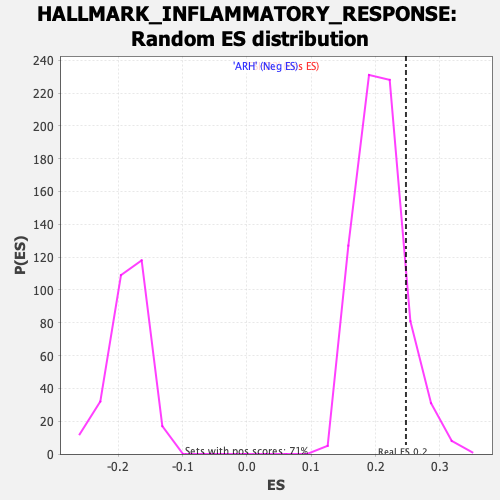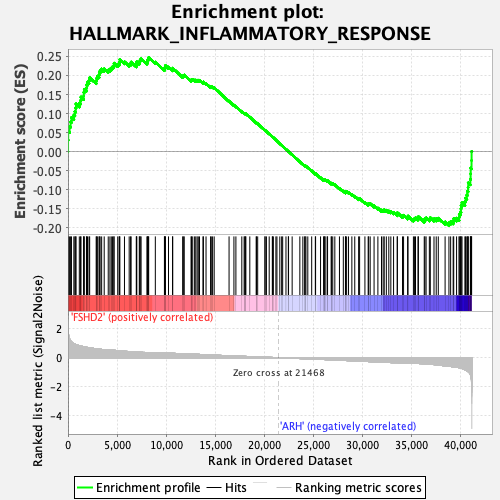
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_INFLAMMATORY_RESPONSE |
| Enrichment Score (ES) | 0.24734537 |
| Normalized Enrichment Score (NES) | 1.189314 |
| Nominal p-value | 0.12219101 |
| FDR q-value | 0.20496924 |
| FWER p-Value | 0.997 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CCRL2 | ENSG00000121797.10 | 4 | 2.444 | 0.0305 | Yes | ||
| 2 | NLRP3 | ENSG00000162711.17 | 20 | 1.857 | 0.0533 | Yes | ||
| 3 | HPN | ENSG00000105707.14 | 162 | 1.301 | 0.0662 | Yes | ||
| 4 | P2RX7 | ENSG00000089041.17 | 265 | 1.162 | 0.0782 | Yes | ||
| 5 | TIMP1 | ENSG00000102265.12 | 352 | 1.090 | 0.0898 | Yes | ||
| 6 | SLC7A2 | ENSG00000003989.17 | 597 | 0.979 | 0.0961 | Yes | ||
| 7 | GABBR1 | ENSG00000204681.11 | 700 | 0.941 | 0.1053 | Yes | ||
| 8 | IFITM1 | ENSG00000185885.16 | 785 | 0.916 | 0.1147 | Yes | ||
| 9 | IRF7 | ENSG00000185507.21 | 812 | 0.910 | 0.1255 | Yes | ||
| 10 | CCR7 | ENSG00000126353.3 | 1153 | 0.825 | 0.1275 | Yes | ||
| 11 | GPC3 | ENSG00000147257.14 | 1281 | 0.805 | 0.1345 | Yes | ||
| 12 | IL1R1 | ENSG00000115594.12 | 1321 | 0.799 | 0.1435 | Yes | ||
| 13 | TLR2 | ENSG00000137462.8 | 1599 | 0.759 | 0.1463 | Yes | ||
| 14 | LIF | ENSG00000128342.5 | 1603 | 0.758 | 0.1557 | Yes | ||
| 15 | LPAR1 | ENSG00000198121.13 | 1657 | 0.752 | 0.1638 | Yes | ||
| 16 | CCL7 | ENSG00000108688.11 | 1881 | 0.720 | 0.1673 | Yes | ||
| 17 | AHR | ENSG00000106546.14 | 1908 | 0.715 | 0.1757 | Yes | ||
| 18 | CX3CL1 | ENSG00000006210.7 | 1988 | 0.707 | 0.1826 | Yes | ||
| 19 | TACR1 | ENSG00000115353.11 | 2141 | 0.689 | 0.1875 | Yes | ||
| 20 | IL2RB | ENSG00000100385.14 | 2208 | 0.683 | 0.1944 | Yes | ||
| 21 | PTGER4 | ENSG00000171522.6 | 2872 | 0.616 | 0.1859 | Yes | ||
| 22 | CXCL6 | ENSG00000124875.10 | 2918 | 0.612 | 0.1925 | Yes | ||
| 23 | BTG2 | ENSG00000159388.6 | 2993 | 0.606 | 0.1983 | Yes | ||
| 24 | RGS16 | ENSG00000143333.7 | 3170 | 0.593 | 0.2014 | Yes | ||
| 25 | RTP4 | ENSG00000136514.3 | 3190 | 0.592 | 0.2083 | Yes | ||
| 26 | TPBG | ENSG00000146242.8 | 3275 | 0.586 | 0.2136 | Yes | ||
| 27 | IL1B | ENSG00000125538.12 | 3430 | 0.572 | 0.2170 | Yes | ||
| 28 | PDPN | ENSG00000162493.16 | 3690 | 0.558 | 0.2177 | Yes | ||
| 29 | PDE4B | ENSG00000184588.18 | 4101 | 0.537 | 0.2144 | Yes | ||
