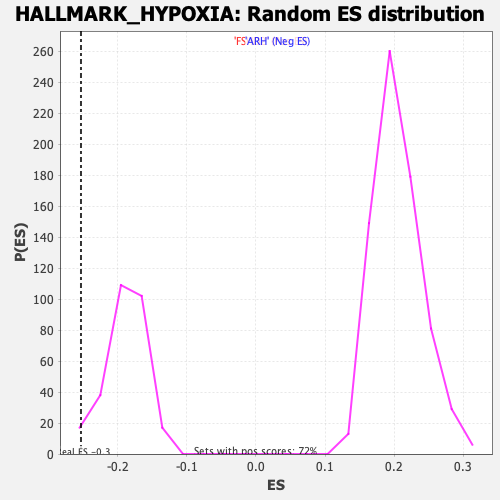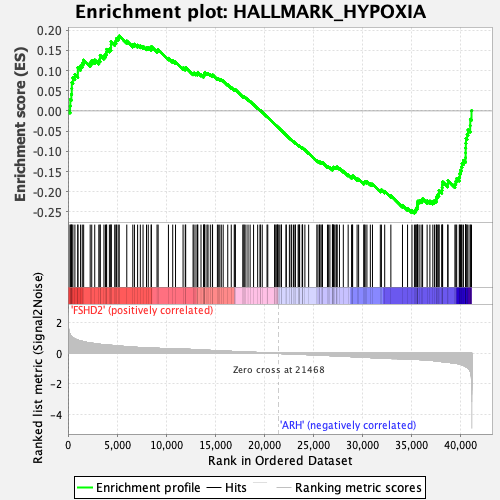
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_HYPOXIA |
| Enrichment Score (ES) | -0.2529327 |
| Normalized Enrichment Score (NES) | -1.3433015 |
| Nominal p-value | 0.017667845 |
| FDR q-value | 0.07220255 |
| FWER p-Value | 0.471 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | STC1 | ENSG00000159167.12 | 211 | 1.227 | 0.0122 | No | ||
| 2 | TGM2 | ENSG00000198959.12 | 237 | 1.183 | 0.0283 | No | ||
| 3 | TPD52 | ENSG00000076554.15 | 340 | 1.096 | 0.0412 | No | ||
| 4 | EFNA1 | ENSG00000169242.12 | 382 | 1.073 | 0.0554 | No | ||
| 5 | INHA | ENSG00000123999.5 | 391 | 1.069 | 0.0703 | No | ||
| 6 | CP | ENSG00000047457.14 | 500 | 1.015 | 0.0819 | No | ||
| 7 | RRAGD | ENSG00000025039.15 | 702 | 0.939 | 0.0903 | No | ||
| 8 | NEDD4L | ENSG00000049759.18 | 1003 | 0.858 | 0.0951 | No | ||
| 9 | GPC4 | ENSG00000076716.9 | 1004 | 0.858 | 0.1072 | No | ||
| 10 | GPC3 | ENSG00000147257.14 | 1281 | 0.805 | 0.1118 | No | ||
| 11 | IER3 | ENSG00000137331.12 | 1454 | 0.779 | 0.1186 | No | ||
| 12 | CDKN1C | ENSG00000129757.13 | 1596 | 0.759 | 0.1259 | No | ||
| 13 | DPYSL4 | ENSG00000151640.13 | 2261 | 0.676 | 0.1192 | No | ||
| 14 | IGFBP3 | ENSG00000146674.15 | 2420 | 0.661 | 0.1247 | No | ||
| 15 | SLC6A6 | ENSG00000131389.17 | 2725 | 0.630 | 0.1261 | No | ||
| 16 | MAP3K1 | ENSG00000095015.6 | 3142 | 0.596 | 0.1244 | No | ||
| 17 | PDGFB | ENSG00000100311.17 | 3269 | 0.586 | 0.1296 | No | ||
| 18 | TPBG | ENSG00000146242.8 | 3275 | 0.586 | 0.1377 | No | ||
| 19 | KIF5A | ENSG00000155980.12 | 3665 | 0.560 | 0.1361 | No | ||
| 20 | CXCR4 | ENSG00000121966.6 | 3819 | 0.548 | 0.1401 | No | ||
| 21 | BRS3 | ENSG00000102239.5 | 3905 | 0.545 | 0.1457 | No | ||
| 22 | MT2A | ENSG00000125148.7 | 3939 | 0.543 | 0.1526 | No | ||
| 23 | DDIT4 | ENSG00000168209.5 | 4218 | 0.529 | 0.1533 | No | ||
| 24 | CDKN1B | ENSG00000111276.11 | 4380 | 0.521 | 0.1567 | No | ||
| 25 | KLHL24 | ENSG00000114796.16 | 4382 | 0.521 | 0.1640 | No | ||
| 26 | BCL2 | ENSG00000171791.13 | 4386 | 0.520 | 0.1713 | No | ||
| 27 | NFIL3 | ENSG00000165030.4 | 4763 | 0.507 | 0.1692 | No | ||
| 28 | ANKZF1 | ENSG00000163516.14 | 4888 | 0.498 | 0.1732 | No | ||
| 29 | PDK3 | ENSG00000067992.15 | 4932 | 0.495 | 0.1792 | No | ||
| 30 | FBP1 | ENSG00000165140.10 | 5107 | 0.483 | 0.1817 | No | ||
