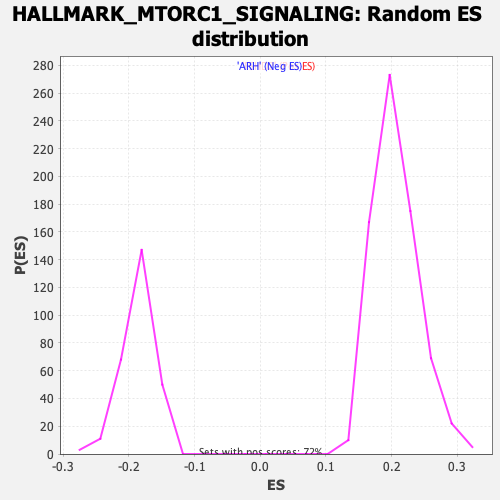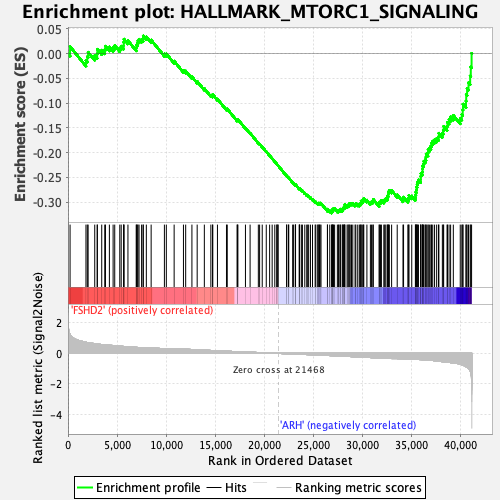
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_MTORC1_SIGNALING |
| Enrichment Score (ES) | -0.32249779 |
| Normalized Enrichment Score (NES) | -1.7364471 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0055784704 |
| FWER p-Value | 0.022 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | STC1 | ENSG00000159167.12 | 211 | 1.227 | 0.0141 | No | ||
| 2 | EGLN3 | ENSG00000129521.14 | 1831 | 0.727 | -0.0140 | No | ||
| 3 | FADS1 | ENSG00000149485.19 | 1968 | 0.709 | -0.0062 | No | ||
| 4 | DAPP1 | ENSG00000070190.13 | 2051 | 0.699 | 0.0028 | No | ||
| 5 | SLC6A6 | ENSG00000131389.17 | 2725 | 0.630 | -0.0037 | No | ||
| 6 | DHCR24 | ENSG00000116133.13 | 2975 | 0.608 | -0.0003 | No | ||
| 7 | BTG2 | ENSG00000159388.6 | 2993 | 0.606 | 0.0088 | No | ||
| 8 | FKBP2 | ENSG00000173486.13 | 3438 | 0.572 | 0.0070 | No | ||
| 9 | CFP | ENSG00000126759.13 | 3748 | 0.554 | 0.0081 | No | ||
| 10 | CXCR4 | ENSG00000121966.6 | 3819 | 0.548 | 0.0150 | No | ||
| 11 | DDIT4 | ENSG00000168209.5 | 4218 | 0.529 | 0.0136 | No | ||
| 12 | DHCR7 | ENSG00000172893.15 | 4598 | 0.514 | 0.0124 | No | ||
| 13 | NFIL3 | ENSG00000165030.4 | 4763 | 0.507 | 0.0164 | No | ||
| 14 | NAMPT | ENSG00000105835.12 | 5278 | 0.473 | 0.0112 | No | ||
| 15 | CD9 | ENSG00000010278.14 | 5422 | 0.464 | 0.0150 | No | ||
| 16 | SLC7A11 | ENSG00000151012.13 | 5674 | 0.449 | 0.0160 | No | ||
| 17 | HMGCR | ENSG00000113161.16 | 5676 | 0.449 | 0.0230 | No | ||
| 18 | FADS2 | ENSG00000134824.14 | 5709 | 0.447 | 0.0292 | No | ||
| 19 | G6PD | ENSG00000160211.19 | 6110 | 0.428 | 0.0262 | No | ||
| 20 | LDLR | ENSG00000130164.13 | 6959 | 0.391 | 0.0116 | No | ||
| 21 | EBP | ENSG00000147155.11 | 6973 | 0.391 | 0.0174 | No | ||
| 22 | PIK3R3 | ENSG00000117461.15 | 7066 | 0.387 | 0.0213 | No | ||
| 23 | SC5D | ENSG00000109929.10 | 7132 | 0.384 | 0.0257 | No | ||
| 24 | ASNS | ENSG00000070669.17 | 7241 | 0.378 | 0.0290 | No | ||
| 25 | SCD | ENSG00000099194.6 | 7501 | 0.371 | 0.0285 | No | ||
| 26 | TM7SF2 | ENSG00000149809.15 | 7658 | 0.364 | 0.0304 | No | ||
| 27 | GCLC | ENSG00000001084.13 | 7671 | 0.364 | 0.0358 | No | ||
| 28 | TXNRD1 | ENSG00000198431.16 | 7984 | 0.362 | 0.0339 | No | ||
| 29 | INSIG1 | ENSG00000186480.13 | 8475 | 0.352 | 0.0275 | No | ||
| 30 | TMEM97 | ENSG00000109084.14 | 9826 | 0.318 | -0.0005 | No | ||