| 30 | IL1A | ENSG00000115008.5 | 4274 | 0.526 | 0.2168 | Yes | ||
| 31 | LCP2 | ENSG00000043462.12 | 4413 | 0.519 | 0.2199 | Yes | ||
| 32 | GPR132 | ENSG00000183484.12 | 4540 | 0.516 | 0.2233 | Yes | ||
| 33 | CCL17 | ENSG00000102970.11 | 4670 | 0.512 | 0.2265 | Yes | ||
| 34 | ABCA1 | ENSG00000165029.16 | 4697 | 0.511 | 0.2323 | Yes | ||
| 35 | IL12B | ENSG00000113302.4 | 5068 | 0.486 | 0.2293 | Yes | ||
| 36 | SEMA4D | ENSG00000187764.11 | 5211 | 0.477 | 0.2318 | Yes | ||
| 37 | KCNA3 | ENSG00000177272.9 | 5262 | 0.474 | 0.2365 | Yes | ||
| 38 | NAMPT | ENSG00000105835.12 | 5278 | 0.473 | 0.2421 | Yes | ||
| 39 | IL18RAP | ENSG00000115607.9 | 5756 | 0.445 | 0.2360 | Yes | ||
| 40 | GCH1 | ENSG00000131979.19 | 6236 | 0.421 | 0.2296 | Yes | ||
| 41 | TNFSF10 | ENSG00000121858.11 | 6380 | 0.413 | 0.2313 | Yes | ||
| 42 | TLR1 | ENSG00000174125.8 | 6430 | 0.410 | 0.2352 | Yes | ||
| 43 | LDLR | ENSG00000130164.13 | 6959 | 0.391 | 0.2272 | Yes | ||
| 44 | PTGER2 | ENSG00000125384.7 | 6983 | 0.390 | 0.2315 | Yes | ||
| 45 | RASGRP1 | ENSG00000172575.12 | 7013 | 0.389 | 0.2357 | Yes | ||
| 46 | CMKLR1 | ENSG00000174600.14 | 7266 | 0.377 | 0.2342 | Yes | ||
| 47 | IL18R1 | ENSG00000115604.10 | 7316 | 0.376 | 0.2377 | Yes | ||
| 48 | GP1BA | ENSG00000185245.8 | 7362 | 0.374 | 0.2413 | Yes | ||
| 49 | CYBB | ENSG00000165168.7 | 7450 | 0.372 | 0.2439 | Yes | ||
| 50 | LTA | ENSG00000226979.8 | 8046 | 0.359 | 0.2338 | Yes | ||
| 51 | P2RY2 | ENSG00000175591.11 | 8099 | 0.358 | 0.2370 | Yes | ||
| 52 | SCARF1 | ENSG00000074660.16 | 8123 | 0.357 | 0.2409 | Yes | ||
| 53 | SLC1A2 | ENSG00000110436.13 | 8193 | 0.354 | 0.2437 | Yes | ||
| 54 | CSF3R | ENSG00000119535.17 | 8224 | 0.352 | 0.2473 | Yes | ||
| 55 | IL4R | ENSG00000077238.14 | 8901 | 0.339 | 0.2351 | No | ||
| 56 | IRF1 | ENSG00000125347.14 | 9828 | 0.318 | 0.2165 | No | ||
| 57 | CCL5 | ENSG00000271503.6 | 9843 | 0.317 | 0.2201 | No | ||
| 58 | TACR3 | ENSG00000169836.5 | 9919 | 0.315 | 0.2222 | No | ||
| 59 | ICAM1 | ENSG00000090339.9 | 9924 | 0.314 | 0.2260 | No | ||
| 60 | NPFFR2 | ENSG00000056291.17 | 10245 | 0.305 | 0.2220 | No | ||
| 61 | ICOSLG | ENSG00000160223.17 | 10654 | 0.304 | 0.2159 | No | ||
| 62 | MXD1 | ENSG00000059728.11 | 10676 | 0.304 | 0.2192 | No | ||
