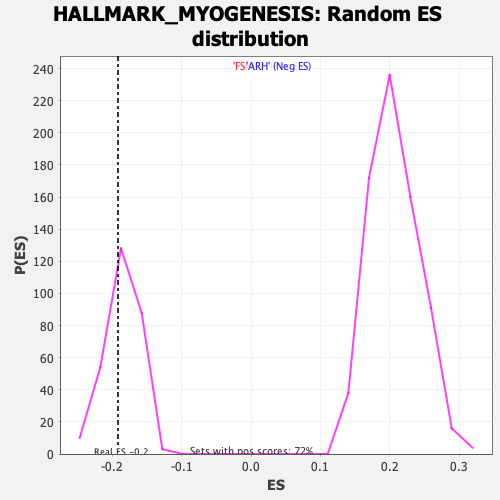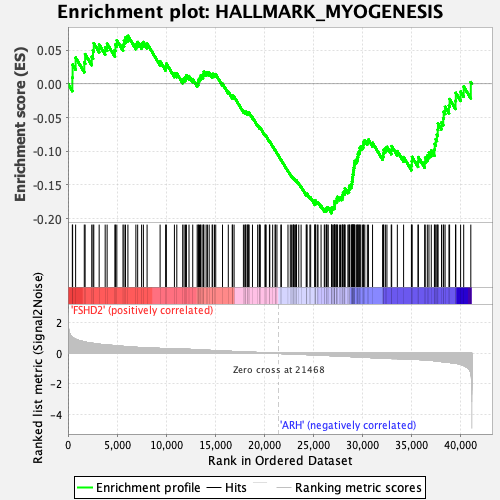
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_MYOGENESIS |
| Enrichment Score (ES) | -0.19178501 |
| Normalized Enrichment Score (NES) | -1.029546 |
| Nominal p-value | 0.36749116 |
| FDR q-value | 0.42255688 |
| FWER p-Value | 0.998 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PTP4A3 | ENSG00000184489.11 | 442 | 1.039 | 0.0093 | No | ||
| 2 | CAMK2B | ENSG00000058404.20 | 471 | 1.027 | 0.0284 | No | ||
| 3 | EFS | ENSG00000100842.13 | 777 | 0.918 | 0.0387 | No | ||
| 4 | CLU | ENSG00000120885.22 | 1659 | 0.752 | 0.0317 | No | ||
| 5 | ACTN3 | ENSG00000248746.6 | 1745 | 0.740 | 0.0440 | No | ||
| 6 | IGFBP3 | ENSG00000146674.15 | 2420 | 0.661 | 0.0403 | No | ||
| 7 | LSP1 | ENSG00000130592.15 | 2548 | 0.649 | 0.0497 | No | ||
| 8 | PTGIS | ENSG00000124212.6 | 2629 | 0.640 | 0.0601 | No | ||
| 9 | CKMT2 | ENSG00000131730.16 | 3176 | 0.593 | 0.0582 | No | ||
| 10 | TCAP | ENSG00000173991.5 | 3789 | 0.550 | 0.0539 | No | ||
| 11 | MAPK12 | ENSG00000188130.14 | 3988 | 0.540 | 0.0595 | No | ||
| 12 | SGCG | ENSG00000102683.7 | 4781 | 0.506 | 0.0500 | No | ||
| 13 | VIPR1 | ENSG00000114812.13 | 4835 | 0.501 | 0.0584 | No | ||
| 14 | PICK1 | ENSG00000100151.16 | 4977 | 0.492 | 0.0644 | No | ||
| 15 | SH2B1 | ENSG00000178188.14 | 5606 | 0.453 | 0.0578 | No | ||
| 16 | ABLIM1 | ENSG00000099204.20 | 5712 | 0.447 | 0.0639 | No | ||
| 17 | ACHE | ENSG00000087085.15 | 5861 | 0.441 | 0.0688 | No | ||
| 18 | NQO1 | ENSG00000181019.13 | 6107 | 0.428 | 0.0711 | No | ||
| 19 | CFD | ENSG00000197766.8 | 6918 | 0.393 | 0.0589 | No | ||
| 20 | PSEN2 | ENSG00000143801.17 | 7101 | 0.385 | 0.0619 | No | ||
| 21 | SCD | ENSG00000099194.6 | 7501 | 0.371 | 0.0593 | No | ||
| 22 | SGCA | ENSG00000108823.16 | 7684 | 0.363 | 0.0619 | No | ||
| 23 | GNAO1 | ENSG00000087258.15 | 8062 | 0.359 | 0.0596 | No | ||
| 24 | MYL7 | ENSG00000106631.8 | 9388 | 0.320 | 0.0335 | No | ||
| 25 | SLC6A8 | ENSG00000130821.16 | 9951 | 0.313 | 0.0258 | No | ||
| 26 | FABP3 | ENSG00000121769.8 | 10026 | 0.310 | 0.0300 | No | ||
| 27 | LPIN1 | ENSG00000134324.11 | 10847 | 0.297 | 0.0157 | No | ||
| 28 | EPHB3 | ENSG00000182580.3 | 11091 | 0.288 | 0.0153 | No | ||
| 29 | TSC2 | ENSG00000103197.18 | 11691 | 0.283 | 0.0062 | No | ||
| 30 | KCNH1 | ENSG00000143473.13 | 11832 | 0.277 | 0.0081 | No | ||
| 31 | CACNA1H | ENSG00000196557.13 | 11974 | 0.272 | 0.0099 | No | ||
| 32 | COL6A3 | ENSG00000163359.15 | 12073 | 0.268 | 0.0127 | No | ||
| 33 | MYOM2 | ENSG00000036448.10 | 12334 | 0.259 | 0.0114 | No | ||