| 31 | CYP51A1 | ENSG00000001630.17 | 10022 | 0.310 | -0.0004 | No | ||
| 32 | PHGDH | ENSG00000092621.12 | 10819 | 0.298 | -0.0152 | No | ||
| 33 | HMGCS1 | ENSG00000112972.15 | 11768 | 0.279 | -0.0339 | No | ||
| 34 | PFKL | ENSG00000141959.17 | 11995 | 0.271 | -0.0352 | No | ||
| 35 | GLA | ENSG00000102393.10 | 12631 | 0.250 | -0.0467 | No | ||
| 36 | IFRD1 | ENSG00000006652.14 | 13167 | 0.233 | -0.0561 | No | ||
| 37 | SLC37A4 | ENSG00000137700.18 | 13897 | 0.209 | -0.0707 | No | ||
| 38 | SHMT2 | ENSG00000182199.11 | 14562 | 0.188 | -0.0839 | No | ||
| 39 | AK4 | ENSG00000162433.15 | 14736 | 0.183 | -0.0853 | No | ||
| 40 | CORO1A | ENSG00000102879.15 | 14755 | 0.182 | -0.0829 | No | ||
| 41 | SQSTM1 | ENSG00000161011.20 | 15233 | 0.167 | -0.0919 | No | ||
| 42 | CTSC | ENSG00000109861.16 | 16154 | 0.141 | -0.1121 | No | ||
| 43 | SQLE | ENSG00000104549.12 | 16232 | 0.139 | -0.1118 | No | ||
| 44 | ACSL3 | ENSG00000123983.14 | 17241 | 0.111 | -0.1347 | No | ||
| 45 | CTH | ENSG00000116761.12 | 17265 | 0.110 | -0.1335 | No | ||
| 46 | ME1 | ENSG00000065833.9 | 17317 | 0.109 | -0.1331 | No | ||
| 47 | FDXR | ENSG00000161513.12 | 18088 | 0.088 | -0.1505 | No | ||
| 48 | ELOVL5 | ENSG00000012660.14 | 18561 | 0.077 | -0.1608 | No | ||
| 49 | PSMA3 | ENSG00000100567.13 | 19401 | 0.054 | -0.1804 | No | ||
| 50 | RDH11 | ENSG00000072042.13 | 19531 | 0.051 | -0.1828 | No | ||
| 51 | SLC2A1 | ENSG00000117394.23 | 19798 | 0.043 | -0.1886 | No | ||
| 52 | SSR1 | ENSG00000124783.14 | 20209 | 0.034 | -0.1981 | No | ||
| 53 | IDI1 | ENSG00000067064.11 | 20552 | 0.023 | -0.2060 | No | ||
| 54 | PSMC4 | ENSG00000013275.8 | 20810 | 0.018 | -0.2120 | No | ||
| 55 | RRP9 | ENSG00000114767.7 | 21079 | 0.009 | -0.2184 | No | ||
| 56 | TUBA4A | ENSG00000127824.14 | 21284 | 0.004 | -0.2234 | No | ||
| 57 | CDKN1A | ENSG00000124762.13 | 21297 | 0.004 | -0.2236 | No | ||
| 58 | DDIT3 | ENSG00000175197.12 | 21390 | 0.001 | -0.2258 | No | ||
| 59 | HK2 | ENSG00000159399.9 | 21414 | 0.001 | -0.2264 | No | ||
| 60 | NUPR1 | ENSG00000176046.8 | 22289 | -0.020 | -0.2474 | No | ||
| 61 | LGMN | ENSG00000100600.15 | 22291 | -0.020 | -0.2471 | No | ||
| 62 | ATP2A2 | ENSG00000174437.17 | 22480 | -0.026 | -0.2513 | No | ||
| 63 | TRIB3 | ENSG00000101255.11 | 22498 | -0.027 | -0.2512 | No | ||
| 64 | XBP1 | ENSG00000100219.16 | 22900 | -0.036 | -0.2605 | No | ||
| 65 | IDH1 | ENSG00000138413.13 | 22996 | -0.039 | -0.2622 | No | ||
