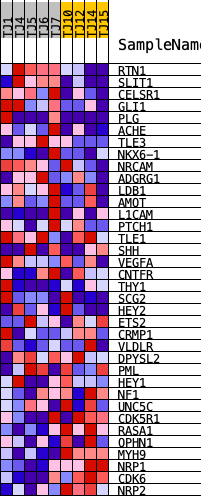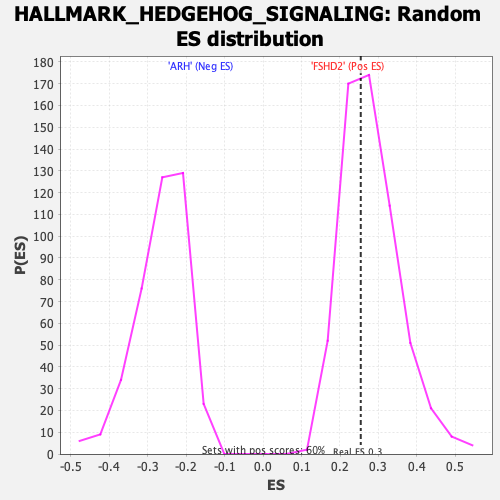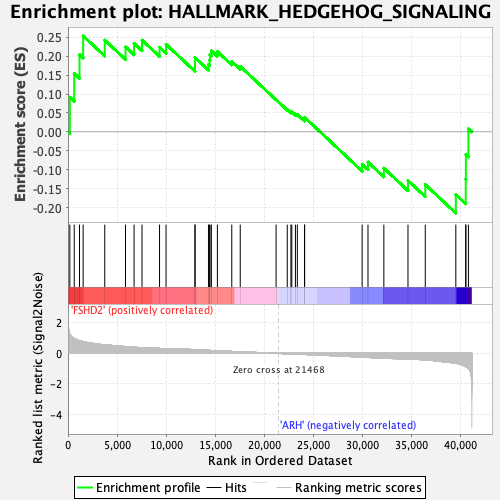
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_HEDGEHOG_SIGNALING |
| Enrichment Score (ES) | 0.25396535 |
| Normalized Enrichment Score (NES) | 0.9060668 |
| Nominal p-value | 0.6006711 |
| FDR q-value | 0.7011138 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RTN1 | ENSG00000139970.17 | 188 | 1.262 | 0.0918 | Yes | ||
| 2 | SLIT1 | ENSG00000187122.17 | 635 | 0.962 | 0.1545 | Yes | ||
| 3 | CELSR1 | ENSG00000075275.16 | 1172 | 0.823 | 0.2044 | Yes | ||
| 4 | GLI1 | ENSG00000111087.10 | 1543 | 0.767 | 0.2540 | Yes | ||
| 5 | PLG | ENSG00000122194.18 | 3746 | 0.554 | 0.2427 | No | ||
| 6 | ACHE | ENSG00000087085.15 | 5861 | 0.441 | 0.2250 | No | ||
| 7 | TLE3 | ENSG00000140332.16 | 6735 | 0.399 | 0.2343 | No | ||
| 8 | NKX6-1 | ENSG00000163623.9 | 7545 | 0.369 | 0.2428 | No | ||
| 9 | NRCAM | ENSG00000091129.20 | 9331 | 0.322 | 0.2240 | No | ||
| 10 | ADGRG1 | ENSG00000205336.12 | 9994 | 0.312 | 0.2317 | No | ||
| 11 | LDB1 | ENSG00000198728.10 | 12945 | 0.239 | 0.1782 | No | ||
| 12 | AMOT | ENSG00000126016.15 | 12946 | 0.239 | 0.1965 | No | ||
| 13 | L1CAM | ENSG00000198910.14 | 14319 | 0.197 | 0.1782 | No | ||
| 14 | PTCH1 | ENSG00000185920.15 | 14439 | 0.193 | 0.1900 | No | ||
| 15 | TLE1 | ENSG00000196781.15 | 14470 | 0.192 | 0.2039 | No | ||
| 16 | SHH | ENSG00000164690.8 | 14613 | 0.187 | 0.2148 | No | ||
| 17 | VEGFA | ENSG00000112715.22 | 15227 | 0.167 | 0.2126 | No | ||
| 18 | CNTFR | ENSG00000122756.15 | 16694 | 0.125 | 0.1866 | No | ||
| 19 | THY1 | ENSG00000154096.13 | 17556 | 0.102 | 0.1734 | No | ||
| 20 | SCG2 | ENSG00000171951.5 | 21211 | 0.006 | 0.0851 | No | ||
| 21 | HEY2 | ENSG00000135547.9 | 22349 | -0.022 | 0.0591 | No | ||
| 22 | ETS2 | ENSG00000157557.13 | 22722 | -0.031 | 0.0524 | No | ||
| 23 | CRMP1 | ENSG00000072832.15 | 22813 | -0.034 | 0.0528 | No | ||
| 24 | VLDLR | ENSG00000147852.16 | 23181 | -0.045 | 0.0473 | No | ||
| 25 | DPYSL2 | ENSG00000092964.18 | 23376 | -0.049 | 0.0463 | No | ||
| 26 | PML | ENSG00000140464.20 | 24111 | -0.067 | 0.0336 | No | ||
| 27 | HEY1 | ENSG00000164683.17 | 24120 | -0.067 | 0.0386 | No | ||
| 28 | NF1 | ENSG00000196712.17 | 29983 | -0.243 | -0.0854 | No | ||
| 29 | UNC5C | ENSG00000182168.15 | 30578 | -0.262 | -0.0798 | No | ||
| 30 | CDK5R1 | ENSG00000176749.9 | 32195 | -0.304 | -0.0959 | No | ||
| 31 | RASA1 | ENSG00000145715.15 | 34655 | -0.354 | -0.1286 | No | ||
| 32 | OPHN1 | ENSG00000079482.13 | 36417 | -0.422 | -0.1392 | No | ||
| 33 | MYH9 | ENSG00000100345.21 | 39531 | -0.641 | -0.1660 | No | ||
| 34 | NRP1 | ENSG00000099250.18 | 40541 | -0.854 | -0.1252 | No | ||
| 35 | CDK6 | ENSG00000105810.9 | 40555 | -0.858 | -0.0600 | No | ||
| 36 | NRP2 | ENSG00000118257.16 | 40814 | -0.978 | 0.0084 | No |