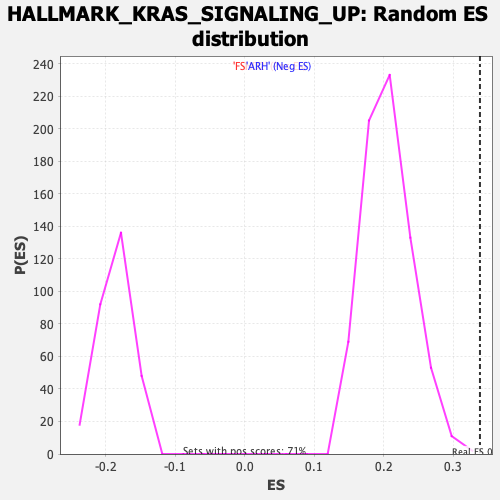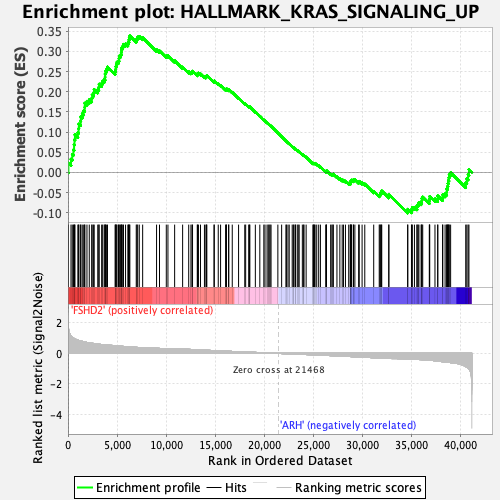
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_KRAS_SIGNALING_UP |
| Enrichment Score (ES) | 0.33839008 |
| Normalized Enrichment Score (NES) | 1.6401646 |
| Nominal p-value | 0.0014164306 |
| FDR q-value | 0.062324457 |
| FWER p-Value | 0.101 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SOX9 | ENSG00000125398.8 | 43 | 1.704 | 0.0233 | Yes | ||
| 2 | ETV1 | ENSG00000006468.14 | 305 | 1.126 | 0.0330 | Yes | ||
| 3 | VWA5A | ENSG00000110002.16 | 433 | 1.042 | 0.0448 | Yes | ||
| 4 | ALDH1A2 | ENSG00000128918.15 | 554 | 0.995 | 0.0561 | Yes | ||
| 5 | PEG3 | ENSG00000198300.14 | 620 | 0.969 | 0.0684 | Yes | ||
| 6 | LCP1 | ENSG00000136167.14 | 660 | 0.955 | 0.0811 | Yes | ||
| 7 | BMP2 | ENSG00000125845.7 | 710 | 0.936 | 0.0933 | Yes | ||
| 8 | CD37 | ENSG00000104894.12 | 1006 | 0.858 | 0.0983 | Yes | ||
| 9 | MALL | ENSG00000144063.3 | 1084 | 0.841 | 0.1085 | Yes | ||
| 10 | PTGS2 | ENSG00000073756.12 | 1105 | 0.837 | 0.1199 | Yes | ||
| 11 | JUP | ENSG00000173801.17 | 1285 | 0.804 | 0.1271 | Yes | ||
| 12 | RELN | ENSG00000189056.14 | 1303 | 0.802 | 0.1381 | Yes | ||
| 13 | IL33 | ENSG00000137033.11 | 1451 | 0.780 | 0.1457 | Yes | ||
| 14 | LIF | ENSG00000128342.5 | 1603 | 0.758 | 0.1528 | Yes | ||
| 15 | ATG10 | ENSG00000152348.16 | 1701 | 0.747 | 0.1611 | Yes | ||
| 16 | FCER1G | ENSG00000158869.11 | 1708 | 0.746 | 0.1716 | Yes | ||
| 17 | LAT2 | ENSG00000086730.17 | 1932 | 0.713 | 0.1764 | Yes | ||
| 18 | CFH | ENSG00000000971.15 | 2168 | 0.686 | 0.1805 | Yes | ||
| 19 | IGFBP3 | ENSG00000146674.15 | 2420 | 0.661 | 0.1838 | Yes | ||
| 20 | GALNT3 | ENSG00000115339.14 | 2460 | 0.658 | 0.1922 | Yes | ||
| 21 | TRIB2 | ENSG00000071575.11 | 2614 | 0.642 | 0.1977 | Yes | ||
| 22 | G0S2 | ENSG00000123689.6 | 2663 | 0.637 | 0.2056 | Yes | ||
| 23 | SATB1 | ENSG00000182568.17 | 3034 | 0.604 | 0.2052 | Yes | ||
| 24 | MAP3K1 | ENSG00000095015.6 | 3142 | 0.596 | 0.2111 | Yes | ||
| 25 | RGS16 | ENSG00000143333.7 | 3170 | 0.593 | 0.2189 | Yes | ||
| 26 | IL1B | ENSG00000125538.12 | 3430 | 0.572 | 0.2208 | Yes | ||
| 27 | SPON1 | ENSG00000262655.4 | 3535 | 0.565 | 0.2263 | Yes | ||
| 28 | SLPI | ENSG00000124107.5 | 3694 | 0.557 | 0.2304 | Yes | ||
| 29 | MMP10 | ENSG00000166670.10 | 3778 | 0.552 | 0.2363 | Yes | ||