| 31 | CCN5 | ENSG00000064205.10 | 5220 | 0.476 | 0.1857 | No | ||
| 32 | CCNG2 | ENSG00000138764.15 | 5991 | 0.434 | 0.1731 | No | ||
| 33 | PNRC1 | ENSG00000146278.11 | 6605 | 0.403 | 0.1638 | No | ||
| 34 | ISG20 | ENSG00000172183.15 | 6772 | 0.397 | 0.1653 | No | ||
| 35 | EFNA3 | ENSG00000143590.14 | 7095 | 0.385 | 0.1629 | No | ||
| 36 | ACKR3 | ENSG00000144476.6 | 7372 | 0.373 | 0.1614 | No | ||
| 37 | BTG1 | ENSG00000133639.5 | 7649 | 0.365 | 0.1599 | No | ||
| 38 | PFKFB3 | ENSG00000170525.21 | 8002 | 0.361 | 0.1564 | No | ||
| 39 | SAP30 | ENSG00000164105.4 | 8180 | 0.354 | 0.1570 | No | ||
| 40 | BGN | ENSG00000182492.16 | 8477 | 0.352 | 0.1548 | No | ||
| 41 | LDHC | ENSG00000166796.12 | 8497 | 0.351 | 0.1593 | No | ||
| 42 | HMOX1 | ENSG00000100292.17 | 9075 | 0.332 | 0.1499 | No | ||
| 43 | PAM | ENSG00000145730.20 | 9186 | 0.328 | 0.1518 | No | ||
| 44 | LALBA | ENSG00000167531.6 | 10237 | 0.305 | 0.1305 | No | ||
| 45 | RORA | ENSG00000069667.16 | 10674 | 0.304 | 0.1241 | No | ||
| 46 | PHKG1 | ENSG00000164776.9 | 10960 | 0.292 | 0.1213 | No | ||
| 47 | RBPJ | ENSG00000168214.20 | 11729 | 0.281 | 0.1065 | No | ||
| 48 | GAPDHS | ENSG00000105679.9 | 11950 | 0.273 | 0.1050 | No | ||
| 49 | PFKL | ENSG00000141959.17 | 11995 | 0.271 | 0.1077 | No | ||
| 50 | BCAN | ENSG00000132692.19 | 12750 | 0.246 | 0.0928 | No | ||
| 51 | ALDOC | ENSG00000109107.14 | 12846 | 0.243 | 0.0939 | No | ||
| 52 | MAFF | ENSG00000185022.12 | 13047 | 0.236 | 0.0923 | No | ||
| 53 | MIF | ENSG00000240972.2 | 13196 | 0.232 | 0.0920 | No | ||
| 54 | FAM162A | ENSG00000114023.15 | 13232 | 0.231 | 0.0944 | No | ||
| 55 | PPFIA4 | ENSG00000143847.15 | 13557 | 0.220 | 0.0896 | No | ||
| 56 | PPARGC1A | ENSG00000109819.9 | 13820 | 0.212 | 0.0862 | No | ||
| 57 | LARGE1 | ENSG00000133424.20 | 13853 | 0.211 | 0.0884 | No | ||
| 58 | FOSL2 | ENSG00000075426.12 | 13886 | 0.209 | 0.0906 | No | ||
| 59 | SLC37A4 | ENSG00000137700.18 | 13897 | 0.209 | 0.0933 | No | ||
| 60 | CAVIN3 | ENSG00000170955.10 | 13955 | 0.207 | 0.0948 | No | ||
| 61 | IRS2 | ENSG00000185950.9 | 14166 | 0.200 | 0.0925 | No | ||
| 62 | ALDOB | ENSG00000136872.20 | 14273 | 0.198 | 0.0927 | No | ||
| 63 | EDN2 | ENSG00000127129.10 | 14507 | 0.190 | 0.0897 | No | ||
| 64 | ILVBL | ENSG00000105135.16 | 14718 | 0.183 | 0.0872 | No | ||
| 65 | AK4 | ENSG00000162433.15 | 14736 | 0.183 | 0.0893 | No | ||
