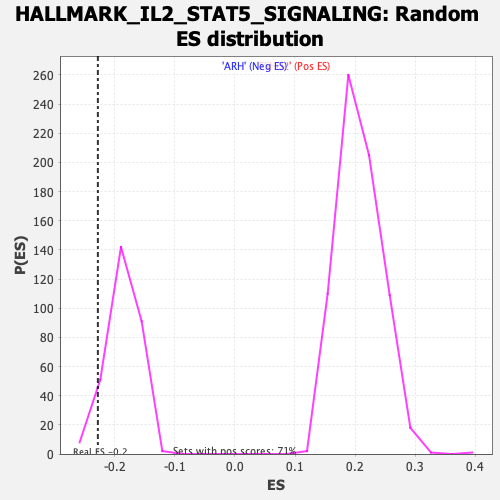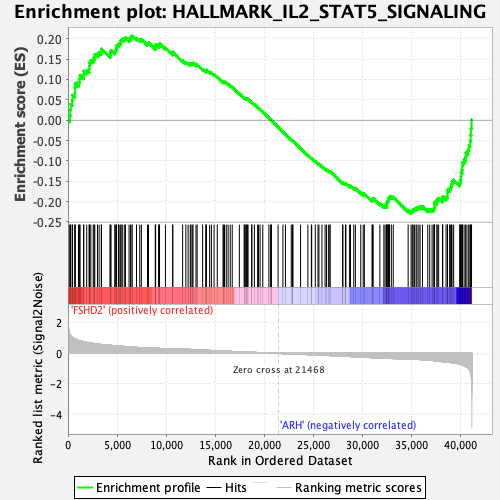
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_IL2_STAT5_SIGNALING |
| Enrichment Score (ES) | -0.22841942 |
| Normalized Enrichment Score (NES) | -1.2217444 |
| Nominal p-value | 0.07482993 |
| FDR q-value | 0.14262362 |
| FWER p-Value | 0.811 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IL13 | ENSG00000169194.9 | 180 | 1.275 | 0.0119 | No | ||
| 2 | TGM2 | ENSG00000198959.12 | 237 | 1.183 | 0.0257 | No | ||
| 3 | IL1R2 | ENSG00000115590.14 | 280 | 1.147 | 0.0393 | No | ||
| 4 | WLS | ENSG00000116729.14 | 435 | 1.042 | 0.0488 | No | ||
| 5 | S100A1 | ENSG00000160678.11 | 463 | 1.030 | 0.0614 | No | ||
| 6 | RRAGD | ENSG00000025039.15 | 702 | 0.939 | 0.0675 | No | ||
| 7 | BMP2 | ENSG00000125845.7 | 710 | 0.936 | 0.0793 | No | ||
| 8 | CISH | ENSG00000114737.15 | 726 | 0.933 | 0.0909 | No | ||
| 9 | SLC39A8 | ENSG00000138821.13 | 1070 | 0.844 | 0.0933 | No | ||
| 10 | PENK | ENSG00000181195.11 | 1184 | 0.821 | 0.1010 | No | ||
| 11 | IRF4 | ENSG00000137265.15 | 1229 | 0.811 | 0.1103 | No | ||
| 12 | CDKN1C | ENSG00000129757.13 | 1596 | 0.759 | 0.1111 | No | ||
| 13 | LIF | ENSG00000128342.5 | 1603 | 0.758 | 0.1207 | No | ||
| 14 | AHR | ENSG00000106546.14 | 1908 | 0.715 | 0.1224 | No | ||
| 15 | PRKCH | ENSG00000027075.15 | 2159 | 0.687 | 0.1251 | No | ||
| 16 | SMPDL3A | ENSG00000172594.13 | 2174 | 0.686 | 0.1335 | No | ||
| 17 | IL2RB | ENSG00000100385.14 | 2208 | 0.683 | 0.1414 | No | ||
| 18 | IFNGR1 | ENSG00000027697.14 | 2329 | 0.669 | 0.1470 | No | ||
| 19 | PTH1R | ENSG00000160801.14 | 2572 | 0.646 | 0.1494 | No | ||
| 20 | MUC1 | ENSG00000185499.16 | 2664 | 0.637 | 0.1553 | No | ||
| 21 | PHLDA1 | ENSG00000139289.13 | 2729 | 0.630 | 0.1618 | No | ||
| 22 | BATF3 | ENSG00000123685.9 | 3004 | 0.605 | 0.1628 | No | ||
| 23 | RGS16 | ENSG00000143333.7 | 3170 | 0.593 | 0.1664 | No | ||
| 24 | LTB | ENSG00000227507.2 | 3397 | 0.576 | 0.1682 | No | ||
| 25 | CDCP1 | ENSG00000163814.8 | 3408 | 0.574 | 0.1753 | No | ||
| 26 | CST7 | ENSG00000077984.6 | 4293 | 0.524 | 0.1604 | No | ||
| 27 | HOPX | ENSG00000171476.22 | 4310 | 0.523 | 0.1667 | No | ||
| 28 | BCL2 | ENSG00000171791.13 | 4386 | 0.520 | 0.1715 | No | ||
| 29 | NFIL3 | ENSG00000165030.4 | 4763 | 0.507 | 0.1688 | No | ||
