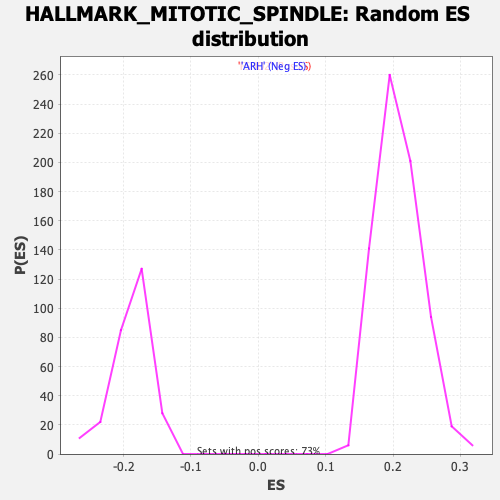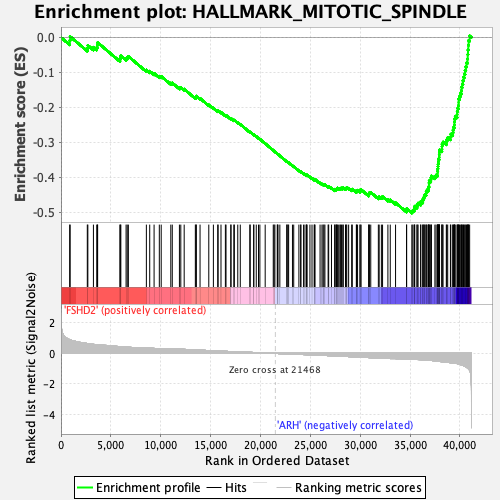
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_MITOTIC_SPINDLE |
| Enrichment Score (ES) | -0.5029521 |
| Normalized Enrichment Score (NES) | -2.6628025 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RHOF | ENSG00000139725.8 | 871 | 0.891 | -0.0087 | No | ||
| 2 | EPB41 | ENSG00000159023.21 | 916 | 0.879 | 0.0027 | No | ||
| 3 | BCL2L11 | ENSG00000153094.23 | 2637 | 0.640 | -0.0303 | No | ||
| 4 | RAPGEF6 | ENSG00000158987.21 | 2692 | 0.634 | -0.0227 | No | ||
| 5 | ABL1 | ENSG00000097007.19 | 3251 | 0.588 | -0.0280 | No | ||
| 6 | CYTH2 | ENSG00000105443.15 | 3585 | 0.563 | -0.0282 | No | ||
| 7 | CDK5RAP2 | ENSG00000136861.18 | 3658 | 0.560 | -0.0220 | No | ||
| 8 | TUBGCP6 | ENSG00000128159.12 | 3674 | 0.559 | -0.0145 | No | ||
| 9 | ATG4B | ENSG00000168397.17 | 5907 | 0.438 | -0.0628 | No | ||
| 10 | NUMA1 | ENSG00000137497.17 | 5945 | 0.436 | -0.0575 | No | ||
| 11 | ABR | ENSG00000159842.15 | 6000 | 0.433 | -0.0527 | No | ||
| 12 | MID1IP1 | ENSG00000165175.15 | 6526 | 0.407 | -0.0598 | No | ||
| 13 | KPTN | ENSG00000118162.14 | 6642 | 0.402 | -0.0569 | No | ||
| 14 | CEP131 | ENSG00000141577.14 | 6755 | 0.398 | -0.0541 | No | ||
| 15 | PREX1 | ENSG00000124126.14 | 8560 | 0.349 | -0.0932 | No | ||
| 16 | SHROOM1 | ENSG00000164403.14 | 8896 | 0.340 | -0.0966 | No | ||
| 17 | RHOT2 | ENSG00000140983.14 | 9334 | 0.322 | -0.1027 | No | ||
| 18 | CDC42EP2 | ENSG00000149798.5 | 9859 | 0.317 | -0.1110 | No | ||
| 19 | SYNPO | ENSG00000171992.13 | 10048 | 0.310 | -0.1112 | No | ||
| 20 | HDAC6 | ENSG00000094631.21 | 11011 | 0.290 | -0.1306 | No | ||
| 21 | SUN2 | ENSG00000100242.15 | 11158 | 0.285 | -0.1301 | No | ||
| 22 | FGD4 | ENSG00000139132.14 | 11874 | 0.275 | -0.1437 | No | ||
| 23 | BCR | ENSG00000186716.21 | 12013 | 0.270 | -0.1432 | No | ||
| 24 | MYH10 | ENSG00000133026.12 | 12352 | 0.259 | -0.1478 | No | ||
| 25 | FSCN1 | ENSG00000075618.18 | 13471 | 0.223 | -0.1720 | No | ||
| 26 | ARHGEF11 | ENSG00000132694.18 | 13549 | 0.221 | -0.1708 | No | ||
| 27 | CNTRL | ENSG00000119397.16 | 13560 | 0.220 | -0.1679 | No | ||
| 28 | ARAP3 | ENSG00000120318.16 | 13943 | 0.208 | -0.1743 | No | ||
| 29 | KLC1 | ENSG00000126214.21 | 14819 | 0.180 | -0.1931 | No | ||
| 30 | AC027237.1 | ENSG00000259191.2 | 15280 | 0.165 | -0.2020 | No | ||
| 31 | CDC42EP4 | ENSG00000179604.10 | 15721 | 0.153 | -0.2106 | No | ||