| 30 | TRIB1 | ENSG00000173334.4 | 3784 | 0.551 | 0.2440 | Yes | ||
| 31 | CXCR4 | ENSG00000121966.6 | 3819 | 0.548 | 0.2510 | Yes | ||
| 32 | IL1RL2 | ENSG00000115598.10 | 3937 | 0.543 | 0.2559 | Yes | ||
| 33 | FUCA1 | ENSG00000179163.11 | 4024 | 0.539 | 0.2616 | Yes | ||
| 34 | CFB | ENSG00000243649.8 | 4800 | 0.504 | 0.2498 | Yes | ||
| 35 | CCND2 | ENSG00000118971.8 | 4862 | 0.500 | 0.2555 | Yes | ||
| 36 | GADD45G | ENSG00000130222.11 | 4871 | 0.500 | 0.2625 | Yes | ||
| 37 | HSD11B1 | ENSG00000117594.10 | 4922 | 0.496 | 0.2683 | Yes | ||
| 38 | CPE | ENSG00000109472.14 | 4995 | 0.492 | 0.2736 | Yes | ||
| 39 | CBX8 | ENSG00000141570.11 | 5149 | 0.481 | 0.2767 | Yes | ||
| 40 | PCSK1N | ENSG00000102109.9 | 5202 | 0.477 | 0.2823 | Yes | ||
| 41 | GYPC | ENSG00000136732.16 | 5219 | 0.476 | 0.2887 | Yes | ||
| 42 | PRKG2 | ENSG00000138669.9 | 5365 | 0.467 | 0.2918 | Yes | ||
| 43 | SEMA3B | ENSG00000012171.20 | 5425 | 0.463 | 0.2970 | Yes | ||
| 44 | MMP11 | ENSG00000099953.10 | 5438 | 0.463 | 0.3033 | Yes | ||
| 45 | LAPTM5 | ENSG00000162511.8 | 5486 | 0.460 | 0.3088 | Yes | ||
| 46 | PTBP2 | ENSG00000117569.18 | 5584 | 0.454 | 0.3129 | Yes | ||
| 47 | MAFB | ENSG00000204103.4 | 5655 | 0.450 | 0.3176 | Yes | ||
| 48 | SERPINA3 | ENSG00000196136.17 | 5867 | 0.440 | 0.3188 | Yes | ||
| 49 | TSPAN1 | ENSG00000117472.10 | 6102 | 0.428 | 0.3192 | Yes | ||
| 50 | SPARCL1 | ENSG00000152583.12 | 6166 | 0.425 | 0.3237 | Yes | ||
| 51 | F2RL1 | ENSG00000164251.5 | 6217 | 0.422 | 0.3285 | Yes | ||
| 52 | RBP4 | ENSG00000138207.14 | 6241 | 0.421 | 0.3340 | Yes | ||
| 53 | KIF5C | ENSG00000168280.17 | 6305 | 0.417 | 0.3384 | Yes | ||
| 54 | HOXD11 | ENSG00000128713.14 | 6956 | 0.392 | 0.3281 | No | ||
| 55 | ITGA2 | ENSG00000164171.11 | 7028 | 0.388 | 0.3319 | No | ||
| 56 | PRDM1 | ENSG00000057657.17 | 7107 | 0.385 | 0.3355 | No | ||
| 57 | CMKLR1 | ENSG00000174600.14 | 7266 | 0.377 | 0.3371 | No | ||
| 58 | ST6GAL1 | ENSG00000073849.15 | 7610 | 0.366 | 0.3339 | No | ||
| 59 | NAP1L2 | ENSG00000186462.9 | 9027 | 0.334 | 0.3041 | No | ||
| 60 | TLR8 | ENSG00000101916.11 | 9327 | 0.322 | 0.3014 | No | ||
| 61 | ANKH | ENSG00000154122.14 | 10019 | 0.311 | 0.2890 | No | ||
| 62 | PPBP | ENSG00000163736.4 | 10187 | 0.305 | 0.2893 | No | ||
| 63 | GABRA3 | ENSG00000011677.13 | 10870 | 0.296 | 0.2769 | No | ||