| 34 | APLNR | ENSG00000134817.10 | 12714 | 0.247 | 0.0069 | No | ||
| 35 | IFRD1 | ENSG00000006652.14 | 13167 | 0.233 | 0.0004 | No | ||
| 36 | FDPS | ENSG00000160752.14 | 13271 | 0.229 | 0.0023 | No | ||
| 37 | OCEL1 | ENSG00000099330.9 | 13295 | 0.229 | 0.0061 | No | ||
| 38 | IGF1 | ENSG00000017427.16 | 13404 | 0.225 | 0.0078 | No | ||
| 39 | APOD | ENSG00000189058.9 | 13505 | 0.221 | 0.0097 | No | ||
| 40 | PPFIA4 | ENSG00000143847.15 | 13557 | 0.220 | 0.0127 | No | ||
| 41 | ITGB5 | ENSG00000082781.12 | 13753 | 0.214 | 0.0121 | No | ||
| 42 | KCNH2 | ENSG00000055118.15 | 13802 | 0.213 | 0.0150 | No | ||
| 43 | LARGE1 | ENSG00000133424.20 | 13853 | 0.211 | 0.0179 | No | ||
| 44 | TNNC2 | ENSG00000101470.10 | 14080 | 0.203 | 0.0163 | No | ||
| 45 | PLXNB2 | ENSG00000196576.15 | 14211 | 0.199 | 0.0169 | No | ||
| 46 | WWTR1 | ENSG00000018408.14 | 14391 | 0.194 | 0.0163 | No | ||
| 47 | ST5 | ENSG00000166444.19 | 14692 | 0.184 | 0.0126 | No | ||
| 48 | LAMA2 | ENSG00000196569.12 | 14748 | 0.182 | 0.0147 | No | ||
| 49 | COL6A2 | ENSG00000142173.15 | 14929 | 0.176 | 0.0137 | No | ||
| 50 | SH3BGR | ENSG00000185437.13 | 15059 | 0.172 | 0.0139 | No | ||
| 51 | ITGB4 | ENSG00000132470.14 | 15750 | 0.152 | 0.0000 | No | ||
| 52 | MAPRE3 | ENSG00000084764.12 | 16339 | 0.136 | -0.0117 | No | ||
| 53 | SYNGR2 | ENSG00000108639.7 | 16740 | 0.124 | -0.0191 | No | ||
| 54 | FOXO4 | ENSG00000184481.17 | 16779 | 0.123 | -0.0176 | No | ||
| 55 | FXYD1 | ENSG00000266964.5 | 16930 | 0.118 | -0.0190 | No | ||
| 56 | NAV2 | ENSG00000166833.21 | 17879 | 0.094 | -0.0404 | No | ||
| 57 | AGRN | ENSG00000188157.15 | 18021 | 0.090 | -0.0421 | No | ||
| 58 | AKT2 | ENSG00000105221.18 | 18110 | 0.088 | -0.0425 | No | ||
| 59 | ENO3 | ENSG00000108515.17 | 18133 | 0.087 | -0.0414 | No | ||
| 60 | ADAM12 | ENSG00000148848.14 | 18322 | 0.083 | -0.0443 | No | ||
| 61 | MYL3 | ENSG00000160808.10 | 18332 | 0.083 | -0.0430 | No | ||
| 62 | HSPB8 | ENSG00000152137.6 | 18377 | 0.082 | -0.0425 | No | ||
| 63 | GPX3 | ENSG00000211445.12 | 18462 | 0.080 | -0.0430 | No | ||
| 64 | SORBS3 | ENSG00000120896.13 | 18804 | 0.070 | -0.0499 | No | ||
| 65 | GAA | ENSG00000171298.13 | 19333 | 0.055 | -0.0617 | No | ||
| 66 | BDKRB2 | ENSG00000168398.6 | 19500 | 0.051 | -0.0648 | No | ||
| 67 | ACSL1 | ENSG00000151726.14 | 19506 | 0.051 | -0.0639 | No | ||
| 68 | STC2 | ENSG00000113739.10 | 19602 | 0.048 | -0.0653 | No | ||
| 69 | PKIA | ENSG00000171033.13 | 20076 | 0.038 | -0.0761 | No | ||
| 70 | MYOM1 | ENSG00000101605.12 | 20128 | 0.036 | -0.0767 | No | ||