| 32 | RAPGEF5 | ENSG00000136237.19 | 15751 | 0.152 | -0.2092 | No | ||
| 33 | TUBD1 | ENSG00000108423.15 | 16043 | 0.144 | -0.2142 | No | ||
| 34 | GEMIN4 | ENSG00000179409.11 | 16483 | 0.131 | -0.2231 | No | ||
| 35 | CLIP2 | ENSG00000106665.15 | 16546 | 0.130 | -0.2228 | No | ||
| 36 | CEP192 | ENSG00000101639.18 | 17033 | 0.117 | -0.2330 | No | ||
| 37 | CEP57 | ENSG00000166037.11 | 17042 | 0.117 | -0.2315 | No | ||
| 38 | NCK2 | ENSG00000071051.14 | 17333 | 0.109 | -0.2371 | No | ||
| 39 | AKAP13 | ENSG00000170776.21 | 17381 | 0.108 | -0.2367 | No | ||
| 40 | TAOK2 | ENSG00000149930.18 | 17725 | 0.098 | -0.2437 | No | ||
| 41 | PLEKHG2 | ENSG00000090924.15 | 17977 | 0.091 | -0.2485 | No | ||
| 42 | CSNK1D | ENSG00000141551.14 | 18935 | 0.067 | -0.2709 | No | ||
| 43 | STK38L | ENSG00000211455.8 | 18970 | 0.066 | -0.2709 | No | ||
| 44 | ARHGAP29 | ENSG00000137962.13 | 19325 | 0.056 | -0.2787 | No | ||
| 45 | MYO1E | ENSG00000157483.9 | 19330 | 0.055 | -0.2780 | No | ||
| 46 | TSC1 | ENSG00000165699.14 | 19580 | 0.049 | -0.2834 | No | ||
| 47 | RICTOR | ENSG00000164327.13 | 19806 | 0.043 | -0.2883 | No | ||
| 48 | ARHGAP4 | ENSG00000089820.15 | 19943 | 0.040 | -0.2910 | No | ||
| 49 | TIAM1 | ENSG00000156299.13 | 20475 | 0.025 | -0.3037 | No | ||
| 50 | TUBA4A | ENSG00000127824.14 | 21284 | 0.004 | -0.3233 | No | ||
| 51 | SAC3D1 | ENSG00000168061.15 | 21362 | 0.002 | -0.3252 | No | ||
| 52 | ARFIP2 | ENSG00000132254.12 | 21447 | 0.000 | -0.3272 | No | ||
| 53 | SMC4 | ENSG00000113810.16 | 21685 | -0.006 | -0.3329 | No | ||
| 54 | ARHGEF7 | ENSG00000102606.18 | 21717 | -0.007 | -0.3336 | No | ||
| 55 | ARHGEF2 | ENSG00000116584.19 | 21758 | -0.008 | -0.3344 | No | ||
| 56 | CNTROB | ENSG00000170037.14 | 21945 | -0.013 | -0.3388 | No | ||
| 57 | KATNB1 | ENSG00000140854.13 | 22637 | -0.029 | -0.3553 | No | ||
| 58 | SOS1 | ENSG00000115904.12 | 22667 | -0.030 | -0.3555 | No | ||
| 59 | TUBGCP2 | ENSG00000130640.13 | 22760 | -0.032 | -0.3573 | No | ||
| 60 | WASF2 | ENSG00000158195.11 | 22827 | -0.034 | -0.3585 | No | ||
| 61 | NUSAP1 | ENSG00000137804.14 | 23210 | -0.045 | -0.3672 | No | ||
| 62 | DYNLL2 | ENSG00000264364.3 | 23336 | -0.048 | -0.3695 | No | ||
| 63 | PCNT | ENSG00000160299.17 | 23855 | -0.062 | -0.3813 | No | ||
| 64 | TBCD | ENSG00000141556.21 | 24035 | -0.065 | -0.3847 | No | ||
| 65 | PXN | ENSG00000089159.16 | 24055 | -0.066 | -0.3843 | No | ||