| 30 | CCND2 | ENSG00000118971.8 | 4862 | 0.500 | 0.1728 | No | ||
| 31 | CD44 | ENSG00000026508.18 | 4934 | 0.495 | 0.1774 | No | ||
| 32 | NOP2 | ENSG00000111641.11 | 4949 | 0.494 | 0.1834 | No | ||
| 33 | SOCS2 | ENSG00000120833.14 | 5140 | 0.481 | 0.1849 | No | ||
| 34 | AGER | ENSG00000204305.14 | 5212 | 0.477 | 0.1893 | No | ||
| 35 | SLC29A2 | ENSG00000174669.12 | 5384 | 0.466 | 0.1911 | No | ||
| 36 | SCN9A | ENSG00000169432.16 | 5398 | 0.465 | 0.1967 | No | ||
| 37 | TNFRSF18 | ENSG00000186891.14 | 5531 | 0.458 | 0.1993 | No | ||
| 38 | F2RL2 | ENSG00000164220.7 | 5734 | 0.446 | 0.2001 | No | ||
| 39 | PLAGL1 | ENSG00000118495.19 | 5858 | 0.441 | 0.2027 | No | ||
| 40 | COCH | ENSG00000100473.18 | 6222 | 0.421 | 0.1993 | No | ||
| 41 | TNFSF10 | ENSG00000121858.11 | 6380 | 0.413 | 0.2007 | No | ||
| 42 | FAH | ENSG00000103876.12 | 6411 | 0.412 | 0.2052 | No | ||
| 43 | SNX9 | ENSG00000130340.16 | 6548 | 0.406 | 0.2071 | No | ||
| 44 | PTGER2 | ENSG00000125384.7 | 6983 | 0.390 | 0.2015 | No | ||
| 45 | IL18R1 | ENSG00000115604.10 | 7316 | 0.376 | 0.1982 | No | ||
| 46 | ENPP1 | ENSG00000197594.13 | 7476 | 0.372 | 0.1991 | No | ||
| 47 | GLIPR2 | ENSG00000122694.16 | 8105 | 0.357 | 0.1883 | No | ||
| 48 | POU2F1 | ENSG00000143190.23 | 8203 | 0.353 | 0.1905 | No | ||
| 49 | IL4R | ENSG00000077238.14 | 8901 | 0.339 | 0.1778 | No | ||
| 50 | SERPINC1 | ENSG00000117601.13 | 8910 | 0.339 | 0.1819 | No | ||
| 51 | PRAF2 | ENSG00000243279.4 | 8931 | 0.338 | 0.1858 | No | ||
| 52 | APLP1 | ENSG00000105290.12 | 9232 | 0.326 | 0.1826 | No | ||
| 53 | IGF1R | ENSG00000140443.15 | 9322 | 0.322 | 0.1845 | No | ||
| 54 | RHOH | ENSG00000168421.13 | 9330 | 0.322 | 0.1885 | No | ||
| 55 | CAPN3 | ENSG00000092529.24 | 9932 | 0.314 | 0.1778 | No | ||
| 56 | RORA | ENSG00000069667.16 | 10674 | 0.304 | 0.1636 | No | ||
| 57 | MXD1 | ENSG00000059728.11 | 10676 | 0.304 | 0.1675 | No | ||
| 58 | ETV4 | ENSG00000175832.13 | 11684 | 0.283 | 0.1465 | No | ||
| 59 | PLSCR1 | ENSG00000188313.13 | 12014 | 0.270 | 0.1419 | No | ||
| 60 | SH3BGRL2 | ENSG00000198478.8 | 12246 | 0.263 | 0.1397 | No | ||
| 61 | GABARAPL1 | ENSG00000139112.11 | 12462 | 0.255 | 0.1377 | No | ||
| 62 | CA2 | ENSG00000104267.10 | 12533 | 0.253 | 0.1392 | No | ||
| 63 | TNFRSF21 | ENSG00000146072.6 | 12662 | 0.249 | 0.1393 | No | ||