| 71 | EIF4A2 | ENSG00000156976.17 | 20197 | 0.034 | -0.0777 | No | ||
| 72 | ITGA7 | ENSG00000135424.16 | 20532 | 0.024 | -0.0854 | No | ||
| 73 | CHRNB1 | ENSG00000170175.11 | 20569 | 0.023 | -0.0858 | No | ||
| 74 | SLN | ENSG00000170290.4 | 20861 | 0.016 | -0.0926 | No | ||
| 75 | MEF2D | ENSG00000116604.18 | 21115 | 0.008 | -0.0986 | No | ||
| 76 | MYL6B | ENSG00000196465.10 | 21130 | 0.007 | -0.0988 | No | ||
| 77 | CDKN1A | ENSG00000124762.13 | 21297 | 0.004 | -0.1028 | No | ||
| 78 | AGL | ENSG00000162688.17 | 21699 | -0.006 | -0.1125 | No | ||
| 79 | DTNA | ENSG00000134769.21 | 21748 | -0.008 | -0.1135 | No | ||
| 80 | SIRT2 | ENSG00000068903.20 | 22423 | -0.024 | -0.1295 | No | ||
| 81 | ADCY9 | ENSG00000162104.10 | 22658 | -0.030 | -0.1346 | No | ||
| 82 | NOTCH1 | ENSG00000148400.12 | 22768 | -0.032 | -0.1367 | No | ||
| 83 | MEF2C | ENSG00000081189.15 | 22923 | -0.037 | -0.1397 | No | ||
| 84 | COX6A2 | ENSG00000156885.6 | 22979 | -0.039 | -0.1403 | No | ||
| 85 | ERBB3 | ENSG00000065361.15 | 22995 | -0.039 | -0.1399 | No | ||
| 86 | ATP2A1 | ENSG00000196296.13 | 23109 | -0.042 | -0.1418 | No | ||
| 87 | COL1A1 | ENSG00000108821.13 | 23206 | -0.045 | -0.1433 | No | ||
| 88 | KLF5 | ENSG00000102554.14 | 23286 | -0.046 | -0.1443 | No | ||
| 89 | COX7A1 | ENSG00000161281.11 | 23295 | -0.047 | -0.1436 | No | ||
| 90 | PFKM | ENSG00000152556.17 | 23521 | -0.053 | -0.1481 | No | ||
| 91 | CHRNG | ENSG00000196811.13 | 23760 | -0.059 | -0.1528 | No | ||
| 92 | BAG1 | ENSG00000107262.23 | 24279 | -0.073 | -0.1640 | No | ||
| 93 | MYH4 | ENSG00000264424.1 | 24340 | -0.074 | -0.1641 | No | ||
| 94 | SPEG | ENSG00000072195.15 | 24357 | -0.075 | -0.1630 | No | ||
| 95 | CAV3 | ENSG00000182533.6 | 24657 | -0.081 | -0.1687 | No | ||
| 96 | COL3A1 | ENSG00000168542.14 | 24737 | -0.084 | -0.1690 | No | ||
| 97 | BIN1 | ENSG00000136717.15 | 25170 | -0.095 | -0.1777 | No | ||
| 98 | SPDEF | ENSG00000124664.11 | 25184 | -0.096 | -0.1762 | No | ||
| 99 | HSPB2 | ENSG00000170276.6 | 25193 | -0.096 | -0.1746 | No | ||
| 100 | RYR1 | ENSG00000196218.12 | 25194 | -0.096 | -0.1727 | No | ||
| 101 | TEAD4 | ENSG00000197905.9 | 25401 | -0.102 | -0.1758 | No | ||
| 102 | BHLHE40 | ENSG00000134107.5 | 25483 | -0.105 | -0.1757 | No | ||
| 103 | PYGM | ENSG00000068976.14 | 25795 | -0.113 | -0.1811 | No | ||
| 104 | MEF2A | ENSG00000068305.17 | 26124 | -0.124 | -0.1867 | No | ||
| 105 | DMD | ENSG00000198947.15 | 26181 | -0.125 | -0.1857 | No | ||
| 106 | DMPK | ENSG00000104936.17 | 26330 | -0.130 | -0.1868 | No | ||
| 107 | COL4A2 | ENSG00000134871.19 | 26354 | -0.131 | -0.1848 | No | ||