| 66 | LRPPRC | ENSG00000138095.19 | 24323 | -0.074 | -0.3898 | No | ||
| 67 | ALMS1 | ENSG00000116127.18 | 24464 | -0.076 | -0.3921 | No | ||
| 68 | MARK4 | ENSG00000007047.15 | 24617 | -0.080 | -0.3947 | No | ||
| 69 | STAU1 | ENSG00000124214.19 | 24627 | -0.081 | -0.3938 | No | ||
| 70 | MAP3K11 | ENSG00000173327.8 | 24676 | -0.082 | -0.3938 | No | ||
| 71 | NIN | ENSG00000100503.23 | 24962 | -0.090 | -0.3995 | No | ||
| 72 | BIN1 | ENSG00000136717.15 | 25170 | -0.095 | -0.4032 | No | ||
| 73 | ITSN1 | ENSG00000205726.15 | 25392 | -0.102 | -0.4071 | No | ||
| 74 | HOOK3 | ENSG00000168172.9 | 25411 | -0.103 | -0.4061 | No | ||
| 75 | CCDC88A | ENSG00000115355.17 | 25474 | -0.104 | -0.4061 | No | ||
| 76 | TUBGCP5 | ENSG00000275835.5 | 25978 | -0.120 | -0.4167 | No | ||
| 77 | RFC1 | ENSG00000035928.16 | 26169 | -0.125 | -0.4196 | No | ||
| 78 | CENPJ | ENSG00000151849.15 | 26281 | -0.128 | -0.4205 | No | ||
| 79 | ARHGEF3 | ENSG00000163947.11 | 26368 | -0.131 | -0.4208 | No | ||
| 80 | PPP4R2 | ENSG00000163605.15 | 26463 | -0.134 | -0.4212 | No | ||
| 81 | RANBP9 | ENSG00000010017.13 | 26787 | -0.143 | -0.4270 | No | ||
| 82 | FARP1 | ENSG00000152767.17 | 26841 | -0.145 | -0.4263 | No | ||
| 83 | ARHGAP10 | ENSG00000071205.12 | 27122 | -0.153 | -0.4310 | No | ||
| 84 | CDC42EP1 | ENSG00000128283.7 | 27451 | -0.163 | -0.4367 | No | ||
| 85 | KATNA1 | ENSG00000186625.14 | 27460 | -0.163 | -0.4346 | No | ||
| 86 | BCAR1 | ENSG00000050820.17 | 27576 | -0.167 | -0.4350 | No | ||
| 87 | LLGL1 | ENSG00000131899.11 | 27619 | -0.168 | -0.4337 | No | ||
| 88 | PCM1 | ENSG00000078674.17 | 27675 | -0.170 | -0.4326 | No | ||
| 89 | KIFAP3 | ENSG00000075945.13 | 27710 | -0.171 | -0.4310 | No | ||
| 90 | MYO9B | ENSG00000099331.13 | 27847 | -0.175 | -0.4319 | No | ||
| 91 | WASL | ENSG00000106299.8 | 27968 | -0.179 | -0.4323 | No | ||
| 92 | CLASP1 | ENSG00000074054.18 | 28043 | -0.182 | -0.4315 | No | ||
| 93 | NOTCH2 | ENSG00000134250.20 | 28133 | -0.185 | -0.4311 | No | ||
| 94 | UXT | ENSG00000126756.12 | 28258 | -0.188 | -0.4314 | No | ||
| 95 | TUBGCP3 | ENSG00000126216.15 | 28303 | -0.190 | -0.4298 | No | ||
| 96 | GSN | ENSG00000148180.19 | 28530 | -0.197 | -0.4326 | No | ||
| 97 | MAP1S | ENSG00000130479.11 | 28590 | -0.199 | -0.4312 | No | ||
| 98 | CTTN | ENSG00000085733.16 | 28654 | -0.201 | -0.4299 | No | ||