| 66 | VEGFA | ENSG00000112715.22 | 15227 | 0.167 | 0.0797 | No | ||
| 67 | STBD1 | ENSG00000118804.8 | 15318 | 0.164 | 0.0799 | No | ||
| 68 | SCARB1 | ENSG00000073060.16 | 15471 | 0.160 | 0.0784 | No | ||
| 69 | SELENBP1 | ENSG00000143416.21 | 15624 | 0.156 | 0.0769 | No | ||
| 70 | ZNF292 | ENSG00000188994.13 | 15818 | 0.150 | 0.0743 | No | ||
| 71 | PKLR | ENSG00000143627.19 | 16283 | 0.138 | 0.0649 | No | ||
| 72 | KDM3A | ENSG00000115548.17 | 16641 | 0.127 | 0.0580 | No | ||
| 73 | CSRP2 | ENSG00000175183.10 | 16944 | 0.118 | 0.0523 | No | ||
| 74 | FOXO3 | ENSG00000118689.15 | 17017 | 0.117 | 0.0522 | No | ||
| 75 | ZFP36 | ENSG00000128016.7 | 17058 | 0.116 | 0.0528 | No | ||
| 76 | SIAH2 | ENSG00000181788.4 | 17816 | 0.096 | 0.0357 | No | ||
| 77 | FOS | ENSG00000170345.10 | 17933 | 0.092 | 0.0342 | No | ||
| 78 | CA12 | ENSG00000074410.14 | 17955 | 0.092 | 0.0350 | No | ||
| 79 | ENO3 | ENSG00000108515.17 | 18133 | 0.087 | 0.0319 | No | ||
| 80 | PIM1 | ENSG00000137193.14 | 18349 | 0.082 | 0.0278 | No | ||
| 81 | VHL | ENSG00000134086.7 | 18565 | 0.077 | 0.0236 | No | ||
| 82 | GCNT2 | ENSG00000111846.17 | 18915 | 0.068 | 0.0161 | No | ||
| 83 | GAA | ENSG00000171298.13 | 19333 | 0.055 | 0.0067 | No | ||
| 84 | NDRG1 | ENSG00000104419.14 | 19578 | 0.049 | 0.0014 | No | ||
| 85 | STC2 | ENSG00000113739.10 | 19602 | 0.048 | 0.0015 | No | ||
| 86 | SLC2A1 | ENSG00000117394.23 | 19798 | 0.043 | -0.0026 | No | ||
| 87 | DCN | ENSG00000011465.17 | 20288 | 0.032 | -0.0141 | No | ||
| 88 | TGFB3 | ENSG00000119699.7 | 20364 | 0.029 | -0.0155 | No | ||
| 89 | GRHPR | ENSG00000137106.18 | 21060 | 0.010 | -0.0324 | No | ||
| 90 | TIPARP | ENSG00000163659.13 | 21091 | 0.009 | -0.0330 | No | ||
| 91 | TMEM45A | ENSG00000181458.10 | 21199 | 0.007 | -0.0355 | No | ||
| 92 | CDKN1A | ENSG00000124762.13 | 21297 | 0.004 | -0.0378 | No | ||
| 93 | NAGK | ENSG00000124357.13 | 21364 | 0.002 | -0.0394 | No | ||
| 94 | DDIT3 | ENSG00000175197.12 | 21390 | 0.001 | -0.0400 | No | ||
| 95 | HK2 | ENSG00000159399.9 | 21414 | 0.001 | -0.0405 | No | ||
| 96 | WSB1 | ENSG00000109046.15 | 21588 | -0.004 | -0.0447 | No | ||
| 97 | DTNA | ENSG00000134769.21 | 21748 | -0.008 | -0.0485 | No | ||
| 98 | SDC4 | ENSG00000124145.6 | 21749 | -0.008 | -0.0484 | No | ||
| 99 | PRKCA | ENSG00000154229.12 | 22219 | -0.019 | -0.0595 | No | ||
| 100 | CCN2 | ENSG00000118523.6 | 22253 | -0.019 | -0.0601 | No | ||
