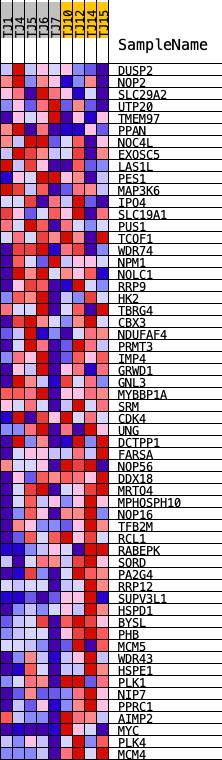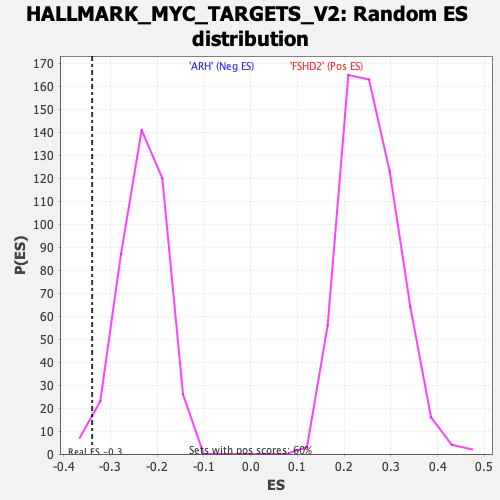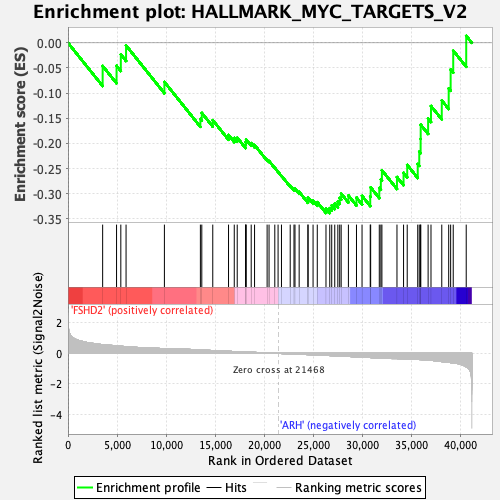
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_MYC_TARGETS_V2 |
| Enrichment Score (ES) | -0.3387217 |
| Normalized Enrichment Score (NES) | -1.4569099 |
| Nominal p-value | 0.01980198 |
| FDR q-value | 0.031148275 |
| FWER p-Value | 0.21 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DUSP2 | ENSG00000158050.5 | 3532 | 0.565 | -0.0460 | No | ||
| 2 | NOP2 | ENSG00000111641.11 | 4949 | 0.494 | -0.0455 | No | ||
| 3 | SLC29A2 | ENSG00000174669.12 | 5384 | 0.466 | -0.0232 | No | ||
| 4 | UTP20 | ENSG00000120800.5 | 5918 | 0.437 | -0.0052 | No | ||
| 5 | TMEM97 | ENSG00000109084.14 | 9826 | 0.318 | -0.0778 | No | ||
| 6 | PPAN | ENSG00000130810.20 | 13494 | 0.222 | -0.1514 | No | ||
| 7 | NOC4L | ENSG00000184967.7 | 13638 | 0.218 | -0.1394 | No | ||
| 8 | EXOSC5 | ENSG00000077348.9 | 14756 | 0.182 | -0.1538 | No | ||
| 9 | LAS1L | ENSG00000001497.16 | 16363 | 0.135 | -0.1833 | No | ||
| 10 | PES1 | ENSG00000100029.18 | 16947 | 0.118 | -0.1891 | No | ||
| 11 | MAP3K6 | ENSG00000142733.17 | 17249 | 0.111 | -0.1886 | No | ||
| 12 | IPO4 | ENSG00000196497.17 | 18103 | 0.088 | -0.2032 | No | ||
| 13 | SLC19A1 | ENSG00000173638.19 | 18117 | 0.088 | -0.1973 | No | ||
| 14 | PUS1 | ENSG00000177192.14 | 18161 | 0.087 | -0.1922 | No | ||
| 15 | TCOF1 | ENSG00000070814.20 | 18674 | 0.074 | -0.1995 | No | ||
| 16 | WDR74 | ENSG00000133316.15 | 19016 | 0.065 | -0.2032 | No | ||
| 17 | NPM1 | ENSG00000181163.13 | 20296 | 0.031 | -0.2321 | No | ||
| 18 | NOLC1 | ENSG00000166197.16 | 20493 | 0.025 | -0.2351 | No | ||
| 19 | RRP9 | ENSG00000114767.7 | 21079 | 0.009 | -0.2487 | No | ||
| 20 | HK2 | ENSG00000159399.9 | 21414 | 0.001 | -0.2567 | No | ||
| 21 | TBRG4 | ENSG00000136270.14 | 21759 | -0.008 | -0.2645 | No | ||
| 22 | CBX3 | ENSG00000122565.19 | 22650 | -0.030 | -0.2841 | No | ||
| 23 | NDUFAF4 | ENSG00000123545.6 | 23035 | -0.040 | -0.2906 | No | ||
| 24 | PRMT3 | ENSG00000185238.13 | 23137 | -0.043 | -0.2900 | No | ||
| 25 | IMP4 | ENSG00000136718.9 | 23562 | -0.054 | -0.2965 | No | ||
| 26 | GRWD1 | ENSG00000105447.13 | 24457 | -0.076 | -0.3128 | No | ||
| 27 | GNL3 | ENSG00000163938.17 | 24468 | -0.076 | -0.3077 | No | ||
