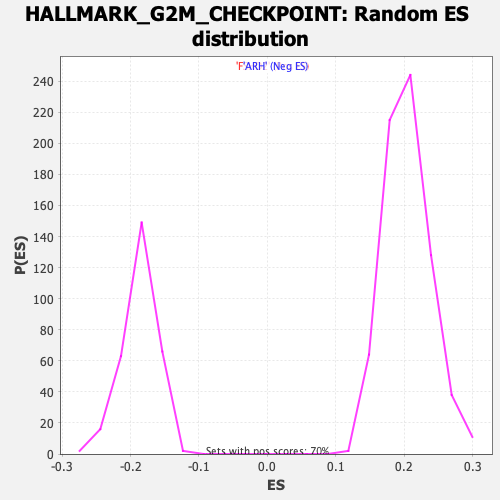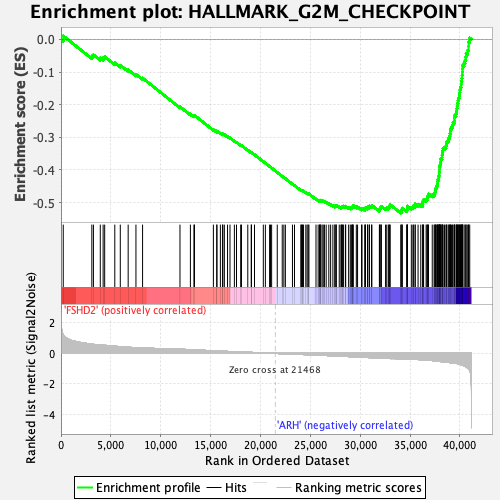
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_G2M_CHECKPOINT |
| Enrichment Score (ES) | -0.5322757 |
| Normalized Enrichment Score (NES) | -2.8461115 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HOXC10 | ENSG00000180818.5 | 225 | 1.204 | 0.0107 | No | ||
| 2 | FOXN3 | ENSG00000053254.15 | 3092 | 0.600 | -0.0512 | No | ||
| 3 | ABL1 | ENSG00000097007.19 | 3251 | 0.588 | -0.0471 | No | ||
| 4 | MT2A | ENSG00000125148.7 | 3939 | 0.543 | -0.0566 | No | ||
| 5 | BCL3 | ENSG00000069399.15 | 4224 | 0.529 | -0.0564 | No | ||
| 6 | CDKN1B | ENSG00000111276.11 | 4380 | 0.521 | -0.0531 | No | ||
| 7 | HIST1H2BK | ENSG00000197903.7 | 5394 | 0.465 | -0.0716 | No | ||
| 8 | NUMA1 | ENSG00000137497.17 | 5945 | 0.436 | -0.0792 | No | ||
| 9 | TLE3 | ENSG00000140332.16 | 6735 | 0.399 | -0.0930 | No | ||
| 10 | LBR | ENSG00000143815.15 | 7515 | 0.370 | -0.1071 | No | ||
| 11 | SAP30 | ENSG00000164105.4 | 8180 | 0.354 | -0.1185 | No | ||
| 12 | SLC38A1 | ENSG00000111371.16 | 11930 | 0.274 | -0.2063 | No | ||
| 13 | HMGN2 | ENSG00000198830.11 | 12975 | 0.238 | -0.2286 | No | ||
| 14 | NASP | ENSG00000132780.17 | 13329 | 0.227 | -0.2342 | No | ||
| 15 | MTF2 | ENSG00000143033.18 | 13387 | 0.226 | -0.2325 | No | ||
| 16 | AC027237.1 | ENSG00000259191.2 | 15280 | 0.165 | -0.2765 | No | ||
| 17 | RPS6KA5 | ENSG00000100784.12 | 15626 | 0.156 | -0.2828 | No | ||
| 18 | CDC25B | ENSG00000101224.17 | 15629 | 0.156 | -0.2808 | No | ||
| 19 | ODF2 | ENSG00000136811.16 | 15981 | 0.146 | -0.2874 | No | ||
| 20 | PDS5B | ENSG00000083642.19 | 16191 | 0.140 | -0.2906 | No | ||
| 21 | SQLE | ENSG00000104549.12 | 16232 | 0.139 | -0.2897 | No | ||
| 22 | EGF | ENSG00000138798.13 | 16379 | 0.135 | -0.2915 | No | ||
| 23 | TENT4A | ENSG00000112941.14 | 16700 | 0.125 | -0.2976 | No | ||
| 24 | RPA2 | ENSG00000117748.10 | 16945 | 0.118 | -0.3019 | No | ||
| 25 | UPF1 | ENSG00000005007.12 | 17385 | 0.107 | -0.3112 | No | ||
| 26 | LIG3 | ENSG00000005156.12 | 17599 | 0.101 | -0.3151 | No | ||
| 27 | HMGA1 | ENSG00000137309.19 | 18018 | 0.090 | -0.3240 | No | ||
| 28 | CHMP1A | ENSG00000131165.15 | 18097 | 0.088 | -0.3248 | No | ||
| 29 | RAD21 | ENSG00000164754.14 | 18730 | 0.073 | -0.3392 | No | ||
| 30 | PRPF4B | ENSG00000112739.17 | 19074 | 0.063 | -0.3467 | No | ||
| 31 | E2F4 | ENSG00000205250.9 | 19102 | 0.062 | -0.3466 | No | ||