| 64 | CD81 | ENSG00000110651.11 | 12744 | 0.247 | 0.1404 | No | ||
| 65 | MAFF | ENSG00000185022.12 | 13047 | 0.236 | 0.1361 | No | ||
| 66 | P2RX4 | ENSG00000135124.14 | 13164 | 0.233 | 0.1362 | No | ||
| 67 | GATA1 | ENSG00000102145.14 | 13724 | 0.215 | 0.1253 | No | ||
| 68 | GPR65 | ENSG00000140030.6 | 14045 | 0.204 | 0.1201 | No | ||
| 69 | DRC1 | ENSG00000157856.12 | 14091 | 0.203 | 0.1216 | No | ||
| 70 | ABCB1 | ENSG00000085563.14 | 14109 | 0.202 | 0.1238 | No | ||
| 71 | PTCH1 | ENSG00000185920.15 | 14439 | 0.193 | 0.1182 | No | ||
| 72 | IGF2R | ENSG00000197081.14 | 14615 | 0.187 | 0.1163 | No | ||
| 73 | CCR4 | ENSG00000183813.7 | 14891 | 0.177 | 0.1119 | No | ||
| 74 | GPX4 | ENSG00000167468.17 | 15215 | 0.168 | 0.1062 | No | ||
| 75 | UMPS | ENSG00000114491.14 | 15817 | 0.150 | 0.0934 | No | ||
| 76 | CD83 | ENSG00000112149.10 | 15825 | 0.150 | 0.0951 | No | ||
| 77 | IL2RA | ENSG00000134460.17 | 15953 | 0.146 | 0.0939 | No | ||
| 78 | TRAF1 | ENSG00000056558.11 | 16131 | 0.142 | 0.0914 | No | ||
| 79 | TTC39B | ENSG00000155158.20 | 16319 | 0.137 | 0.0886 | No | ||
| 80 | RHOB | ENSG00000143878.10 | 16535 | 0.130 | 0.0850 | No | ||
| 81 | SYNGR2 | ENSG00000108639.7 | 16740 | 0.124 | 0.0816 | No | ||
| 82 | IFITM3 | ENSG00000142089.16 | 17463 | 0.105 | 0.0653 | No | ||
| 83 | CSF1 | ENSG00000184371.14 | 17952 | 0.092 | 0.0546 | No | ||
| 84 | PDCD2L | ENSG00000126249.8 | 18009 | 0.090 | 0.0544 | No | ||
| 85 | ENO3 | ENSG00000108515.17 | 18133 | 0.087 | 0.0525 | No | ||
| 86 | PUS1 | ENSG00000177192.14 | 18161 | 0.087 | 0.0529 | No | ||
| 87 | COL6A1 | ENSG00000142156.14 | 18165 | 0.087 | 0.0540 | No | ||
| 88 | IKZF4 | ENSG00000123411.15 | 18307 | 0.084 | 0.0516 | No | ||
| 89 | PIM1 | ENSG00000137193.14 | 18349 | 0.082 | 0.0516 | No | ||
| 90 | IKZF2 | ENSG00000030419.16 | 18729 | 0.073 | 0.0433 | No | ||
| 91 | ST3GAL4 | ENSG00000110080.18 | 18736 | 0.072 | 0.0441 | No | ||
| 92 | EOMES | ENSG00000163508.13 | 18998 | 0.065 | 0.0386 | No | ||
| 93 | MYO1E | ENSG00000157483.9 | 19330 | 0.055 | 0.0312 | No | ||
| 94 | NFKBIZ | ENSG00000144802.11 | 19416 | 0.053 | 0.0298 | No | ||
| 95 | NDRG1 | ENSG00000104419.14 | 19578 | 0.049 | 0.0265 | No | ||
| 96 | MAP3K8 | ENSG00000107968.10 | 19857 | 0.042 | 0.0202 | No | ||
| 97 | TIAM1 | ENSG00000156299.13 | 20475 | 0.025 | 0.0055 | No | ||