| 108 | MYH1 | ENSG00000109061.10 | 26406 | -0.132 | -0.1835 | No | ||
| 109 | PVALB | ENSG00000100362.13 | 26525 | -0.136 | -0.1838 | No | ||
| 110 | TGFB1 | ENSG00000105329.10 | 26855 | -0.145 | -0.1890 | Yes | ||
| 111 | TPD52L1 | ENSG00000111907.21 | 26861 | -0.145 | -0.1863 | Yes | ||
| 112 | PDE4DIP | ENSG00000178104.19 | 26870 | -0.145 | -0.1837 | Yes | ||
| 113 | CKM | ENSG00000104879.5 | 26984 | -0.149 | -0.1836 | Yes | ||
| 114 | NOS1 | ENSG00000089250.19 | 27089 | -0.152 | -0.1832 | Yes | ||
| 115 | HRC | ENSG00000130528.12 | 27175 | -0.154 | -0.1823 | Yes | ||
| 116 | TNNT1 | ENSG00000105048.17 | 27178 | -0.154 | -0.1793 | Yes | ||
| 117 | GJA5 | ENSG00000265107.3 | 27185 | -0.155 | -0.1765 | Yes | ||
| 118 | GABARAPL2 | ENSG00000034713.8 | 27201 | -0.155 | -0.1739 | Yes | ||
| 119 | ATP6AP1 | ENSG00000071553.17 | 27324 | -0.159 | -0.1738 | Yes | ||
| 120 | COL15A1 | ENSG00000204291.11 | 27409 | -0.161 | -0.1727 | Yes | ||
| 121 | MYH2 | ENSG00000125414.19 | 27417 | -0.161 | -0.1698 | Yes | ||
| 122 | MYLPF | ENSG00000180209.12 | 27454 | -0.163 | -0.1675 | Yes | ||
| 123 | CTF1 | ENSG00000150281.6 | 27662 | -0.169 | -0.1693 | Yes | ||
| 124 | SCHIP1 | ENSG00000151967.18 | 27775 | -0.173 | -0.1687 | Yes | ||
| 125 | MYH3 | ENSG00000109063.15 | 27927 | -0.177 | -0.1690 | Yes | ||
| 126 | CACNG1 | ENSG00000108878.5 | 27982 | -0.180 | -0.1668 | Yes | ||
| 127 | SPARC | ENSG00000113140.11 | 28014 | -0.181 | -0.1641 | Yes | ||
| 128 | MYL2 | ENSG00000111245.15 | 28045 | -0.182 | -0.1613 | Yes | ||
| 129 | MYH8 | ENSG00000133020.4 | 28178 | -0.185 | -0.1609 | Yes | ||
| 130 | MYBPH | ENSG00000133055.9 | 28219 | -0.187 | -0.1583 | Yes | ||
| 131 | HDAC5 | ENSG00000108840.15 | 28234 | -0.187 | -0.1550 | Yes | ||
| 132 | GSN | ENSG00000148180.19 | 28530 | -0.197 | -0.1584 | Yes | ||
| 133 | ACTA1 | ENSG00000143632.14 | 28635 | -0.200 | -0.1571 | Yes | ||
| 134 | REEP1 | ENSG00000068615.18 | 28658 | -0.201 | -0.1537 | Yes | ||
| 135 | PRNP | ENSG00000171867.17 | 28702 | -0.202 | -0.1509 | Yes | ||
| 136 | SOD3 | ENSG00000109610.6 | 28863 | -0.207 | -0.1508 | Yes | ||
| 137 | AK1 | ENSG00000106992.19 | 28916 | -0.209 | -0.1480 | Yes | ||
| 138 | CASQ1 | ENSG00000143318.13 | 28953 | -0.210 | -0.1448 | Yes | ||
| 139 | TNNI1 | ENSG00000159173.19 | 28979 | -0.211 | -0.1413 | Yes | ||
| 140 | FLII | ENSG00000177731.16 | 29008 | -0.212 | -0.1379 | Yes | ||
| 141 | CRAT | ENSG00000095321.17 | 29033 | -0.213 | -0.1344 | Yes | ||
| 142 | LDB3 | ENSG00000122367.19 | 29081 | -0.215 | -0.1314 | Yes | ||
| 143 | APP | ENSG00000142192.21 | 29094 | -0.215 | -0.1275 | Yes | ||