| 32 | POLE | ENSG00000177084.17 | 19403 | 0.054 | -0.3532 | No | ||
| 33 | YTHDC1 | ENSG00000083896.12 | 20287 | 0.032 | -0.3743 | No | ||
| 34 | NOLC1 | ENSG00000166197.16 | 20493 | 0.025 | -0.3790 | No | ||
| 35 | CBX1 | ENSG00000108468.15 | 20917 | 0.014 | -0.3891 | No | ||
| 36 | SFPQ | ENSG00000116560.11 | 21006 | 0.012 | -0.3911 | No | ||
| 37 | PTTG3P | ENSG00000213005.3 | 21121 | 0.008 | -0.3938 | No | ||
| 38 | SMC4 | ENSG00000113810.16 | 21685 | -0.006 | -0.4074 | No | ||
| 39 | CTCF | ENSG00000102974.16 | 22185 | -0.018 | -0.4194 | No | ||
| 40 | HNRNPD | ENSG00000138668.19 | 22348 | -0.022 | -0.4230 | No | ||
| 41 | TRA2B | ENSG00000136527.18 | 22499 | -0.027 | -0.4263 | No | ||
| 42 | NUSAP1 | ENSG00000137804.14 | 23210 | -0.045 | -0.4431 | No | ||
| 43 | ORC5 | ENSG00000164815.11 | 23413 | -0.050 | -0.4473 | No | ||
| 44 | CCNT1 | ENSG00000129315.11 | 24047 | -0.066 | -0.4619 | No | ||
| 45 | PML | ENSG00000140464.20 | 24111 | -0.067 | -0.4625 | No | ||
| 46 | SMARCC1 | ENSG00000173473.11 | 24189 | -0.069 | -0.4635 | No | ||
| 47 | WRN | ENSG00000165392.10 | 24224 | -0.071 | -0.4633 | No | ||
| 48 | SMAD3 | ENSG00000166949.16 | 24321 | -0.074 | -0.4647 | No | ||
| 49 | CUL1 | ENSG00000055130.17 | 24550 | -0.079 | -0.4692 | No | ||
| 50 | EWSR1 | ENSG00000182944.17 | 24678 | -0.082 | -0.4712 | No | ||
| 51 | TNPO2 | ENSG00000105576.15 | 24787 | -0.085 | -0.4727 | No | ||
| 52 | POLA2 | ENSG00000014138.9 | 24862 | -0.087 | -0.4733 | No | ||
| 53 | DDX39A | ENSG00000123136.15 | 25570 | -0.107 | -0.4891 | No | ||
| 54 | SLC7A5 | ENSG00000103257.9 | 25788 | -0.113 | -0.4929 | No | ||
| 55 | TOP1 | ENSG00000198900.6 | 25928 | -0.118 | -0.4947 | No | ||
| 56 | CUL3 | ENSG00000036257.13 | 25973 | -0.119 | -0.4942 | No | ||
| 57 | DBF4 | ENSG00000006634.8 | 25980 | -0.120 | -0.4927 | No | ||
| 58 | ILF3 | ENSG00000129351.17 | 26047 | -0.122 | -0.4927 | No | ||
| 59 | DMD | ENSG00000198947.15 | 26181 | -0.125 | -0.4942 | No | ||
| 60 | CDK4 | ENSG00000135446.17 | 26296 | -0.129 | -0.4953 | No | ||
| 61 | MAPK14 | ENSG00000112062.10 | 26416 | -0.133 | -0.4964 | No | ||
| 62 | H2AFX | ENSG00000188486.3 | 26590 | -0.138 | -0.4988 | No | ||
| 63 | TGFB1 | ENSG00000105329.10 | 26855 | -0.145 | -0.5033 | No | ||
| 64 | CUL5 | ENSG00000166266.14 | 27037 | -0.150 | -0.5057 | No | ||
| 65 | FANCC | ENSG00000158169.13 | 27263 | -0.157 | -0.5090 | No | ||