| 99 | MARCKS | ENSG00000277443.3 | 28843 | -0.207 | -0.4316 | No | ||
| 100 | SPTBN1 | ENSG00000115306.16 | 29149 | -0.216 | -0.4360 | No | ||
| 101 | SSH2 | ENSG00000141298.19 | 29207 | -0.218 | -0.4343 | No | ||
| 102 | ARL8A | ENSG00000143862.8 | 29618 | -0.231 | -0.4410 | No | ||
| 103 | SPTAN1 | ENSG00000197694.16 | 29628 | -0.232 | -0.4380 | No | ||
| 104 | CDC27 | ENSG00000004897.12 | 29738 | -0.236 | -0.4373 | No | ||
| 105 | NF1 | ENSG00000196712.17 | 29983 | -0.243 | -0.4398 | No | ||
| 106 | CEP72 | ENSG00000112877.8 | 29993 | -0.243 | -0.4366 | No | ||
| 107 | SHROOM2 | ENSG00000146950.13 | 30092 | -0.247 | -0.4355 | No | ||
| 108 | LATS1 | ENSG00000131023.13 | 30857 | -0.271 | -0.4503 | No | ||
| 109 | DST | ENSG00000151914.20 | 30866 | -0.272 | -0.4467 | No | ||
| 110 | KIF22 | ENSG00000079616.12 | 30925 | -0.274 | -0.4442 | No | ||
| 111 | KIF3C | ENSG00000084731.15 | 31061 | -0.279 | -0.4436 | No | ||
| 112 | DYNC1H1 | ENSG00000197102.12 | 31822 | -0.290 | -0.4580 | No | ||
| 113 | ARHGEF12 | ENSG00000196914.9 | 31905 | -0.293 | -0.4559 | No | ||
| 114 | INCENP | ENSG00000149503.13 | 32126 | -0.301 | -0.4570 | No | ||
| 115 | ARHGAP27 | ENSG00000159314.11 | 32245 | -0.307 | -0.4556 | No | ||
| 116 | ARFGEF1 | ENSG00000066777.9 | 32780 | -0.327 | -0.4640 | No | ||
| 117 | TLK1 | ENSG00000198586.14 | 33016 | -0.336 | -0.4650 | No | ||
| 118 | NEDD9 | ENSG00000111859.17 | 33548 | -0.338 | -0.4732 | No | ||
| 119 | KIF1B | ENSG00000054523.17 | 34654 | -0.354 | -0.4951 | No | ||
| 120 | RASA1 | ENSG00000145715.15 | 34655 | -0.354 | -0.4901 | No | ||
| 121 | CEP250 | ENSG00000126001.16 | 35181 | -0.373 | -0.4977 | Yes | ||
| 122 | DLG1 | ENSG00000075711.21 | 35280 | -0.377 | -0.4948 | Yes | ||
| 123 | CAPZB | ENSG00000077549.18 | 35414 | -0.384 | -0.4926 | Yes | ||
| 124 | RAB3GAP1 | ENSG00000115839.17 | 35438 | -0.385 | -0.4877 | Yes | ||
| 125 | DOCK4 | ENSG00000128512.22 | 35473 | -0.387 | -0.4831 | Yes | ||
| 126 | WASF1 | ENSG00000112290.13 | 35700 | -0.397 | -0.4830 | Yes | ||
| 127 | RALBP1 | ENSG00000017797.13 | 35709 | -0.398 | -0.4775 | Yes | ||
| 128 | NCK1 | ENSG00000158092.7 | 35806 | -0.402 | -0.4742 | Yes | ||
| 129 | RASA2 | ENSG00000155903.12 | 36064 | -0.407 | -0.4747 | Yes | ||
| 130 | PAFAH1B1 | ENSG00000007168.13 | 36076 | -0.408 | -0.4693 | Yes | ||
| 131 | BIRC5 | ENSG00000089685.15 | 36279 | -0.415 | -0.4683 | Yes | ||
| 132 | ESPL1 | ENSG00000135476.11 | 36287 | -0.415 | -0.4626 | Yes | ||
