | No | ||
| 65 | NELFE | ENSG00000204356.14 | 21837 | -0.011 | -0.1764 | No | ||
| 66 | BRF2 | ENSG00000104221.13 | 22069 | -0.016 | -0.1815 | No | ||
| 67 | POLD4 | ENSG00000175482.9 | 22165 | -0.017 | -0.1834 | No | ||
| 68 | XPC | ENSG00000154767.14 | 22181 | -0.018 | -0.1832 | No | ||
| 69 | TMED2 | ENSG00000086598.11 | 22331 | -0.022 | -0.1862 | No | ||
| 70 | SF3A3 | ENSG00000183431.12 | 22336 | -0.022 | -0.1856 | No | ||
| 71 | ERCC1 | ENSG00000012061.15 | 22428 | -0.024 | -0.1871 | No | ||
| 72 | POLR2C | ENSG00000102978.13 | 22510 | -0.027 | -0.1883 | No | ||
| 73 | ITPA | ENSG00000125877.13 | 22703 | -0.031 | -0.1921 | No | ||
| 74 | NFX1 | ENSG00000086102.19 | 22748 | -0.032 | -0.1922 | No | ||
| 75 | DGUOK | ENSG00000114956.20 | 22858 | -0.035 | -0.1938 | No | ||
| 76 | RAE1 | ENSG00000101146.13 | 23056 | -0.041 | -0.1974 | No | ||
| 77 | SDCBP | ENSG00000137575.12 | 23102 | -0.042 | -0.1973 | No | ||
| 78 | ARL6IP1 | ENSG00000170540.15 | 23188 | -0.045 | -0.1980 | No | ||
| 79 | ADA | ENSG00000196839.13 | 23425 | -0.051 | -0.2023 | No | ||
| 80 | IMPDH2 | ENSG00000178035.12 | 24066 | -0.066 | -0.2160 | No | ||
| 81 | REV3L | ENSG00000009413.15 | 24276 | -0.072 | -0.2189 | No | ||
| 82 | EIF1B | ENSG00000114784.4 | 24510 | -0.077 | -0.2223 | No | ||
| 83 | NUDT21 | ENSG00000167005.14 | 24528 | -0.078 | -0.2204 | No | ||
| 84 | NUDT9 | ENSG00000170502.13 | 24770 | -0.085 | -0.2238 | No | ||
| 85 | ELOA | ENSG00000011007.12 | 24844 | -0.087 | -0.2230 | No | ||
| 86 | POLA2 | ENSG00000014138.9 | 24862 | -0.087 | -0.2209 | No | ||
| 87 | CLP1 | ENSG00000172409.6 | 25179 | -0.095 | -0.2257 | No | ||
| 88 | MPC2 | ENSG00000143158.11 | 25364 | -0.101 | -0.2272 | No | ||
| 89 | AGO4 | ENSG00000134698.11 | 25374 | -0.101 | -0.2245 | No | ||
| 90 | GTF2B | ENSG00000137947.12 | 25384 | -0.102 | -0.2217 | No | ||
| 91 | GSDME | ENSG00000105928.16 | 25472 | -0.104 | -0.2207 | No | ||
| 92 | POLD1 | ENSG00000062822.14 | 25827 | -0.115 | -0.2260 | No | ||
| 93 | POLD3 | ENSG00000077514.9 | 26261 | -0.128 | -0.2327 | No | ||
| 94 | POLA1 | ENSG00000101868.11 | 26537 | -0.136 | -0.2354 | Yes | ||
| 95 | RFC2 | ENSG00000049541.11 | 26606 | -0.138 | -0.2330 | Yes | ||
| 96 | UPF3B | ENSG00000125351.11 | 26703 | -0.141 | -0.2312 | Yes | ||
| 97 | CSTF3 | ENSG00000176102.13 | 27011 | -0.149 | -0.2342 | Yes | ||
| 98 | POLR3C | ENSG00000186141.9 | 27226 | -0.156 | -0.2349 | Yes | ||
| 99 | TP53 | ENSG00000141510.17 | 27245 | -0.156 | -0.2307 | Yes | ||
| 100 | VPS37B | ENSG00000139722.7 | 27258 | -0.157 | -0.2263 | Yes | ||
| 101 | DUT | ENSG00000128951.14 | 27355 | -0.160 | -0.2240 | Yes | ||
| 102 | POLR1C | ENSG00000171453.20 | 27553 | -0.166 | -0.2239 | Yes | ||
| 103 | POLR2K | ENSG00000147669.11 | 27760 | -0.172 | -0.2238 | Yes | ||
| 104 | NCBP2 | ENSG00000114503.11 | 27882 | -0.176 | -0.2215 | Yes | ||
| 105 | GTF2A2 | ENSG00000140307.11 | 28194 | -0.186 | -0.2236 | Yes | ||
| 106 | LIG1 | ENSG00000105486.14 | 28401 | -0.193 | -0.2230 | Yes | ||
| 107 | DCTN4 | ENSG00000132912.12 | 28497 | -0.197 | -0.2195 | Yes | ||