| 64 | ETV4 | ENSG00000175832.13 | 11684 | 0.283 | 0.2611 | No | ||
| 65 | ADAMDEC1 | ENSG00000134028.14 | 12304 | 0.261 | 0.2497 | No | ||
| 66 | CA2 | ENSG00000104267.10 | 12533 | 0.253 | 0.2478 | No | ||
| 67 | GPNMB | ENSG00000136235.16 | 12650 | 0.250 | 0.2485 | No | ||
| 68 | CTSS | ENSG00000163131.11 | 12694 | 0.248 | 0.2510 | No | ||
| 69 | KLF4 | ENSG00000136826.15 | 13176 | 0.233 | 0.2426 | No | ||
| 70 | ANO1 | ENSG00000131620.17 | 13204 | 0.232 | 0.2452 | No | ||
| 71 | CIDEA | ENSG00000176194.18 | 13297 | 0.229 | 0.2462 | No | ||
| 72 | APOD | ENSG00000189058.9 | 13505 | 0.221 | 0.2444 | No | ||
| 73 | MAP4K1 | ENSG00000104814.13 | 13908 | 0.209 | 0.2375 | No | ||
| 74 | PSMB8 | ENSG00000204264.11 | 14058 | 0.204 | 0.2368 | No | ||
| 75 | ABCB1 | ENSG00000085563.14 | 14109 | 0.202 | 0.2385 | No | ||
| 76 | EPHB2 | ENSG00000133216.16 | 14152 | 0.201 | 0.2403 | No | ||
| 77 | IGF2 | ENSG00000167244.21 | 14882 | 0.177 | 0.2251 | No | ||
| 78 | ANXA10 | ENSG00000109511.12 | 14912 | 0.177 | 0.2269 | No | ||
| 79 | TMEM100 | ENSG00000166292.12 | 15315 | 0.164 | 0.2194 | No | ||
| 80 | PRRX1 | ENSG00000116132.12 | 15553 | 0.158 | 0.2159 | No | ||
| 81 | CDADC1 | ENSG00000102543.14 | 16066 | 0.143 | 0.2054 | No | ||
| 82 | TRAF1 | ENSG00000056558.11 | 16131 | 0.142 | 0.2059 | No | ||
| 83 | DUSP6 | ENSG00000139318.8 | 16150 | 0.141 | 0.2075 | No | ||
| 84 | ACE | ENSG00000159640.16 | 16370 | 0.135 | 0.2041 | No | ||
| 85 | RBM4 | ENSG00000173933.20 | 16388 | 0.134 | 0.2056 | No | ||
| 86 | GFPT2 | ENSG00000131459.13 | 16742 | 0.124 | 0.1987 | No | ||
| 87 | IKZF1 | ENSG00000185811.19 | 17387 | 0.107 | 0.1845 | No | ||
| 88 | TSPAN13 | ENSG00000106537.8 | 18017 | 0.090 | 0.1705 | No | ||
| 89 | AKT2 | ENSG00000105221.18 | 18110 | 0.088 | 0.1695 | No | ||
| 90 | ADAM17 | ENSG00000151694.14 | 18419 | 0.082 | 0.1631 | No | ||
| 91 | WNT7A | ENSG00000154764.6 | 18492 | 0.079 | 0.1625 | No | ||
| 92 | FGF9 | ENSG00000102678.7 | 18537 | 0.078 | 0.1625 | No | ||
| 93 | ADGRA2 | ENSG00000020181.18 | 19096 | 0.063 | 0.1498 | No | ||
| 94 | MMD | ENSG00000108960.9 | 19552 | 0.050 | 0.1394 | No | ||
| 95 | CROT | ENSG00000005469.11 | 19961 | 0.039 | 0.1300 | No | ||
| 96 | CCL20 | ENSG00000115009.13 | 20079 | 0.037 | 0.1277 | No | ||
| 97 | TFPI | ENSG00000003436.16 | 20290 | 0.032 | 0.1230 | No | ||