| 63 | CD82 | ENSG00000085117.12 | 11688 | 0.283 | 0.1980 | No | ||
| 64 | C5AR1 | ENSG00000197405.8 | 11756 | 0.280 | 0.1999 | No | ||
| 65 | LCK | ENSG00000182866.17 | 11848 | 0.276 | 0.2011 | No | ||
| 66 | LY6E | ENSG00000160932.11 | 12542 | 0.253 | 0.1874 | No | ||
| 67 | MMP14 | ENSG00000157227.13 | 12590 | 0.252 | 0.1894 | No | ||
| 68 | APLNR | ENSG00000134817.10 | 12714 | 0.247 | 0.1895 | No | ||
| 69 | IL15RA | ENSG00000134470.21 | 12907 | 0.241 | 0.1878 | No | ||
| 70 | STAB1 | ENSG00000010327.10 | 13026 | 0.237 | 0.1879 | No | ||
| 71 | P2RX4 | ENSG00000135124.14 | 13164 | 0.233 | 0.1875 | No | ||
| 72 | CCL24 | ENSG00000106178.6 | 13280 | 0.229 | 0.1875 | No | ||
| 73 | MEFV | ENSG00000103313.12 | 13403 | 0.225 | 0.1874 | No | ||
| 74 | ACVR1B | ENSG00000135503.13 | 13757 | 0.214 | 0.1814 | No | ||
| 75 | CD40 | ENSG00000101017.14 | 13803 | 0.213 | 0.1830 | No | ||
| 76 | TNFAIP6 | ENSG00000123610.5 | 14070 | 0.204 | 0.1790 | No | ||
| 77 | EBI3 | ENSG00000105246.6 | 14531 | 0.189 | 0.1702 | No | ||
| 78 | OPRK1 | ENSG00000082556.12 | 14604 | 0.187 | 0.1708 | No | ||
| 79 | CCL2 | ENSG00000108691.9 | 14738 | 0.183 | 0.1698 | No | ||
| 80 | LAMP3 | ENSG00000078081.8 | 14900 | 0.177 | 0.1681 | No | ||
| 81 | CXCL9 | ENSG00000138755.6 | 16417 | 0.133 | 0.1328 | No | ||
| 82 | IFNGR2 | ENSG00000159128.14 | 16912 | 0.119 | 0.1222 | No | ||
| 83 | ROS1 | ENSG00000047936.10 | 17114 | 0.114 | 0.1187 | No | ||
| 84 | ATP2B1 | ENSG00000070961.15 | 17717 | 0.099 | 0.1052 | No | ||
| 85 | CSF1 | ENSG00000184371.14 | 17952 | 0.092 | 0.1007 | No | ||
| 86 | CXCR6 | ENSG00000172215.6 | 18020 | 0.090 | 0.1002 | No | ||
| 87 | OSMR | ENSG00000145623.13 | 18084 | 0.088 | 0.0997 | No | ||
| 88 | LYN | ENSG00000254087.8 | 18130 | 0.087 | 0.0997 | No | ||
| 89 | ADRM1 | ENSG00000130706.13 | 18532 | 0.078 | 0.0909 | No | ||
| 90 | TLR3 | ENSG00000164342.12 | 19180 | 0.060 | 0.0759 | No | ||
| 91 | NFKB1 | ENSG00000109320.13 | 19218 | 0.059 | 0.0757 | No | ||
| 92 | ITGB3 | ENSG00000259207.7 | 19335 | 0.055 | 0.0736 | No | ||
| 93 | CCL20 | ENSG00000115009.13 | 20079 | 0.037 | 0.0559 | No | ||
| 94 | FZD5 | ENSG00000163251.4 | 20082 | 0.037 | 0.0563 | No | ||
| 95 | GNA15 | ENSG00000060558.4 | 20226 | 0.033 | 0.0532 | No | ||
| 96 | EREG | ENSG00000124882.4 | 20508 | 0.024 | 0.0467 | No | ||