| 108 | RRM2B | ENSG00000048392.11 | 28542 | -0.198 | -0.2147 | Yes | ||
| 109 | MRPL40 | ENSG00000185608.8 | 28602 | -0.199 | -0.2102 | Yes | ||
| 110 | AK1 | ENSG00000106992.19 | 28916 | -0.209 | -0.2117 | Yes | ||
| 111 | TK2 | ENSG00000166548.16 | 29736 | -0.236 | -0.2247 | Yes | ||
| 112 | STX3 | ENSG00000166900.17 | 29784 | -0.237 | -0.2188 | Yes | ||
| 113 | RALA | ENSG00000006451.8 | 29860 | -0.240 | -0.2136 | Yes | ||
| 114 | CDA | ENSG00000158825.6 | 29864 | -0.240 | -0.2066 | Yes | ||
| 115 | SNAPC5 | ENSG00000174446.13 | 29914 | -0.241 | -0.2006 | Yes | ||
| 116 | POLR2D | ENSG00000144231.11 | 29937 | -0.242 | -0.1940 | Yes | ||
| 117 | POLR2E | ENSG00000099817.12 | 30161 | -0.249 | -0.1921 | Yes | ||
| 118 | RFC4 | ENSG00000163918.10 | 30672 | -0.265 | -0.1967 | Yes | ||
| 119 | ALYREF | ENSG00000183684.8 | 31002 | -0.277 | -0.1965 | Yes | ||
| 120 | ERCC8 | ENSG00000049167.14 | 31065 | -0.279 | -0.1898 | Yes | ||
| 121 | NME4 | ENSG00000103202.13 | 31135 | -0.282 | -0.1831 | Yes | ||
| 122 | POLR3GL | ENSG00000121851.13 | 31740 | -0.287 | -0.1894 | Yes | ||
| 123 | TSG101 | ENSG00000074319.13 | 32189 | -0.304 | -0.1913 | Yes | ||
| 124 | RBX1 | ENSG00000100387.9 | 32483 | -0.316 | -0.1891 | Yes | ||
| 125 | SUPT4H1 | ENSG00000213246.7 | 32484 | -0.316 | -0.1798 | Yes | ||
| 126 | DAD1 | ENSG00000129562.11 | 34059 | -0.343 | -0.2080 | Yes | ||
| 127 | TAF12 | ENSG00000120656.11 | 34072 | -0.344 | -0.1982 | Yes | ||
| 128 | COX17 | ENSG00000138495.6 | 35127 | -0.370 | -0.2129 | Yes | ||
| 129 | GTF2H3 | ENSG00000111358.14 | 35764 | -0.400 | -0.2166 | Yes | ||
| 130 | PNP | ENSG00000198805.11 | 36135 | -0.411 | -0.2135 | Yes | ||
| 131 | RPA3 | ENSG00000106399.11 | 36661 | -0.434 | -0.2134 | Yes | ||
| 132 | HPRT1 | ENSG00000165704.15 | 36862 | -0.444 | -0.2052 | Yes | ||
| 133 | PCNA | ENSG00000132646.11 | 37166 | -0.461 | -0.1990 | Yes | ||
| 134 | RFC3 | ENSG00000133119.13 | 37532 | -0.487 | -0.1935 | Yes | ||
| 135 | AK3 | ENSG00000147853.17 | 37860 | -0.517 | -0.1862 | Yes | ||
| 136 | TAF13 | ENSG00000197780.10 | 37892 | -0.520 | -0.1716 | Yes | ||
| 137 | NME1 | ENSG00000239672.7 | 38038 | -0.530 | -0.1594 | Yes | ||
| 138 | CETN2 | ENSG00000147400.9 | 38135 | -0.537 | -0.1459 | Yes | ||
| 139 | POLH | ENSG00000170734.11 | 38286 | -0.550 | -0.1333 | Yes | ||
| 140 | PRIM1 | ENSG00000198056.15 | 38325 | -0.554 | -0.1178 | Yes | ||
| 141 | FEN1 | ENSG00000168496.4 | 38636 | -0.565 | -0.1087 | Yes | ||
| 142 | TAF9 | ENSG00000273841.5 | 38720 | -0.572 | -0.0938 | Yes | ||
| 143 | RAD51 | ENSG00000051180.17 | 38806 | -0.582 | -0.0787 | Yes | ||
| 144 | ZWINT | ENSG00000122952.17 | 39173 | -0.616 | -0.0694 | Yes | ||
| 145 | NT5C3A | ENSG00000122643.21 | 39751 | -0.667 | -0.0638 | Yes | ||
| 146 | GTF2H1 | ENSG00000110768.12 | 40144 | -0.740 | -0.0515 | Yes | ||
| 147 | ERCC2 | ENSG00000104884.15 | 40308 | -0.779 | -0.0324 | Yes | ||
| 148 | TYMS | ENSG00000176890.16 | 40602 | -0.877 | -0.0136 | Yes | ||
| 149 | GTF2H5 | ENSG00000272047.2 | 40695 | -0.922 | 0.0114 | Yes |