| 133 | ABI1 | ENSG00000136754.17 | 36378 | -0.420 | -0.4589 | Yes | ||
| 134 | OPHN1 | ENSG00000079482.13 | 36417 | -0.422 | -0.4539 | Yes | ||
| 135 | YWHAE | ENSG00000108953.17 | 36492 | -0.426 | -0.4497 | Yes | ||
| 136 | CCNB2 | ENSG00000157456.8 | 36613 | -0.432 | -0.4465 | Yes | ||
| 137 | ARF6 | ENSG00000165527.7 | 36619 | -0.432 | -0.4405 | Yes | ||
| 138 | PLK1 | ENSG00000166851.15 | 36707 | -0.436 | -0.4365 | Yes | ||
| 139 | DOCK2 | ENSG00000134516.17 | 36842 | -0.442 | -0.4335 | Yes | ||
| 140 | KIF2C | ENSG00000142945.13 | 36852 | -0.443 | -0.4275 | Yes | ||
| 141 | TRIO | ENSG00000038382.20 | 36905 | -0.446 | -0.4225 | Yes | ||
| 142 | FGD6 | ENSG00000180263.14 | 36907 | -0.447 | -0.4162 | Yes | ||
| 143 | SMC3 | ENSG00000108055.9 | 36921 | -0.447 | -0.4102 | Yes | ||
| 144 | ARHGDIA | ENSG00000141522.12 | 37082 | -0.457 | -0.4076 | Yes | ||
| 145 | MAPRE1 | ENSG00000101367.9 | 37101 | -0.458 | -0.4016 | Yes | ||
| 146 | SASS6 | ENSG00000156876.10 | 37143 | -0.460 | -0.3961 | Yes | ||
| 147 | PALLD | ENSG00000129116.19 | 37495 | -0.484 | -0.3978 | Yes | ||
| 148 | KIF4A | ENSG00000090889.12 | 37615 | -0.495 | -0.3938 | Yes | ||
| 149 | DLGAP5 | ENSG00000126787.13 | 37762 | -0.508 | -0.3902 | Yes | ||
| 150 | BUB1 | ENSG00000169679.15 | 37763 | -0.508 | -0.3830 | Yes | ||
| 151 | FBXO5 | ENSG00000112029.10 | 37782 | -0.511 | -0.3762 | Yes | ||
| 152 | KIF3B | ENSG00000101350.8 | 37801 | -0.512 | -0.3694 | Yes | ||
| 153 | CKAP5 | ENSG00000175216.15 | 37814 | -0.514 | -0.3624 | Yes | ||
| 154 | FLNA | ENSG00000196924.17 | 37839 | -0.515 | -0.3558 | Yes | ||
| 155 | CD2AP | ENSG00000198087.7 | 37848 | -0.516 | -0.3487 | Yes | ||
| 156 | SMC1A | ENSG00000072501.17 | 37922 | -0.522 | -0.3431 | Yes | ||
| 157 | NDC80 | ENSG00000080986.13 | 37927 | -0.522 | -0.3358 | Yes | ||
| 158 | VCL | ENSG00000035403.17 | 37935 | -0.523 | -0.3286 | Yes | ||
| 159 | NEK2 | ENSG00000117650.13 | 37958 | -0.525 | -0.3217 | Yes | ||
| 160 | CDC42 | ENSG00000070831.17 | 38194 | -0.542 | -0.3198 | Yes | ||
| 161 | KIF15 | ENSG00000163808.17 | 38204 | -0.543 | -0.3123 | Yes | ||
| 162 | TTK | ENSG00000112742.10 | 38213 | -0.544 | -0.3049 | Yes | ||
| 163 | ALS2 | ENSG00000003393.15 | 38310 | -0.553 | -0.2994 | Yes | ||
| 164 | AURKA | ENSG00000087586.17 | 38638 | -0.565 | -0.2994 | Yes | ||
| 165 | TPX2 | ENSG00000088325.16 | 38681 | -0.569 | -0.2924 | Yes | ||