| 97 | CXCL11 | ENSG00000169248.13 | 20828 | 0.017 | 0.0391 | No | ||
| 98 | PTAFR | ENSG00000169403.12 | 20842 | 0.016 | 0.0390 | No | ||
| 99 | KCNJ2 | ENSG00000123700.4 | 20935 | 0.014 | 0.0369 | No | ||
| 100 | RIPK2 | ENSG00000104312.8 | 21192 | 0.007 | 0.0308 | No | ||
| 101 | CDKN1A | ENSG00000124762.13 | 21297 | 0.004 | 0.0283 | No | ||
| 102 | CALCRL | ENSG00000064989.13 | 21590 | -0.004 | 0.0212 | No | ||
| 103 | CD55 | ENSG00000196352.15 | 21778 | -0.009 | 0.0167 | No | ||
| 104 | PROK2 | ENSG00000163421.9 | 21882 | -0.011 | 0.0144 | No | ||
| 105 | SCN1B | ENSG00000105711.12 | 22218 | -0.018 | 0.0064 | No | ||
| 106 | EIF2AK2 | ENSG00000055332.18 | 22459 | -0.025 | 0.0009 | No | ||
| 107 | ATP2A2 | ENSG00000174437.17 | 22480 | -0.026 | 0.0007 | No | ||
| 108 | NFKBIA | ENSG00000100906.10 | 22848 | -0.035 | -0.0078 | No | ||
| 109 | RELA | ENSG00000173039.19 | 23632 | -0.056 | -0.0262 | No | ||
| 110 | RGS1 | ENSG00000090104.12 | 23939 | -0.064 | -0.0329 | No | ||
| 111 | SRI | ENSG00000075142.13 | 24123 | -0.067 | -0.0365 | No | ||
| 112 | BEST1 | ENSG00000167995.16 | 24168 | -0.069 | -0.0367 | No | ||
| 113 | IFNAR1 | ENSG00000142166.13 | 24251 | -0.072 | -0.0378 | No | ||
| 114 | PSEN1 | ENSG00000080815.19 | 24443 | -0.076 | -0.0415 | No | ||
| 115 | SELL | ENSG00000188404.10 | 24840 | -0.087 | -0.0501 | No | ||
| 116 | OSM | ENSG00000099985.4 | 25211 | -0.096 | -0.0580 | No | ||
| 117 | ACVR2A | ENSG00000121989.15 | 25237 | -0.097 | -0.0573 | No | ||
| 118 | GPR183 | ENSG00000169508.7 | 25743 | -0.112 | -0.0683 | No | ||
| 119 | SGMS2 | ENSG00000164023.14 | 26053 | -0.122 | -0.0743 | No | ||
| 120 | BST2 | ENSG00000130303.13 | 26105 | -0.124 | -0.0740 | No | ||
| 121 | PCDH7 | ENSG00000169851.15 | 26172 | -0.125 | -0.0740 | No | ||
| 122 | AQP9 | ENSG00000103569.10 | 26201 | -0.126 | -0.0731 | No | ||
| 123 | TAPBP | ENSG00000231925.11 | 26413 | -0.133 | -0.0766 | No | ||
| 124 | PVR | ENSG00000073008.15 | 26458 | -0.134 | -0.0760 | No | ||
| 125 | IL18 | ENSG00000150782.12 | 26822 | -0.144 | -0.0831 | No | ||
| 126 | RHOG | ENSG00000177105.10 | 26871 | -0.145 | -0.0824 | No | ||
| 127 | NOD2 | ENSG00000167207.13 | 26987 | -0.149 | -0.0834 | No | ||
| 128 | NMI | ENSG00000123609.11 | 27198 | -0.155 | -0.0866 | No | ||
| 129 | CHST2 | ENSG00000175040.6 | 27669 | -0.170 | -0.0959 | No | ||