| 98 | SPRY4 | ENSG00000187678.9 | 20655 | 0.021 | 0.0014 | No | ||
| 99 | MAPKAPK2 | ENSG00000162889.10 | 20713 | 0.019 | 0.0003 | No | ||
| 100 | ETFBKMT | ENSG00000139160.13 | 20732 | 0.019 | 0.0001 | No | ||
| 101 | HK2 | ENSG00000159399.9 | 21414 | 0.001 | -0.0166 | No | ||
| 102 | AMACR | ENSG00000242110.8 | 21907 | -0.012 | -0.0284 | No | ||
| 103 | CD79B | ENSG00000007312.12 | 22164 | -0.017 | -0.0344 | No | ||
| 104 | IRF6 | ENSG00000117595.12 | 22754 | -0.032 | -0.0484 | No | ||
| 105 | GUCY1B1 | ENSG00000061918.13 | 22867 | -0.035 | -0.0507 | No | ||
| 106 | XBP1 | ENSG00000100219.16 | 22900 | -0.036 | -0.0510 | No | ||
| 107 | ALCAM | ENSG00000170017.12 | 22924 | -0.037 | -0.0511 | No | ||
| 108 | FAM126B | ENSG00000155744.9 | 23709 | -0.058 | -0.0695 | No | ||
| 109 | NT5E | ENSG00000135318.12 | 24453 | -0.076 | -0.0867 | No | ||
| 110 | DHRS3 | ENSG00000162496.9 | 24804 | -0.086 | -0.0941 | No | ||
| 111 | SELL | ENSG00000188404.10 | 24840 | -0.087 | -0.0939 | No | ||
| 112 | ITGAE | ENSG00000083457.12 | 25206 | -0.096 | -0.1015 | No | ||
| 113 | BHLHE40 | ENSG00000134107.5 | 25483 | -0.105 | -0.1069 | No | ||
| 114 | GPR83 | ENSG00000123901.9 | 25578 | -0.107 | -0.1079 | No | ||
| 115 | TLR7 | ENSG00000196664.5 | 25886 | -0.117 | -0.1139 | No | ||
| 116 | SYT11 | ENSG00000132718.9 | 26204 | -0.126 | -0.1200 | No | ||
| 117 | TWSG1 | ENSG00000128791.12 | 26343 | -0.130 | -0.1217 | No | ||
| 118 | FURIN | ENSG00000140564.12 | 26542 | -0.136 | -0.1248 | No | ||
| 119 | SOCS1 | ENSG00000185338.5 | 26705 | -0.141 | -0.1269 | No | ||
| 120 | SERPINB6 | ENSG00000124570.20 | 26707 | -0.141 | -0.1252 | No | ||
| 121 | HUWE1 | ENSG00000086758.16 | 27995 | -0.180 | -0.1543 | No | ||
| 122 | SLC1A5 | ENSG00000105281.12 | 28017 | -0.181 | -0.1525 | No | ||
| 123 | TNFRSF1B | ENSG00000028137.19 | 28280 | -0.189 | -0.1565 | No | ||
| 124 | DCPS | ENSG00000110063.10 | 28326 | -0.190 | -0.1551 | No | ||
| 125 | PRNP | ENSG00000171867.17 | 28702 | -0.202 | -0.1617 | No | ||
| 126 | EEF1AKMT1 | ENSG00000150456.11 | 28797 | -0.205 | -0.1614 | No | ||
| 127 | SNX14 | ENSG00000135317.13 | 29130 | -0.216 | -0.1667 | No | ||
| 128 | RNH1 | ENSG00000023191.17 | 29292 | -0.221 | -0.1678 | No | ||
| 129 | ITIH5 | ENSG00000123243.15 | 29842 | -0.239 | -0.1782 | No | ||
| 130 | FLT3LG | ENSG00000090554.13 | 30091 | -0.247 | -0.1811 | No | ||