| 101 | COL5A1 | ENSG00000130635.15 | 22567 | -0.028 | -0.0673 | No | ||
| 102 | KDELR3 | ENSG00000100196.11 | 22740 | -0.032 | -0.0711 | No | ||
| 103 | MT1E | ENSG00000169715.15 | 22802 | -0.033 | -0.0721 | No | ||
| 104 | NDST2 | ENSG00000166507.18 | 22990 | -0.039 | -0.0761 | No | ||
| 105 | PLAC8 | ENSG00000145287.10 | 23135 | -0.043 | -0.0790 | No | ||
| 106 | VLDLR | ENSG00000147852.16 | 23181 | -0.045 | -0.0795 | No | ||
| 107 | PKP1 | ENSG00000081277.12 | 23458 | -0.051 | -0.0855 | No | ||
| 108 | HEXA | ENSG00000213614.9 | 23544 | -0.054 | -0.0868 | No | ||
| 109 | GLRX | ENSG00000173221.14 | 23563 | -0.054 | -0.0865 | No | ||
| 110 | GPI | ENSG00000105220.17 | 23611 | -0.056 | -0.0869 | No | ||
| 111 | GALK1 | ENSG00000108479.12 | 23871 | -0.062 | -0.0923 | No | ||
| 112 | GBE1 | ENSG00000114480.13 | 23894 | -0.063 | -0.0920 | No | ||
| 113 | SLC25A1 | ENSG00000100075.10 | 23913 | -0.064 | -0.0915 | No | ||
| 114 | EXT1 | ENSG00000182197.11 | 24146 | -0.068 | -0.0962 | No | ||
| 115 | ALDOA | ENSG00000149925.20 | 24530 | -0.078 | -0.1044 | No | ||
| 116 | GCK | ENSG00000106633.17 | 25365 | -0.101 | -0.1234 | No | ||
| 117 | BHLHE40 | ENSG00000134107.5 | 25483 | -0.105 | -0.1248 | No | ||
| 118 | LXN | ENSG00000079257.8 | 25642 | -0.110 | -0.1271 | No | ||
| 119 | HAS1 | ENSG00000105509.10 | 25668 | -0.110 | -0.1261 | No | ||
| 120 | PYGM | ENSG00000068976.14 | 25795 | -0.113 | -0.1276 | No | ||
| 121 | SULT2B1 | ENSG00000088002.11 | 25907 | -0.117 | -0.1287 | No | ||
| 122 | HS3ST1 | ENSG00000002587.10 | 25942 | -0.118 | -0.1278 | No | ||
| 123 | GAPDH | ENSG00000111640.15 | 26448 | -0.134 | -0.1383 | No | ||
| 124 | GPC1 | ENSG00000063660.9 | 26538 | -0.136 | -0.1385 | No | ||
| 125 | IDS | ENSG00000010404.18 | 26695 | -0.141 | -0.1403 | No | ||
| 126 | TPI1 | ENSG00000111669.15 | 26955 | -0.148 | -0.1446 | No | ||
| 127 | B3GALT6 | ENSG00000176022.7 | 26956 | -0.148 | -0.1425 | No | ||
| 128 | PPP1R15A | ENSG00000087074.8 | 27000 | -0.149 | -0.1414 | No | ||
| 129 | TNFAIP3 | ENSG00000118503.15 | 27043 | -0.151 | -0.1403 | No | ||
| 130 | ATF3 | ENSG00000162772.17 | 27076 | -0.151 | -0.1390 | No | ||
| 131 | BNIP3L | ENSG00000104765.16 | 27191 | -0.155 | -0.1396 | No | ||
| 132 | CASP6 | ENSG00000138794.10 | 27327 | -0.159 | -0.1406 | No | ||
| 133 | NCAN | ENSG00000130287.14 | 27413 | -0.161 | -0.1404 | No | ||