| 130 | SELENOS | ENSG00000131871.15 | 28063 | -0.183 | -0.1032 | No | ||
| 131 | TNFRSF1B | ENSG00000028137.19 | 28280 | -0.189 | -0.1061 | No | ||
| 132 | SLC31A1 | ENSG00000136868.11 | 28289 | -0.189 | -0.1040 | No | ||
| 133 | CSF3 | ENSG00000108342.12 | 28399 | -0.193 | -0.1042 | No | ||
| 134 | MSR1 | ENSG00000038945.15 | 28595 | -0.199 | -0.1065 | No | ||
| 135 | PTPRE | ENSG00000132334.16 | 28934 | -0.210 | -0.1121 | No | ||
| 136 | HIF1A | ENSG00000100644.17 | 29230 | -0.218 | -0.1166 | No | ||
| 137 | IL6 | ENSG00000136244.12 | 29624 | -0.231 | -0.1233 | No | ||
| 138 | MARCO | ENSG00000019169.10 | 29725 | -0.235 | -0.1228 | No | ||
| 139 | ITGB8 | ENSG00000105855.10 | 30265 | -0.253 | -0.1328 | No | ||
| 140 | SLC11A2 | ENSG00000110911.16 | 30597 | -0.262 | -0.1376 | No | ||
| 141 | EMP3 | ENSG00000142227.11 | 30646 | -0.264 | -0.1354 | No | ||
| 142 | RAF1 | ENSG00000132155.11 | 30802 | -0.270 | -0.1359 | No | ||
| 143 | AXL | ENSG00000167601.12 | 31198 | -0.285 | -0.1419 | No | ||
| 144 | CCL22 | ENSG00000102962.5 | 31587 | -0.285 | -0.1478 | No | ||
| 145 | KCNMB2 | ENSG00000197584.12 | 31955 | -0.295 | -0.1531 | No | ||
| 146 | NDP | ENSG00000124479.11 | 32120 | -0.301 | -0.1533 | No | ||
| 147 | SLC28A2 | ENSG00000137860.12 | 32233 | -0.306 | -0.1522 | No | ||
| 148 | GNAI3 | ENSG00000065135.11 | 32448 | -0.315 | -0.1535 | No | ||
| 149 | CXCL10 | ENSG00000169245.6 | 32700 | -0.324 | -0.1556 | No | ||
| 150 | IRAK2 | ENSG00000134070.5 | 32907 | -0.332 | -0.1565 | No | ||
| 151 | VIP | ENSG00000146469.13 | 33186 | -0.336 | -0.1590 | No | ||
| 152 | HRH1 | ENSG00000196639.6 | 33562 | -0.338 | -0.1640 | No | ||
| 153 | BDKRB1 | ENSG00000100739.11 | 33570 | -0.339 | -0.1599 | No | ||
| 154 | CD14 | ENSG00000170458.14 | 34097 | -0.345 | -0.1684 | No | ||
| 155 | PIK3R5 | ENSG00000141506.13 | 34205 | -0.350 | -0.1667 | No | ||
| 156 | C3AR1 | ENSG00000171860.5 | 34621 | -0.353 | -0.1724 | No | ||
| 157 | KIF1B | ENSG00000054523.17 | 34654 | -0.354 | -0.1687 | No | ||
| 158 | NMUR1 | ENSG00000171596.7 | 35208 | -0.374 | -0.1775 | No | ||
| 159 | PLAUR | ENSG00000011422.12 | 35312 | -0.379 | -0.1753 | No | ||
| 160 | RNF144B | ENSG00000137393.9 | 35421 | -0.385 | -0.1731 | No | ||
| 161 | ATP2C1 | ENSG00000017260.19 | 35680 | -0.397 | -0.1745 | No | ||