| 98 | DNMBP | ENSG00000107554.17 | 20411 | 0.028 | 0.1205 | No | ||
| 99 | EREG | ENSG00000124882.4 | 20508 | 0.024 | 0.1185 | No | ||
| 100 | EPB41L3 | ENSG00000082397.17 | 20603 | 0.022 | 0.1165 | No | ||
| 101 | PECAM1 | ENSG00000261371.6 | 20696 | 0.020 | 0.1145 | No | ||
| 102 | CLEC4A | ENSG00000111729.14 | 21391 | 0.001 | 0.0976 | No | ||
| 103 | MTMR10 | ENSG00000166912.17 | 21776 | -0.009 | 0.0884 | No | ||
| 104 | SCN1B | ENSG00000105711.12 | 22218 | -0.018 | 0.0779 | No | ||
| 105 | TPH1 | ENSG00000129167.9 | 22259 | -0.020 | 0.0772 | No | ||
| 106 | FLT4 | ENSG00000037280.16 | 22421 | -0.024 | 0.0736 | No | ||
| 107 | USH1C | ENSG00000006611.16 | 22548 | -0.027 | 0.0709 | No | ||
| 108 | ADGRL4 | ENSG00000162618.15 | 22853 | -0.035 | 0.0640 | No | ||
| 109 | CBR4 | ENSG00000145439.12 | 22970 | -0.038 | 0.0617 | No | ||
| 110 | HDAC9 | ENSG00000048052.21 | 23030 | -0.040 | 0.0608 | No | ||
| 111 | GPRC5B | ENSG00000167191.12 | 23172 | -0.044 | 0.0580 | No | ||
| 112 | ETV5 | ENSG00000244405.8 | 23177 | -0.044 | 0.0586 | No | ||
| 113 | HIST1H2BB | ENSG00000276410.3 | 23366 | -0.049 | 0.0547 | No | ||
| 114 | WDR33 | ENSG00000136709.12 | 23466 | -0.051 | 0.0530 | No | ||
| 115 | GLRX | ENSG00000173221.14 | 23563 | -0.054 | 0.0514 | No | ||
| 116 | ENG | ENSG00000106991.13 | 23886 | -0.063 | 0.0445 | No | ||
| 117 | TMEM176A | ENSG00000002933.9 | 24023 | -0.065 | 0.0421 | No | ||
| 118 | ZNF277 | ENSG00000198839.10 | 24048 | -0.066 | 0.0424 | No | ||
| 119 | ARG1 | ENSG00000118520.15 | 24273 | -0.072 | 0.0380 | No | ||
| 120 | NIN | ENSG00000100503.23 | 24962 | -0.090 | 0.0225 | No | ||
| 121 | CBL | ENSG00000110395.6 | 25021 | -0.092 | 0.0224 | No | ||
| 122 | PLVAP | ENSG00000130300.9 | 25056 | -0.093 | 0.0229 | No | ||
| 123 | TMEM176B | ENSG00000106565.18 | 25133 | -0.094 | 0.0224 | No | ||
| 124 | MMP9 | ENSG00000100985.7 | 25181 | -0.096 | 0.0226 | No | ||
| 125 | SCG3 | ENSG00000104112.9 | 25353 | -0.101 | 0.0199 | No | ||
| 126 | MAP7 | ENSG00000135525.18 | 25553 | -0.107 | 0.0165 | No | ||
| 127 | GNG11 | ENSG00000127920.6 | 25729 | -0.112 | 0.0139 | No | ||
| 128 | CFHR2 | ENSG00000080910.13 | 26270 | -0.128 | 0.0025 | No | ||
| 129 | PTCD2 | ENSG00000049883.15 | 26339 | -0.130 | 0.0027 | No | ||
| 130 | BTC | ENSG00000174808.12 | 26349 | -0.131 | 0.0043 | No | ||