| 134 | HDLBP | ENSG00000115677.17 | 27421 | -0.162 | -0.1383 | No | ||
| 135 | CHST2 | ENSG00000175040.6 | 27669 | -0.170 | -0.1419 | No | ||
| 136 | MXI1 | ENSG00000119950.21 | 28064 | -0.183 | -0.1490 | No | ||
| 137 | IGFBP1 | ENSG00000146678.10 | 28557 | -0.198 | -0.1582 | No | ||
| 138 | ANGPTL4 | ENSG00000167772.12 | 28905 | -0.208 | -0.1637 | No | ||
| 139 | ATP7A | ENSG00000165240.20 | 28987 | -0.212 | -0.1627 | No | ||
| 140 | SLC2A5 | ENSG00000142583.18 | 29013 | -0.212 | -0.1603 | No | ||
| 141 | PPP1R3C | ENSG00000119938.9 | 29469 | -0.226 | -0.1683 | No | ||
| 142 | IL6 | ENSG00000136244.12 | 29624 | -0.231 | -0.1687 | No | ||
| 143 | PRDX5 | ENSG00000126432.14 | 30144 | -0.249 | -0.1779 | No | ||
| 144 | PLIN2 | ENSG00000147872.10 | 30207 | -0.250 | -0.1759 | No | ||
| 145 | CCN1 | ENSG00000142871.17 | 30290 | -0.253 | -0.1743 | No | ||
| 146 | PGM1 | ENSG00000079739.17 | 30468 | -0.259 | -0.1750 | No | ||
| 147 | XPNPEP1 | ENSG00000108039.18 | 30817 | -0.270 | -0.1797 | No | ||
| 148 | SRPX | ENSG00000101955.15 | 31035 | -0.278 | -0.1810 | No | ||
| 149 | ETS1 | ENSG00000134954.14 | 31835 | -0.291 | -0.1964 | No | ||
| 150 | ERO1A | ENSG00000197930.13 | 31946 | -0.294 | -0.1950 | No | ||
| 151 | HSPA5 | ENSG00000044574.8 | 32282 | -0.308 | -0.1988 | No | ||
| 152 | EGFR | ENSG00000146648.18 | 32910 | -0.332 | -0.2094 | No | ||
| 153 | PGF | ENSG00000119630.14 | 34094 | -0.345 | -0.2334 | No | ||
| 154 | JMJD6 | ENSG00000070495.14 | 34612 | -0.352 | -0.2411 | No | ||
| 155 | B4GALNT2 | ENSG00000167080.8 | 35061 | -0.368 | -0.2468 | No | ||
| 156 | PLAUR | ENSG00000011422.12 | 35312 | -0.379 | -0.2476 | Yes | ||
| 157 | SDC3 | ENSG00000162512.16 | 35439 | -0.385 | -0.2452 | Yes | ||
| 158 | PFKP | ENSG00000067057.17 | 35494 | -0.388 | -0.2411 | Yes | ||
| 159 | PDK1 | ENSG00000152256.13 | 35589 | -0.393 | -0.2378 | Yes | ||
| 160 | P4HA2 | ENSG00000072682.18 | 35623 | -0.395 | -0.2331 | Yes | ||
| 161 | ENO1 | ENSG00000074800.16 | 35638 | -0.395 | -0.2278 | Yes | ||
| 162 | PGM2 | ENSG00000169299.14 | 35669 | -0.396 | -0.2230 | Yes | ||
| 163 | PCK1 | ENSG00000124253.11 | 35868 | -0.402 | -0.2221 | Yes | ||
| 164 | ANXA2 | ENSG00000182718.16 | 36074 | -0.407 | -0.2214 | Yes | ||
| 165 | TKTL1 | ENSG00000007350.17 | 36144 | -0.411 | -0.2173 | Yes | ||
| 166 | ENO2 | ENSG00000111674.9 | 36606 | -0.431 | -0.2224 | Yes | ||