| 66 | MTHFD2L | ENSG00000163738.18 | 23178 | -0.044 | -0.2659 | No | ||
| 67 | VLDLR | ENSG00000147852.16 | 23181 | -0.045 | -0.2652 | No | ||
| 68 | RPN1 | ENSG00000163902.12 | 23194 | -0.045 | -0.2648 | No | ||
| 69 | GLRX | ENSG00000173221.14 | 23563 | -0.054 | -0.2730 | No | ||
| 70 | COPS5 | ENSG00000121022.14 | 23597 | -0.055 | -0.2729 | No | ||
| 71 | GPI | ENSG00000105220.17 | 23611 | -0.056 | -0.2723 | No | ||
| 72 | SEC11A | ENSG00000140612.14 | 23787 | -0.060 | -0.2757 | No | ||
| 73 | GBE1 | ENSG00000114480.13 | 23894 | -0.063 | -0.2773 | No | ||
| 74 | ADD3 | ENSG00000148700.15 | 24136 | -0.068 | -0.2821 | No | ||
| 75 | CANX | ENSG00000127022.15 | 24334 | -0.074 | -0.2857 | No | ||
| 76 | YKT6 | ENSG00000106636.8 | 24433 | -0.075 | -0.2869 | No | ||
| 77 | ALDOA | ENSG00000149925.20 | 24530 | -0.078 | -0.2881 | No | ||
| 78 | LTA4H | ENSG00000111144.10 | 24714 | -0.083 | -0.2912 | No | ||
| 79 | PSPH | ENSG00000146733.14 | 24924 | -0.089 | -0.2949 | No | ||
| 80 | NFYC | ENSG00000066136.20 | 25192 | -0.096 | -0.3000 | No | ||
| 81 | PSAT1 | ENSG00000135069.14 | 25261 | -0.098 | -0.3001 | No | ||
| 82 | MTHFD2 | ENSG00000065911.12 | 25466 | -0.104 | -0.3034 | No | ||
| 83 | BHLHE40 | ENSG00000134107.5 | 25483 | -0.105 | -0.3022 | No | ||
| 84 | DDX39A | ENSG00000123136.15 | 25570 | -0.107 | -0.3026 | No | ||
| 85 | HSP90B1 | ENSG00000166598.15 | 25632 | -0.109 | -0.3024 | No | ||
| 86 | ARPC5L | ENSG00000136950.13 | 25696 | -0.111 | -0.3022 | No | ||
| 87 | SLC7A5 | ENSG00000103257.9 | 25788 | -0.113 | -0.3026 | No | ||
| 88 | BCAT1 | ENSG00000060982.15 | 26443 | -0.133 | -0.3165 | No | ||
| 89 | GAPDH | ENSG00000111640.15 | 26448 | -0.134 | -0.3145 | No | ||
| 90 | UNG | ENSG00000076248.10 | 26682 | -0.140 | -0.3180 | No | ||
| 91 | PDAP1 | ENSG00000106244.13 | 26869 | -0.145 | -0.3202 | Yes | ||
| 92 | GMPS | ENSG00000163655.16 | 26897 | -0.146 | -0.3186 | Yes | ||
| 93 | TPI1 | ENSG00000111669.15 | 26955 | -0.148 | -0.3176 | Yes | ||
| 94 | SLC1A4 | ENSG00000115902.11 | 26994 | -0.149 | -0.3162 | Yes | ||
| 95 | PPP1R15A | ENSG00000087074.8 | 27000 | -0.149 | -0.3140 | Yes | ||
| 96 | IGFBP5 | ENSG00000115461.5 | 27066 | -0.151 | -0.3132 | Yes | ||
| 97 | TFRC | ENSG00000072274.13 | 27176 | -0.154 | -0.3135 | Yes | ||
| 98 | SERPINH1 | ENSG00000149257.14 | 27457 | -0.163 | -0.3178 | Yes | ||
| 99 | SLC9A3R1 | ENSG00000109062.12 | 27565 | -0.167 | -0.3178 | Yes | ||