| 131 | NR1H4 | ENSG00000012504.15 | 26772 | -0.143 | -0.0039 | No | ||
| 132 | CAB39L | ENSG00000102547.19 | 26895 | -0.146 | -0.0048 | No | ||
| 133 | PPP1R15A | ENSG00000087074.8 | 27000 | -0.149 | -0.0052 | No | ||
| 134 | TNFAIP3 | ENSG00000118503.15 | 27043 | -0.151 | -0.0041 | No | ||
| 135 | PCP4 | ENSG00000183036.11 | 27414 | -0.161 | -0.0108 | No | ||
| 136 | TOR1AIP2 | ENSG00000169905.12 | 27711 | -0.171 | -0.0156 | No | ||
| 137 | ITGB2 | ENSG00000160255.18 | 27949 | -0.179 | -0.0188 | No | ||
| 138 | PIGR | ENSG00000162896.6 | 28078 | -0.183 | -0.0193 | No | ||
| 139 | TNFRSF1B | ENSG00000028137.19 | 28280 | -0.189 | -0.0215 | No | ||
| 140 | SNAP25 | ENSG00000132639.12 | 28613 | -0.200 | -0.0268 | No | ||
| 141 | ZNF639 | ENSG00000121864.10 | 28764 | -0.204 | -0.0275 | No | ||
| 142 | SPRY2 | ENSG00000136158.12 | 28819 | -0.206 | -0.0259 | No | ||
| 143 | KCNN4 | ENSG00000104783.14 | 28826 | -0.206 | -0.0231 | No | ||
| 144 | BTBD3 | ENSG00000132640.15 | 28845 | -0.207 | -0.0206 | No | ||
| 145 | ANGPTL4 | ENSG00000167772.12 | 28905 | -0.208 | -0.0191 | No | ||
| 146 | NR0B2 | ENSG00000131910.5 | 29084 | -0.215 | -0.0203 | No | ||
| 147 | BPGM | ENSG00000172331.12 | 29104 | -0.215 | -0.0177 | No | ||
| 148 | TNNT2 | ENSG00000118194.20 | 29241 | -0.219 | -0.0179 | No | ||
| 149 | PDCD1LG2 | ENSG00000197646.8 | 29627 | -0.232 | -0.0240 | No | ||
| 150 | STRN | ENSG00000115808.12 | 29685 | -0.234 | -0.0220 | No | ||
| 151 | SNAP91 | ENSG00000065609.14 | 29977 | -0.243 | -0.0257 | No | ||
| 152 | SDCCAG8 | ENSG00000054282.16 | 30251 | -0.252 | -0.0287 | No | ||
| 153 | GUCY1A1 | ENSG00000164116.16 | 31161 | -0.283 | -0.0469 | No | ||
| 154 | TMEM158 | ENSG00000249992.2 | 31724 | -0.286 | -0.0565 | No | ||
| 155 | HKDC1 | ENSG00000156510.13 | 31806 | -0.290 | -0.0543 | No | ||
| 156 | ETS1 | ENSG00000134954.14 | 31835 | -0.291 | -0.0509 | No | ||
| 157 | ERO1A | ENSG00000197930.13 | 31946 | -0.294 | -0.0493 | No | ||
| 158 | LY96 | ENSG00000154589.7 | 31980 | -0.295 | -0.0459 | No | ||
| 159 | CXCL10 | ENSG00000169245.6 | 32700 | -0.324 | -0.0588 | No | ||
| 160 | PLAU | ENSG00000122861.16 | 32703 | -0.324 | -0.0543 | No | ||
| 161 | C3AR1 | ENSG00000171860.5 | 34621 | -0.353 | -0.0960 | No | ||
| 162 | ALDH1A3 | ENSG00000184254.17 | 34635 | -0.353 | -0.0913 | No | ||