| 131 | PLIN2 | ENSG00000147872.10 | 30207 | -0.250 | -0.1807 | No | ||
| 132 | CKAP4 | ENSG00000136026.14 | 30989 | -0.277 | -0.1962 | No | ||
| 133 | PTRH2 | ENSG00000141378.14 | 31046 | -0.279 | -0.1940 | No | ||
| 134 | CTSZ | ENSG00000101160.14 | 31105 | -0.281 | -0.1918 | No | ||
| 135 | CYFIP1 | ENSG00000273749.5 | 31788 | -0.289 | -0.2048 | No | ||
| 136 | CCNE1 | ENSG00000105173.14 | 32197 | -0.305 | -0.2108 | No | ||
| 137 | TNFRSF4 | ENSG00000186827.11 | 32325 | -0.310 | -0.2100 | No | ||
| 138 | AHNAK | ENSG00000124942.14 | 32497 | -0.317 | -0.2101 | No | ||
| 139 | SHE | ENSG00000169291.10 | 32504 | -0.317 | -0.2062 | No | ||
| 140 | MYO1C | ENSG00000197879.17 | 32509 | -0.317 | -0.2023 | No | ||
| 141 | CCND3 | ENSG00000112576.12 | 32532 | -0.317 | -0.1987 | No | ||
| 142 | FGL2 | ENSG00000127951.7 | 32655 | -0.323 | -0.1976 | No | ||
| 143 | CTLA4 | ENSG00000163599.17 | 32687 | -0.324 | -0.1942 | No | ||
| 144 | CXCL10 | ENSG00000169245.6 | 32700 | -0.324 | -0.1904 | No | ||
| 145 | CAPG | ENSG00000042493.16 | 32769 | -0.326 | -0.1879 | No | ||
| 146 | ODC1 | ENSG00000115758.13 | 32958 | -0.334 | -0.1882 | No | ||
| 147 | TNFSF11 | ENSG00000120659.15 | 33152 | -0.336 | -0.1886 | No | ||
| 148 | SPRED2 | ENSG00000198369.10 | 34671 | -0.355 | -0.2211 | No | ||
| 149 | GALM | ENSG00000143891.17 | 34972 | -0.364 | -0.2238 | Yes | ||
| 150 | ADAM19 | ENSG00000135074.15 | 35096 | -0.368 | -0.2221 | Yes | ||
| 151 | AHCY | ENSG00000101444.13 | 35184 | -0.373 | -0.2194 | Yes | ||
| 152 | CASP3 | ENSG00000164305.19 | 35329 | -0.380 | -0.2181 | Yes | ||
| 153 | IL1RL1 | ENSG00000115602.17 | 35484 | -0.388 | -0.2169 | Yes | ||
| 154 | RABGAP1L | ENSG00000152061.23 | 35578 | -0.393 | -0.2141 | Yes | ||
| 155 | GSTO1 | ENSG00000148834.13 | 35765 | -0.400 | -0.2136 | Yes | ||
| 156 | CD86 | ENSG00000114013.16 | 35887 | -0.402 | -0.2114 | Yes | ||
| 157 | PNP | ENSG00000198805.11 | 36135 | -0.411 | -0.2121 | Yes | ||
| 158 | LCLAT1 | ENSG00000172954.13 | 36670 | -0.434 | -0.2196 | Yes | ||
| 159 | EMP1 | ENSG00000134531.10 | 36872 | -0.444 | -0.2189 | Yes | ||
| 160 | GBP4 | ENSG00000162654.9 | 37113 | -0.458 | -0.2189 | Yes | ||
| 161 | LRIG1 | ENSG00000144749.13 | 37286 | -0.470 | -0.2171 | Yes | ||
| 162 | PLPP1 | ENSG00000067113.17 | 37340 | -0.474 | -0.2123 | Yes | ||