| 100 | PSMD13 | ENSG00000185627.18 | 27620 | -0.168 | -0.3164 | Yes | ||
| 101 | NFKBIB | ENSG00000104825.17 | 27749 | -0.172 | -0.3169 | Yes | ||
| 102 | CYB5B | ENSG00000103018.16 | 27762 | -0.172 | -0.3144 | Yes | ||
| 103 | ITGB2 | ENSG00000160255.18 | 27949 | -0.179 | -0.3162 | Yes | ||
| 104 | SLC1A5 | ENSG00000105281.12 | 28017 | -0.181 | -0.3150 | Yes | ||
| 105 | ACLY | ENSG00000131473.17 | 28035 | -0.182 | -0.3125 | Yes | ||
| 106 | PSMG1 | ENSG00000183527.12 | 28095 | -0.184 | -0.3111 | Yes | ||
| 107 | USO1 | ENSG00000138768.14 | 28192 | -0.186 | -0.3105 | Yes | ||
| 108 | NUFIP1 | ENSG00000083635.8 | 28207 | -0.186 | -0.3079 | Yes | ||
| 109 | GSR | ENSG00000104687.14 | 28212 | -0.187 | -0.3051 | Yes | ||
| 110 | PSMC6 | ENSG00000100519.12 | 28449 | -0.195 | -0.3078 | Yes | ||
| 111 | MLLT11 | ENSG00000213190.3 | 28545 | -0.198 | -0.3070 | Yes | ||
| 112 | ABCF2 | ENSG00000033050.9 | 28644 | -0.201 | -0.3063 | Yes | ||
| 113 | NMT1 | ENSG00000136448.12 | 28656 | -0.201 | -0.3034 | Yes | ||
| 114 | WARS | ENSG00000140105.18 | 28765 | -0.204 | -0.3028 | Yes | ||
| 115 | CALR | ENSG00000179218.13 | 28898 | -0.208 | -0.3028 | Yes | ||
| 116 | ELOVL6 | ENSG00000170522.9 | 29029 | -0.213 | -0.3026 | Yes | ||
| 117 | EEF1E1 | ENSG00000124802.12 | 29262 | -0.220 | -0.3048 | Yes | ||
| 118 | ETF1 | ENSG00000120705.13 | 29337 | -0.222 | -0.3031 | Yes | ||
| 119 | SDF2L1 | ENSG00000128228.5 | 29533 | -0.228 | -0.3043 | Yes | ||
| 120 | EDEM1 | ENSG00000134109.11 | 29697 | -0.234 | -0.3046 | Yes | ||
| 121 | CCT6A | ENSG00000146731.11 | 29797 | -0.238 | -0.3033 | Yes | ||
| 122 | UBE2D3 | ENSG00000109332.20 | 29865 | -0.240 | -0.3011 | Yes | ||
| 123 | GGA2 | ENSG00000103365.15 | 29882 | -0.241 | -0.2978 | Yes | ||
| 124 | ADIPOR2 | ENSG00000006831.10 | 30043 | -0.245 | -0.2978 | Yes | ||
| 125 | PSMA4 | ENSG00000041357.16 | 30107 | -0.247 | -0.2955 | Yes | ||
| 126 | RAB1A | ENSG00000138069.18 | 30150 | -0.249 | -0.2926 | Yes | ||
| 127 | PGM1 | ENSG00000079739.17 | 30468 | -0.259 | -0.2963 | Yes | ||
| 128 | ACACA | ENSG00000278540.5 | 30823 | -0.270 | -0.3007 | Yes | ||
| 129 | CCNG1 | ENSG00000113328.19 | 30943 | -0.275 | -0.2992 | Yes | ||
| 130 | TOMM40 | ENSG00000130204.13 | 31075 | -0.280 | -0.2980 | Yes | ||
| 131 | RPA1 | ENSG00000132383.12 | 31119 | -0.282 | -0.2947 | Yes | ||
| 132 | UCHL5 | ENSG00000116750.13 | 31722 | -0.286 | -0.3049 | Yes | ||