| 28 | MYBBP1A | ENSG00000132382.14 | 24980 | -0.091 | -0.3137 | No | ||
| 29 | SRM | ENSG00000116649.10 | 25409 | -0.102 | -0.3169 | No | ||
| 30 | CDK4 | ENSG00000135446.17 | 26296 | -0.129 | -0.3294 | Yes | ||
| 31 | UNG | ENSG00000076248.10 | 26682 | -0.140 | -0.3288 | Yes | ||
| 32 | DCTPP1 | ENSG00000179958.9 | 26881 | -0.146 | -0.3233 | Yes | ||
| 33 | FARSA | ENSG00000179115.11 | 27190 | -0.155 | -0.3199 | Yes | ||
| 34 | NOP56 | ENSG00000101361.17 | 27506 | -0.165 | -0.3159 | Yes | ||
| 35 | DDX18 | ENSG00000088205.13 | 27696 | -0.170 | -0.3085 | Yes | ||
| 36 | MRTO4 | ENSG00000053372.5 | 27844 | -0.175 | -0.2997 | Yes | ||
| 37 | MPHOSPH10 | ENSG00000124383.9 | 28582 | -0.199 | -0.3036 | Yes | ||
| 38 | NOP16 | ENSG00000048162.20 | 29405 | -0.225 | -0.3077 | Yes | ||
| 39 | TFB2M | ENSG00000162851.8 | 29972 | -0.243 | -0.3044 | Yes | ||
| 40 | RCL1 | ENSG00000120158.12 | 30807 | -0.270 | -0.3056 | Yes | ||
| 41 | RABEPK | ENSG00000136933.17 | 30851 | -0.271 | -0.2875 | Yes | ||
| 42 | SORD | ENSG00000140263.14 | 31730 | -0.287 | -0.2886 | Yes | ||
| 43 | PA2G4 | ENSG00000170515.14 | 31890 | -0.293 | -0.2718 | Yes | ||
| 44 | RRP12 | ENSG00000052749.13 | 31997 | -0.296 | -0.2534 | Yes | ||
| 45 | SUPV3L1 | ENSG00000156502.14 | 33532 | -0.337 | -0.2669 | Yes | ||
| 46 | HSPD1 | ENSG00000144381.17 | 34204 | -0.350 | -0.2585 | Yes | ||
| 47 | BYSL | ENSG00000112578.10 | 34577 | -0.350 | -0.2429 | Yes | ||
| 48 | PHB | ENSG00000167085.11 | 35649 | -0.396 | -0.2410 | Yes | ||
| 49 | MCM5 | ENSG00000100297.16 | 35805 | -0.402 | -0.2163 | Yes | ||
| 50 | WDR43 | ENSG00000163811.12 | 35944 | -0.403 | -0.1912 | Yes | ||
| 51 | HSPE1 | ENSG00000115541.11 | 35951 | -0.403 | -0.1629 | Yes | ||
| 52 | PLK1 | ENSG00000166851.15 | 36707 | -0.436 | -0.1505 | Yes | ||
| 53 | NIP7 | ENSG00000132603.15 | 37000 | -0.452 | -0.1256 | Yes | ||
| 54 | PPRC1 | ENSG00000148840.11 | 38099 | -0.535 | -0.1146 | Yes | ||
| 55 | AIMP2 | ENSG00000106305.10 | 38805 | -0.581 | -0.0907 | Yes | ||
| 56 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.0529 | Yes | ||
| 57 | PLK4 | ENSG00000142731.11 | 39267 | -0.620 | -0.0155 | Yes | ||
| 58 | MCM4 | ENSG00000104738.18 | 40591 | -0.872 | 0.0139 | Yes |