| 66 | KATNA1 | ENSG00000186625.14 | 27460 | -0.163 | -0.5116 | No | ||
| 67 | ATRX | ENSG00000085224.22 | 27481 | -0.164 | -0.5099 | No | ||
| 68 | RBM14 | ENSG00000239306.4 | 27595 | -0.167 | -0.5104 | No | ||
| 69 | HNRNPU | ENSG00000153187.20 | 27597 | -0.167 | -0.5082 | No | ||
| 70 | SUV39H1 | ENSG00000101945.16 | 27901 | -0.176 | -0.5132 | No | ||
| 71 | PRMT5 | ENSG00000100462.16 | 28066 | -0.183 | -0.5147 | No | ||
| 72 | NOTCH2 | ENSG00000134250.20 | 28133 | -0.185 | -0.5139 | No | ||
| 73 | NUP98 | ENSG00000110713.17 | 28185 | -0.186 | -0.5126 | No | ||
| 74 | STAG1 | ENSG00000118007.13 | 28245 | -0.188 | -0.5115 | No | ||
| 75 | SRSF10 | ENSG00000188529.15 | 28319 | -0.190 | -0.5107 | No | ||
| 76 | HIRA | ENSG00000100084.14 | 28517 | -0.197 | -0.5129 | No | ||
| 77 | NCL | ENSG00000115053.16 | 28548 | -0.198 | -0.5110 | No | ||
| 78 | MARCKS | ENSG00000277443.3 | 28843 | -0.207 | -0.5154 | No | ||
| 79 | MCM3 | ENSG00000112118.20 | 29007 | -0.212 | -0.5165 | No | ||
| 80 | SRSF2 | ENSG00000161547.16 | 29115 | -0.215 | -0.5162 | No | ||
| 81 | G3BP1 | ENSG00000145907.15 | 29198 | -0.217 | -0.5153 | No | ||
| 82 | HIF1A | ENSG00000100644.17 | 29230 | -0.218 | -0.5131 | No | ||
| 83 | DR1 | ENSG00000117505.13 | 29261 | -0.220 | -0.5109 | No | ||
| 84 | PURA | ENSG00000185129.7 | 29308 | -0.221 | -0.5090 | No | ||
| 85 | EZH2 | ENSG00000106462.10 | 29613 | -0.231 | -0.5133 | No | ||
| 86 | CDC27 | ENSG00000004897.12 | 29738 | -0.236 | -0.5132 | No | ||
| 87 | E2F2 | ENSG00000007968.7 | 30138 | -0.249 | -0.5196 | No | ||
| 88 | CKS2 | ENSG00000123975.5 | 30183 | -0.250 | -0.5173 | No | ||
| 89 | CKS1B | ENSG00000173207.13 | 30473 | -0.259 | -0.5208 | No | ||
| 90 | CDKN3 | ENSG00000100526.20 | 30484 | -0.259 | -0.5176 | No | ||
| 91 | MEIS1 | ENSG00000143995.20 | 30535 | -0.260 | -0.5153 | No | ||
| 92 | CDKN2C | ENSG00000123080.12 | 30743 | -0.267 | -0.5167 | No | ||
| 93 | CASP8AP2 | ENSG00000118412.12 | 30749 | -0.268 | -0.5133 | No | ||
| 94 | ARID4A | ENSG00000032219.19 | 30913 | -0.274 | -0.5135 | No | ||
| 95 | KIF22 | ENSG00000079616.12 | 30925 | -0.274 | -0.5101 | No | ||
| 96 | MAP3K20 | ENSG00000091436.17 | 31138 | -0.282 | -0.5115 | No | ||
| 97 | KMT5A | ENSG00000183955.13 | 31173 | -0.284 | -0.5085 | No | ||
| 98 | SYNCRIP | ENSG00000135316.18 | 31926 | -0.294 | -0.5229 | No | ||
| 99 | PTTG1 | ENSG00000164611.13 | 31965 | -0.295 | -0.5198 | No | ||