| 167 | CHST3 | ENSG00000122863.6 | 36888 | -0.445 | -0.2230 | Yes | ||
| 168 | CITED2 | ENSG00000164442.10 | 37189 | -0.463 | -0.2238 | Yes | ||
| 169 | P4HA1 | ENSG00000122884.12 | 37364 | -0.475 | -0.2213 | Yes | ||
| 170 | SERPINE1 | ENSG00000106366.8 | 37533 | -0.487 | -0.2186 | Yes | ||
| 171 | ERRFI1 | ENSG00000116285.13 | 37565 | -0.490 | -0.2124 | Yes | ||
| 172 | AKAP12 | ENSG00000131016.17 | 37685 | -0.501 | -0.2082 | Yes | ||
| 173 | PGK1 | ENSG00000102144.15 | 37804 | -0.512 | -0.2039 | Yes | ||
| 174 | TGFBI | ENSG00000120708.17 | 37818 | -0.514 | -0.1970 | Yes | ||
| 175 | GYS1 | ENSG00000104812.15 | 38103 | -0.535 | -0.1963 | Yes | ||
| 176 | TPST2 | ENSG00000128294.16 | 38137 | -0.537 | -0.1896 | Yes | ||
| 177 | AMPD3 | ENSG00000133805.15 | 38154 | -0.539 | -0.1824 | Yes | ||
| 178 | SDC2 | ENSG00000169439.12 | 38187 | -0.541 | -0.1755 | Yes | ||
| 179 | NOCT | ENSG00000151014.6 | 38706 | -0.571 | -0.1801 | Yes | ||
| 180 | KLF7 | ENSG00000118263.15 | 38734 | -0.574 | -0.1726 | Yes | ||
| 181 | HK1 | ENSG00000156515.23 | 39440 | -0.630 | -0.1810 | Yes | ||
| 182 | MYH9 | ENSG00000100345.21 | 39531 | -0.641 | -0.1741 | Yes | ||
| 183 | DUSP1 | ENSG00000120129.6 | 39618 | -0.648 | -0.1671 | Yes | ||
| 184 | ADORA2B | ENSG00000170425.3 | 39882 | -0.692 | -0.1637 | Yes | ||
| 185 | HOXB9 | ENSG00000170689.10 | 39931 | -0.700 | -0.1550 | Yes | ||
| 186 | F3 | ENSG00000117525.14 | 40005 | -0.714 | -0.1467 | Yes | ||
| 187 | CAVIN1 | ENSG00000177469.13 | 40101 | -0.731 | -0.1387 | Yes | ||
| 188 | KLF6 | ENSG00000067082.15 | 40162 | -0.745 | -0.1297 | Yes | ||
| 189 | JUN | ENSG00000177606.7 | 40305 | -0.779 | -0.1222 | Yes | ||
| 190 | ADM | ENSG00000148926.10 | 40523 | -0.847 | -0.1155 | Yes | ||
| 191 | S100A4 | ENSG00000196154.12 | 40530 | -0.850 | -0.1037 | Yes | ||
| 192 | NDST1 | ENSG00000070614.15 | 40535 | -0.851 | -0.0918 | Yes | ||
| 193 | NR3C1 | ENSG00000113580.15 | 40550 | -0.857 | -0.0800 | Yes | ||
| 194 | SLC2A3 | ENSG00000059804.16 | 40569 | -0.866 | -0.0682 | Yes | ||
| 195 | LDHA | ENSG00000134333.14 | 40682 | -0.914 | -0.0581 | Yes | ||
| 196 | UGP2 | ENSG00000169764.15 | 40772 | -0.945 | -0.0469 | Yes | ||
| 197 | TES | ENSG00000135269.18 | 40994 | -1.130 | -0.0364 | Yes | ||
| 198 | CAV1 | ENSG00000105974.12 | 41003 | -1.149 | -0.0204 | Yes | ||
| 199 | LOX | ENSG00000113083.14 | 41130 | -1.714 | 0.0008 | Yes |