| 133 | SORD | ENSG00000140263.14 | 31730 | -0.287 | -0.3005 | Yes | ||
| 134 | M6PR | ENSG00000003056.8 | 31850 | -0.291 | -0.2989 | Yes | ||
| 135 | ERO1A | ENSG00000197930.13 | 31946 | -0.294 | -0.2966 | Yes | ||
| 136 | GOT1 | ENSG00000120053.12 | 32193 | -0.304 | -0.2978 | Yes | ||
| 137 | HSPA5 | ENSG00000044574.8 | 32282 | -0.308 | -0.2951 | Yes | ||
| 138 | STARD4 | ENSG00000164211.13 | 32393 | -0.313 | -0.2929 | Yes | ||
| 139 | MCM2 | ENSG00000073111.14 | 32546 | -0.318 | -0.2916 | Yes | ||
| 140 | CACYBP | ENSG00000116161.18 | 32580 | -0.319 | -0.2874 | Yes | ||
| 141 | FGL2 | ENSG00000127951.7 | 32655 | -0.323 | -0.2841 | Yes | ||
| 142 | SERP1 | ENSG00000120742.11 | 32659 | -0.323 | -0.2791 | Yes | ||
| 143 | PSMD12 | ENSG00000197170.10 | 32756 | -0.326 | -0.2764 | Yes | ||
| 144 | EIF2S2 | ENSG00000125977.7 | 32987 | -0.335 | -0.2767 | Yes | ||
| 145 | IMMT | ENSG00000132305.20 | 33557 | -0.338 | -0.2853 | Yes | ||
| 146 | UFM1 | ENSG00000120686.12 | 34146 | -0.347 | -0.2942 | Yes | ||
| 147 | HSPD1 | ENSG00000144381.17 | 34204 | -0.350 | -0.2901 | Yes | ||
| 148 | TCEA1 | ENSG00000187735.14 | 34643 | -0.353 | -0.2952 | Yes | ||
| 149 | SKAP2 | ENSG00000005020.13 | 34726 | -0.357 | -0.2916 | Yes | ||
| 150 | HSPA9 | ENSG00000113013.15 | 34748 | -0.358 | -0.2865 | Yes | ||
| 151 | PSMB5 | ENSG00000100804.19 | 35039 | -0.367 | -0.2878 | Yes | ||
| 152 | ATP5MC1 | ENSG00000159199.14 | 35409 | -0.384 | -0.2908 | Yes | ||
| 153 | PSME3 | ENSG00000131467.11 | 35430 | -0.385 | -0.2853 | Yes | ||
| 154 | ATP6V1D | ENSG00000100554.12 | 35460 | -0.386 | -0.2799 | Yes | ||
| 155 | CCNF | ENSG00000162063.13 | 35514 | -0.389 | -0.2751 | Yes | ||
| 156 | POLR3G | ENSG00000113356.12 | 35541 | -0.390 | -0.2696 | Yes | ||
| 157 | PDK1 | ENSG00000152256.13 | 35589 | -0.393 | -0.2646 | Yes | ||
| 158 | ENO1 | ENSG00000074800.16 | 35638 | -0.395 | -0.2595 | Yes | ||
| 159 | HMBS | ENSG00000256269.10 | 35722 | -0.399 | -0.2553 | Yes | ||
| 160 | PSMC2 | ENSG00000161057.12 | 35948 | -0.403 | -0.2545 | Yes | ||
| 161 | HSPE1 | ENSG00000115541.11 | 35951 | -0.403 | -0.2482 | Yes | ||
| 162 | STIP1 | ENSG00000168439.16 | 35973 | -0.404 | -0.2423 | Yes | ||
| 163 | EPRS | ENSG00000136628.18 | 36119 | -0.410 | -0.2394 | Yes | ||
| 164 | PNP | ENSG00000198805.11 | 36135 | -0.411 | -0.2334 | Yes | ||
| 165 | PRDX1 | ENSG00000117450.14 | 36141 | -0.411 | -0.2270 | Yes | ||
| 166 | TBK1 | ENSG00000183735.10 | 36229 | -0.412 | -0.2227 | Yes | ||