| 100 | KPNB1 | ENSG00000108424.10 | 31990 | -0.296 | -0.5165 | No | ||
| 101 | BUB3 | ENSG00000154473.18 | 32087 | -0.300 | -0.5148 | No | ||
| 102 | INCENP | ENSG00000149503.13 | 32126 | -0.301 | -0.5116 | No | ||
| 103 | MCM2 | ENSG00000073111.14 | 32546 | -0.318 | -0.5176 | No | ||
| 104 | HMGB3 | ENSG00000029993.15 | 32666 | -0.323 | -0.5161 | No | ||
| 105 | DKC1 | ENSG00000130826.18 | 32829 | -0.329 | -0.5157 | No | ||
| 106 | TMPO | ENSG00000120802.13 | 32905 | -0.332 | -0.5130 | No | ||
| 107 | ODC1 | ENSG00000115758.13 | 32958 | -0.334 | -0.5098 | No | ||
| 108 | MEIS2 | ENSG00000134138.20 | 33007 | -0.336 | -0.5064 | No | ||
| 109 | EFNA5 | ENSG00000184349.13 | 34067 | -0.343 | -0.5276 | Yes | ||
| 110 | CUL4A | ENSG00000139842.15 | 34180 | -0.348 | -0.5257 | Yes | ||
| 111 | CDC20 | ENSG00000117399.14 | 34187 | -0.349 | -0.5211 | Yes | ||
| 112 | SRSF1 | ENSG00000136450.13 | 34211 | -0.350 | -0.5170 | Yes | ||
| 113 | HUS1 | ENSG00000136273.13 | 34665 | -0.355 | -0.5233 | Yes | ||
| 114 | H2AFV | ENSG00000105968.18 | 34723 | -0.357 | -0.5199 | Yes | ||
| 115 | MCM6 | ENSG00000076003.5 | 34728 | -0.357 | -0.5151 | Yes | ||
| 116 | SNRPD1 | ENSG00000167088.11 | 34734 | -0.358 | -0.5104 | Yes | ||
| 117 | H2AFZ | ENSG00000164032.12 | 35129 | -0.370 | -0.5151 | Yes | ||
| 118 | TACC3 | ENSG00000013810.19 | 35296 | -0.378 | -0.5140 | Yes | ||
| 119 | MNAT1 | ENSG00000020426.11 | 35332 | -0.380 | -0.5098 | Yes | ||
| 120 | TFDP1 | ENSG00000198176.13 | 35512 | -0.389 | -0.5089 | Yes | ||
| 121 | CCNF | ENSG00000162063.13 | 35514 | -0.389 | -0.5037 | Yes | ||
| 122 | MCM5 | ENSG00000100297.16 | 35805 | -0.402 | -0.5053 | Yes | ||
| 123 | PAFAH1B1 | ENSG00000007168.13 | 36076 | -0.408 | -0.5064 | Yes | ||
| 124 | BIRC5 | ENSG00000089685.15 | 36279 | -0.415 | -0.5058 | Yes | ||
| 125 | ESPL1 | ENSG00000135476.11 | 36287 | -0.415 | -0.5004 | Yes | ||
| 126 | DTYMK | ENSG00000168393.13 | 36297 | -0.416 | -0.4950 | Yes | ||
| 127 | TROAP | ENSG00000135451.13 | 36359 | -0.419 | -0.4908 | Yes | ||
| 128 | CCNB2 | ENSG00000157456.8 | 36613 | -0.432 | -0.4912 | Yes | ||
| 129 | PLK1 | ENSG00000166851.15 | 36707 | -0.436 | -0.4876 | Yes | ||
| 130 | TRAIP | ENSG00000183763.9 | 36731 | -0.437 | -0.4822 | Yes | ||
| 131 | UBE2S | ENSG00000108106.14 | 36803 | -0.440 | -0.4780 | Yes | ||
| 132 | KIF2C | ENSG00000142945.13 | 36852 | -0.443 | -0.4733 | Yes | ||