| 163 | BMPR2 | ENSG00000204217.15 | 37352 | -0.475 | -0.2065 | Yes | ||
| 164 | P4HA1 | ENSG00000122884.12 | 37364 | -0.475 | -0.2007 | Yes | ||
| 165 | HIPK2 | ENSG00000064393.16 | 37609 | -0.494 | -0.2003 | Yes | ||
| 166 | PLEC | ENSG00000178209.15 | 37624 | -0.496 | -0.1943 | Yes | ||
| 167 | ITGAV | ENSG00000138448.12 | 37784 | -0.511 | -0.1917 | Yes | ||
| 168 | BATF | ENSG00000156127.7 | 38174 | -0.540 | -0.1943 | Yes | ||
| 169 | CSF2 | ENSG00000164400.6 | 38203 | -0.543 | -0.1880 | Yes | ||
| 170 | TNFRSF8 | ENSG00000120949.15 | 38537 | -0.560 | -0.1890 | Yes | ||
| 171 | BCL2L1 | ENSG00000171552.13 | 38679 | -0.568 | -0.1852 | Yes | ||
| 172 | ANXA4 | ENSG00000196975.16 | 38680 | -0.569 | -0.1779 | Yes | ||
| 173 | SPP1 | ENSG00000118785.14 | 38686 | -0.569 | -0.1707 | Yes | ||
| 174 | LRRC8C | ENSG00000171488.15 | 38900 | -0.590 | -0.1684 | Yes | ||
| 175 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.1632 | Yes | ||
| 176 | DENND5A | ENSG00000184014.8 | 39087 | -0.611 | -0.1574 | Yes | ||
| 177 | SELP | ENSG00000174175.17 | 39143 | -0.614 | -0.1509 | Yes | ||
| 178 | CDC42SE2 | ENSG00000158985.13 | 39311 | -0.624 | -0.1470 | Yes | ||
| 179 | ECM1 | ENSG00000143369.15 | 39921 | -0.699 | -0.1530 | Yes | ||
| 180 | GADD45B | ENSG00000099860.9 | 40019 | -0.716 | -0.1462 | Yes | ||
| 181 | ARL4A | ENSG00000122644.13 | 40051 | -0.722 | -0.1377 | Yes | ||
| 182 | CDC6 | ENSG00000094804.12 | 40060 | -0.723 | -0.1287 | Yes | ||
| 183 | KLF6 | ENSG00000067082.15 | 40162 | -0.745 | -0.1216 | Yes | ||
| 184 | NCS1 | ENSG00000107130.10 | 40188 | -0.750 | -0.1126 | Yes | ||
| 185 | MAP6 | ENSG00000171533.11 | 40217 | -0.756 | -0.1036 | Yes | ||
| 186 | PHTF2 | ENSG00000006576.16 | 40375 | -0.798 | -0.0973 | Yes | ||
| 187 | NRP1 | ENSG00000099250.18 | 40541 | -0.854 | -0.0904 | Yes | ||
| 188 | SLC2A3 | ENSG00000059804.16 | 40569 | -0.866 | -0.0800 | Yes | ||
| 189 | IL10RA | ENSG00000110324.10 | 40784 | -0.953 | -0.0730 | Yes | ||
| 190 | UCK2 | ENSG00000143179.16 | 40869 | -1.012 | -0.0621 | Yes | ||
| 191 | SWAP70 | ENSG00000133789.15 | 40989 | -1.124 | -0.0507 | Yes | ||
| 192 | NCOA3 | ENSG00000124151.19 | 41048 | -1.221 | -0.0365 | Yes | ||
| 193 | ITGA6 | ENSG00000091409.15 | 41076 | -1.311 | -0.0204 | Yes | ||
| 194 | TNFRSF9 | ENSG00000049249.8 | 41133 | -1.755 | 0.0007 | Yes |