| 162 | SPHK1 | ENSG00000176170.13 | 35713 | -0.398 | -0.1703 | No | ||
| 163 | FPR1 | ENSG00000171051.8 | 36312 | -0.416 | -0.1797 | No | ||
| 164 | ABI1 | ENSG00000136754.17 | 36378 | -0.420 | -0.1760 | No | ||
| 165 | ITGA5 | ENSG00000161638.11 | 36500 | -0.427 | -0.1736 | No | ||
| 166 | INHBA | ENSG00000122641.11 | 36833 | -0.442 | -0.1762 | No | ||
| 167 | TNFSF15 | ENSG00000181634.8 | 36930 | -0.448 | -0.1729 | No | ||
| 168 | PTGIR | ENSG00000160013.9 | 37306 | -0.471 | -0.1762 | No | ||
| 169 | SERPINE1 | ENSG00000106366.8 | 37533 | -0.487 | -0.1756 | No | ||
| 170 | TNFSF9 | ENSG00000125657.5 | 37744 | -0.507 | -0.1744 | No | ||
| 171 | HBEGF | ENSG00000113070.8 | 38438 | -0.558 | -0.1843 | No | ||
| 172 | IL7R | ENSG00000168685.15 | 38829 | -0.583 | -0.1866 | No | ||
| 173 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.1833 | No | ||
| 174 | OLR1 | ENSG00000173391.9 | 39260 | -0.619 | -0.1818 | No | ||
| 175 | MET | ENSG00000105976.15 | 39317 | -0.624 | -0.1753 | No | ||
| 176 | SLC7A1 | ENSG00000139514.13 | 39608 | -0.647 | -0.1743 | No | ||
| 177 | EDN1 | ENSG00000078401.7 | 39866 | -0.688 | -0.1720 | No | ||
| 178 | ADORA2B | ENSG00000170425.3 | 39882 | -0.692 | -0.1637 | No | ||
| 179 | F3 | ENSG00000117525.14 | 40005 | -0.714 | -0.1577 | No | ||
| 180 | CXCL8 | ENSG00000169429.11 | 40055 | -0.722 | -0.1499 | No | ||
| 181 | CD70 | ENSG00000125726.11 | 40070 | -0.726 | -0.1411 | No | ||
| 182 | KLF6 | ENSG00000067082.15 | 40162 | -0.745 | -0.1340 | No | ||
| 183 | SLC31A2 | ENSG00000136867.11 | 40458 | -0.827 | -0.1309 | No | ||
| 184 | ADM | ENSG00000148926.10 | 40523 | -0.847 | -0.1219 | No | ||
| 185 | DCBLD2 | ENSG00000057019.16 | 40653 | -0.900 | -0.1138 | No | ||
| 186 | MEP1A | ENSG00000112818.10 | 40712 | -0.927 | -0.1036 | No | ||
| 187 | IL10RA | ENSG00000110324.10 | 40784 | -0.953 | -0.0934 | No | ||
| 188 | SLC4A4 | ENSG00000080493.17 | 40795 | -0.961 | -0.0816 | No | ||
| 189 | IL15 | ENSG00000164136.17 | 41012 | -1.156 | -0.0724 | No | ||
| 190 | CD69 | ENSG00000110848.8 | 41040 | -1.206 | -0.0580 | No | ||
| 191 | HAS2 | ENSG00000170961.7 | 41046 | -1.217 | -0.0429 | No | ||
| 192 | TNFRSF9 | ENSG00000049249.8 | 41133 | -1.755 | -0.0230 | No | ||
| 193 | ADGRE1 | ENSG00000174837.15 | 41139 | -1.894 | 0.0005 | No |