| 133 | XPO1 | ENSG00000082898.17 | 37227 | -0.465 | -0.4761 | Yes | ||
| 134 | HMMR | ENSG00000072571.20 | 37412 | -0.479 | -0.4742 | Yes | ||
| 135 | AURKB | ENSG00000178999.13 | 37485 | -0.484 | -0.4694 | Yes | ||
| 136 | RBL1 | ENSG00000080839.12 | 37521 | -0.487 | -0.4637 | Yes | ||
| 137 | CHAF1A | ENSG00000167670.16 | 37553 | -0.489 | -0.4579 | Yes | ||
| 138 | KIF4A | ENSG00000090889.12 | 37615 | -0.495 | -0.4527 | Yes | ||
| 139 | NUP50 | ENSG00000093000.19 | 37691 | -0.501 | -0.4478 | Yes | ||
| 140 | BUB1 | ENSG00000169679.15 | 37763 | -0.508 | -0.4427 | Yes | ||
| 141 | SLC12A2 | ENSG00000064651.14 | 37771 | -0.510 | -0.4360 | Yes | ||
| 142 | FBXO5 | ENSG00000112029.10 | 37782 | -0.511 | -0.4293 | Yes | ||
| 143 | HSPA8 | ENSG00000109971.14 | 37882 | -0.519 | -0.4248 | Yes | ||
| 144 | MAD2L1 | ENSG00000164109.14 | 37893 | -0.520 | -0.4180 | Yes | ||
| 145 | SMC1A | ENSG00000072501.17 | 37922 | -0.522 | -0.4117 | Yes | ||
| 146 | NDC80 | ENSG00000080986.13 | 37927 | -0.522 | -0.4047 | Yes | ||
| 147 | GINS2 | ENSG00000131153.9 | 37954 | -0.524 | -0.3983 | Yes | ||
| 148 | NEK2 | ENSG00000117650.13 | 37958 | -0.525 | -0.3913 | Yes | ||
| 149 | CENPA | ENSG00000115163.15 | 37993 | -0.527 | -0.3850 | Yes | ||
| 150 | CDC7 | ENSG00000097046.13 | 38052 | -0.531 | -0.3793 | Yes | ||
| 151 | E2F1 | ENSG00000101412.13 | 38085 | -0.533 | -0.3729 | Yes | ||
| 152 | UBE2C | ENSG00000175063.17 | 38091 | -0.534 | -0.3658 | Yes | ||
| 153 | KIF15 | ENSG00000163808.17 | 38204 | -0.543 | -0.3612 | Yes | ||
| 154 | TTK | ENSG00000112742.10 | 38213 | -0.544 | -0.3541 | Yes | ||
| 155 | NSD2 | ENSG00000109685.18 | 38242 | -0.546 | -0.3474 | Yes | ||
| 156 | CHEK1 | ENSG00000149554.13 | 38279 | -0.549 | -0.3409 | Yes | ||
| 157 | CCND1 | ENSG00000110092.3 | 38307 | -0.552 | -0.3341 | Yes | ||
| 158 | ATF5 | ENSG00000169136.11 | 38458 | -0.558 | -0.3303 | Yes | ||
| 159 | PBK | ENSG00000168078.10 | 38606 | -0.562 | -0.3263 | Yes | ||
| 160 | AURKA | ENSG00000087586.17 | 38638 | -0.565 | -0.3194 | Yes | ||
| 161 | TPX2 | ENSG00000088325.16 | 38681 | -0.569 | -0.3128 | Yes | ||
| 162 | CDC45 | ENSG00000093009.10 | 38822 | -0.583 | -0.3084 | Yes | ||
| 163 | ORC6 | ENSG00000091651.9 | 38888 | -0.589 | -0.3020 | Yes | ||
| 164 | STMN1 | ENSG00000117632.23 | 38971 | -0.598 | -0.2960 | Yes | ||
| 165 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.2886 | Yes | ||
| 166 | RAD54L | ENSG00000085999.13 | 39046 | -0.606 | -0.2815 | Yes | ||