| 163 | PTPRR | ENSG00000153233.13 | 35009 | -0.365 | -0.0952 | No | ||
| 164 | BIRC3 | ENSG00000023445.14 | 35047 | -0.367 | -0.0908 | No | ||
| 165 | SCG5 | ENSG00000166922.8 | 35094 | -0.368 | -0.0867 | No | ||
| 166 | PLAUR | ENSG00000011422.12 | 35312 | -0.379 | -0.0866 | No | ||
| 167 | PLAT | ENSG00000104368.18 | 35576 | -0.392 | -0.0874 | No | ||
| 168 | RABGAP1L | ENSG00000152061.23 | 35578 | -0.393 | -0.0818 | No | ||
| 169 | EVI5 | ENSG00000067208.14 | 35690 | -0.397 | -0.0788 | No | ||
| 170 | AVL9 | ENSG00000105778.19 | 35769 | -0.400 | -0.0750 | No | ||
| 171 | FBXO4 | ENSG00000151876.13 | 35992 | -0.405 | -0.0746 | No | ||
| 172 | F13A1 | ENSG00000124491.15 | 36056 | -0.406 | -0.0703 | No | ||
| 173 | ID2 | ENSG00000115738.10 | 36059 | -0.406 | -0.0646 | No | ||
| 174 | IL2RG | ENSG00000147168.12 | 36157 | -0.412 | -0.0611 | No | ||
| 175 | INHBA | ENSG00000122641.11 | 36833 | -0.442 | -0.0712 | No | ||
| 176 | DOCK2 | ENSG00000134516.17 | 36842 | -0.442 | -0.0651 | No | ||
| 177 | EMP1 | ENSG00000134531.10 | 36872 | -0.444 | -0.0595 | No | ||
| 178 | USP12 | ENSG00000152484.14 | 37406 | -0.478 | -0.0656 | No | ||
| 179 | AKAP12 | ENSG00000131016.17 | 37685 | -0.501 | -0.0653 | No | ||
| 180 | YRDC | ENSG00000196449.4 | 37693 | -0.502 | -0.0583 | No | ||
| 181 | AMMECR1 | ENSG00000101935.10 | 38178 | -0.541 | -0.0624 | No | ||
| 182 | CSF2 | ENSG00000164400.6 | 38203 | -0.543 | -0.0552 | No | ||
| 183 | HBEGF | ENSG00000113070.8 | 38438 | -0.558 | -0.0529 | No | ||
| 184 | PRELID3B | ENSG00000101166.16 | 38605 | -0.562 | -0.0489 | No | ||
| 185 | NGF | ENSG00000134259.3 | 38608 | -0.563 | -0.0410 | No | ||
| 186 | SPP1 | ENSG00000118785.14 | 38686 | -0.569 | -0.0347 | No | ||
| 187 | TSPAN7 | ENSG00000156298.12 | 38729 | -0.573 | -0.0275 | No | ||
| 188 | CCSER2 | ENSG00000107771.17 | 38766 | -0.577 | -0.0202 | No | ||
| 189 | IL7R | ENSG00000168685.15 | 38829 | -0.583 | -0.0133 | No | ||
| 190 | ITGBL1 | ENSG00000198542.14 | 38854 | -0.585 | -0.0056 | No | ||
| 191 | ADAM8 | ENSG00000151651.16 | 39002 | -0.601 | -0.0006 | No | ||
| 192 | NRP1 | ENSG00000099250.18 | 40541 | -0.854 | -0.0259 | No | ||
| 193 | DCBLD2 | ENSG00000057019.16 | 40653 | -0.900 | -0.0157 | No | ||
| 194 | IL10RA | ENSG00000110324.10 | 40784 | -0.953 | -0.0053 | No | ||
| 195 | PLEK2 | ENSG00000100558.9 | 40868 | -1.012 | 0.0072 | No |