| 166 | CDC42BPA | ENSG00000143776.18 | 38777 | -0.578 | -0.2866 | Yes | ||
| 167 | ROCK1 | ENSG00000067900.8 | 39050 | -0.606 | -0.2846 | Yes | ||
| 168 | PIF1 | ENSG00000140451.13 | 39086 | -0.611 | -0.2769 | Yes | ||
| 169 | CLIP1 | ENSG00000130779.20 | 39284 | -0.622 | -0.2729 | Yes | ||
| 170 | TOP2A | ENSG00000131747.15 | 39313 | -0.624 | -0.2648 | Yes | ||
| 171 | ARHGAP5 | ENSG00000100852.13 | 39363 | -0.626 | -0.2571 | Yes | ||
| 172 | CENPF | ENSG00000117724.13 | 39444 | -0.631 | -0.2502 | Yes | ||
| 173 | CDK1 | ENSG00000170312.16 | 39453 | -0.632 | -0.2414 | Yes | ||
| 174 | RACGAP1 | ENSG00000161800.13 | 39469 | -0.633 | -0.2329 | Yes | ||
| 175 | MYH9 | ENSG00000100345.21 | 39531 | -0.641 | -0.2253 | Yes | ||
| 176 | LMNB1 | ENSG00000113368.12 | 39725 | -0.665 | -0.2207 | Yes | ||
| 177 | SORBS2 | ENSG00000154556.18 | 39730 | -0.665 | -0.2114 | Yes | ||
| 178 | PRC1 | ENSG00000198901.14 | 39769 | -0.668 | -0.2029 | Yes | ||
| 179 | APC | ENSG00000134982.17 | 39848 | -0.685 | -0.1951 | Yes | ||
| 180 | BRCA2 | ENSG00000139618.15 | 39873 | -0.690 | -0.1859 | Yes | ||
| 181 | KNTC1 | ENSG00000184445.12 | 39875 | -0.690 | -0.1762 | Yes | ||
| 182 | ANLN | ENSG00000011426.11 | 39965 | -0.707 | -0.1684 | Yes | ||
| 183 | MID1 | ENSG00000101871.14 | 40040 | -0.721 | -0.1601 | Yes | ||
| 184 | CENPE | ENSG00000138778.12 | 40145 | -0.740 | -0.1521 | Yes | ||
| 185 | PDLIM5 | ENSG00000163110.15 | 40169 | -0.746 | -0.1422 | Yes | ||
| 186 | RASAL2 | ENSG00000075391.16 | 40239 | -0.762 | -0.1331 | Yes | ||
| 187 | EPB41L2 | ENSG00000079819.19 | 40303 | -0.778 | -0.1237 | Yes | ||
| 188 | EZR | ENSG00000092820.18 | 40361 | -0.794 | -0.1138 | Yes | ||
| 189 | PCGF5 | ENSG00000180628.15 | 40443 | -0.821 | -0.1042 | Yes | ||
| 190 | KIF11 | ENSG00000138160.6 | 40519 | -0.845 | -0.0941 | Yes | ||
| 191 | ACTN4 | ENSG00000130402.12 | 40593 | -0.872 | -0.0836 | Yes | ||
| 192 | PKD2 | ENSG00000118762.8 | 40664 | -0.904 | -0.0725 | Yes | ||
| 193 | ECT2 | ENSG00000114346.13 | 40756 | -0.937 | -0.0615 | Yes | ||
| 194 | KIF5B | ENSG00000170759.11 | 40785 | -0.954 | -0.0487 | Yes | ||
| 195 | NET1 | ENSG00000173848.19 | 40812 | -0.977 | -0.0356 | Yes | ||
| 196 | RABGAP1 | ENSG00000011454.17 | 40846 | -0.994 | -0.0223 | Yes | ||
| 197 | KIF20B | ENSG00000138182.14 | 40873 | -1.017 | -0.0086 | Yes | ||
| 198 | FLNB | ENSG00000136068.14 | 40977 | -1.106 | 0.0045 | Yes |