| 167 | HSPA4 | ENSG00000170606.15 | 36280 | -0.415 | -0.2174 | Yes | ||
| 168 | PSMD14 | ENSG00000115233.12 | 36419 | -0.422 | -0.2141 | Yes | ||
| 169 | NUP205 | ENSG00000155561.15 | 36490 | -0.426 | -0.2092 | Yes | ||
| 170 | PPA1 | ENSG00000180817.12 | 36510 | -0.427 | -0.2029 | Yes | ||
| 171 | QDPR | ENSG00000151552.12 | 36694 | -0.435 | -0.2005 | Yes | ||
| 172 | PLK1 | ENSG00000166851.15 | 36707 | -0.436 | -0.1940 | Yes | ||
| 173 | HPRT1 | ENSG00000165704.15 | 36862 | -0.444 | -0.1908 | Yes | ||
| 174 | PITPNB | ENSG00000180957.18 | 36991 | -0.452 | -0.1868 | Yes | ||
| 175 | PPIA | ENSG00000196262.14 | 37055 | -0.456 | -0.1812 | Yes | ||
| 176 | ACTR3 | ENSG00000115091.12 | 37178 | -0.462 | -0.1769 | Yes | ||
| 177 | P4HA1 | ENSG00000122884.12 | 37364 | -0.475 | -0.1740 | Yes | ||
| 178 | GSK3B | ENSG00000082701.15 | 37577 | -0.491 | -0.1714 | Yes | ||
| 179 | BUB1 | ENSG00000169679.15 | 37763 | -0.508 | -0.1680 | Yes | ||
| 180 | PGK1 | ENSG00000102144.15 | 37804 | -0.512 | -0.1609 | Yes | ||
| 181 | ACTR2 | ENSG00000138071.13 | 38158 | -0.539 | -0.1611 | Yes | ||
| 182 | RIT1 | ENSG00000143622.11 | 38248 | -0.546 | -0.1547 | Yes | ||
| 183 | TUBG1 | ENSG00000131462.8 | 38291 | -0.550 | -0.1470 | Yes | ||
| 184 | AURKA | ENSG00000087586.17 | 38638 | -0.565 | -0.1466 | Yes | ||
| 185 | SRD5A1 | ENSG00000145545.12 | 38684 | -0.569 | -0.1388 | Yes | ||
| 186 | MAP2K3 | ENSG00000034152.19 | 38842 | -0.584 | -0.1334 | Yes | ||
| 187 | PNO1 | ENSG00000115946.8 | 39000 | -0.601 | -0.1278 | Yes | ||
| 188 | IFI30 | ENSG00000216490.4 | 39276 | -0.621 | -0.1248 | Yes | ||
| 189 | RRM2 | ENSG00000171848.15 | 39983 | -0.710 | -0.1309 | Yes | ||
| 190 | GTF2H1 | ENSG00000110768.12 | 40144 | -0.740 | -0.1232 | Yes | ||
| 191 | SLA | ENSG00000155926.14 | 40233 | -0.760 | -0.1134 | Yes | ||
| 192 | CDC25A | ENSG00000164045.12 | 40263 | -0.768 | -0.1020 | Yes | ||
| 193 | SLC2A3 | ENSG00000059804.16 | 40569 | -0.866 | -0.0959 | Yes | ||
| 194 | MCM4 | ENSG00000104738.18 | 40591 | -0.872 | -0.0827 | Yes | ||
| 195 | LDHA | ENSG00000134333.14 | 40682 | -0.914 | -0.0706 | Yes | ||
| 196 | DHFR | ENSG00000228716.7 | 40826 | -0.981 | -0.0587 | Yes | ||
| 197 | TES | ENSG00000135269.18 | 40994 | -1.130 | -0.0450 | Yes | ||
| 198 | PLOD2 | ENSG00000152952.12 | 41057 | -1.247 | -0.0269 | Yes | ||
| 199 | SYTL2 | ENSG00000137501.17 | 41138 | -1.875 | 0.0006 | Yes |