| 167 | SMC2 | ENSG00000136824.19 | 39061 | -0.608 | -0.2737 | Yes | ||
| 168 | EXO1 | ENSG00000174371.17 | 39163 | -0.615 | -0.2678 | Yes | ||
| 169 | PLK4 | ENSG00000142731.11 | 39267 | -0.620 | -0.2620 | Yes | ||
| 170 | TOP2A | ENSG00000131747.15 | 39313 | -0.624 | -0.2547 | Yes | ||
| 171 | CENPF | ENSG00000117724.13 | 39444 | -0.631 | -0.2494 | Yes | ||
| 172 | CDK1 | ENSG00000170312.16 | 39453 | -0.632 | -0.2411 | Yes | ||
| 173 | RACGAP1 | ENSG00000161800.13 | 39469 | -0.633 | -0.2329 | Yes | ||
| 174 | SLC7A1 | ENSG00000139514.13 | 39608 | -0.647 | -0.2276 | Yes | ||
| 175 | GSPT1 | ENSG00000103342.13 | 39655 | -0.655 | -0.2199 | Yes | ||
| 176 | BARD1 | ENSG00000138376.11 | 39685 | -0.660 | -0.2117 | Yes | ||
| 177 | LMNB1 | ENSG00000113368.12 | 39725 | -0.665 | -0.2037 | Yes | ||
| 178 | PRC1 | ENSG00000198901.14 | 39769 | -0.668 | -0.1957 | Yes | ||
| 179 | MYBL2 | ENSG00000101057.16 | 39815 | -0.677 | -0.1877 | Yes | ||
| 180 | BRCA2 | ENSG00000139618.15 | 39873 | -0.690 | -0.1798 | Yes | ||
| 181 | PRIM2 | ENSG00000146143.19 | 39937 | -0.702 | -0.1719 | Yes | ||
| 182 | JPT1 | ENSG00000189159.16 | 39972 | -0.708 | -0.1632 | Yes | ||
| 183 | POLQ | ENSG00000051341.14 | 40007 | -0.714 | -0.1544 | Yes | ||
| 184 | CDC6 | ENSG00000094804.12 | 40060 | -0.723 | -0.1459 | Yes | ||
| 185 | KNL1 | ENSG00000137812.19 | 40136 | -0.739 | -0.1378 | Yes | ||
| 186 | CENPE | ENSG00000138778.12 | 40145 | -0.740 | -0.1280 | Yes | ||
| 187 | AMD1 | ENSG00000123505.17 | 40193 | -0.751 | -0.1191 | Yes | ||
| 188 | MKI67 | ENSG00000148773.14 | 40220 | -0.757 | -0.1095 | Yes | ||
| 189 | RASAL2 | ENSG00000075391.16 | 40239 | -0.762 | -0.0997 | Yes | ||
| 190 | CDC25A | ENSG00000164045.12 | 40263 | -0.768 | -0.0899 | Yes | ||
| 191 | E2F3 | ENSG00000112242.15 | 40268 | -0.769 | -0.0796 | Yes | ||
| 192 | STIL | ENSG00000123473.15 | 40411 | -0.811 | -0.0722 | Yes | ||
| 193 | KIF11 | ENSG00000138160.6 | 40519 | -0.845 | -0.0634 | Yes | ||
| 194 | CCNA2 | ENSG00000145386.10 | 40598 | -0.874 | -0.0536 | Yes | ||
| 195 | KPNA2 | ENSG00000182481.9 | 40625 | -0.887 | -0.0422 | Yes | ||
| 196 | KIF5B | ENSG00000170759.11 | 40785 | -0.954 | -0.0333 | Yes | ||
| 197 | UCK2 | ENSG00000143179.16 | 40869 | -1.012 | -0.0217 | Yes | ||
| 198 | KIF20B | ENSG00000138182.14 | 40873 | -1.017 | -0.0080 | Yes | ||
| 199 | RAD23B | ENSG00000119318.13 | 40986 | -1.117 | 0.0043 | Yes |