| 144 | PDLIM7 | ENSG00000196923.14 | 29135 | -0.216 | -0.1244 | Yes | ||
| 145 | MYH7 | ENSG00000092054.13 | 29177 | -0.217 | -0.1212 | Yes | ||
| 146 | KIFC3 | ENSG00000140859.16 | 29188 | -0.217 | -0.1172 | Yes | ||
| 147 | TNNT2 | ENSG00000118194.20 | 29241 | -0.219 | -0.1143 | Yes | ||
| 148 | MB | ENSG00000198125.13 | 29390 | -0.224 | -0.1135 | Yes | ||
| 149 | PPP1R3C | ENSG00000119938.9 | 29469 | -0.226 | -0.1111 | Yes | ||
| 150 | MYL4 | ENSG00000198336.9 | 29531 | -0.228 | -0.1082 | Yes | ||
| 151 | TNNT3 | ENSG00000130595.19 | 29558 | -0.229 | -0.1044 | Yes | ||
| 152 | SPTAN1 | ENSG00000197694.16 | 29628 | -0.232 | -0.1016 | Yes | ||
| 153 | DES | ENSG00000175084.12 | 29731 | -0.235 | -0.0995 | Yes | ||
| 154 | ACTN2 | ENSG00000077522.13 | 29748 | -0.236 | -0.0954 | Yes | ||
| 155 | NCAM1 | ENSG00000149294.16 | 29840 | -0.239 | -0.0930 | Yes | ||
| 156 | PC | ENSG00000173599.15 | 30008 | -0.244 | -0.0923 | Yes | ||
| 157 | TNNC1 | ENSG00000114854.8 | 30104 | -0.247 | -0.0899 | Yes | ||
| 158 | MYOG | ENSG00000122180.5 | 30136 | -0.248 | -0.0858 | Yes | ||
| 159 | TPM2 | ENSG00000198467.15 | 30252 | -0.252 | -0.0838 | Yes | ||
| 160 | TNNI2 | ENSG00000130598.16 | 30512 | -0.260 | -0.0851 | Yes | ||
| 161 | DAPK2 | ENSG00000035664.11 | 30624 | -0.263 | -0.0827 | Yes | ||
| 162 | CHRNA1 | ENSG00000138435.16 | 31043 | -0.279 | -0.0875 | Yes | ||
| 163 | FHL1 | ENSG00000022267.19 | 32067 | -0.299 | -0.1067 | Yes | ||
| 164 | CNN3 | ENSG00000117519.16 | 32157 | -0.302 | -0.1031 | Yes | ||
| 165 | CASQ2 | ENSG00000118729.11 | 32183 | -0.303 | -0.0978 | Yes | ||
| 166 | SORBS1 | ENSG00000095637.22 | 32350 | -0.311 | -0.0958 | Yes | ||
| 167 | MYO1C | ENSG00000197879.17 | 32509 | -0.317 | -0.0936 | Yes | ||
| 168 | FKBP1B | ENSG00000119782.14 | 32949 | -0.334 | -0.0978 | Yes | ||
| 169 | CD36 | ENSG00000135218.19 | 32994 | -0.336 | -0.0924 | Yes | ||
| 170 | RB1 | ENSG00000139687.15 | 33569 | -0.339 | -0.0999 | Yes | ||
| 171 | FGF2 | ENSG00000138685.15 | 34208 | -0.350 | -0.1087 | Yes | ||
| 172 | MRAS | ENSG00000158186.13 | 35004 | -0.365 | -0.1211 | Yes | ||
| 173 | MYL1 | ENSG00000168530.16 | 35045 | -0.367 | -0.1150 | Yes | ||
| 174 | MYLK | ENSG00000065534.18 | 35071 | -0.368 | -0.1085 | Yes | ||
| 175 | TPM3 | ENSG00000143549.21 | 35675 | -0.397 | -0.1155 | Yes | ||
| 176 | SPHK1 | ENSG00000176170.13 | 35713 | -0.398 | -0.1087 | Yes | ||
| 177 | SMTN | ENSG00000183963.18 | 36357 | -0.419 | -0.1163 | Yes | ||
| 178 | IGFBP7 | ENSG00000163453.11 | 36410 | -0.421 | -0.1095 | Yes | ||
| 179 | AEBP1 | ENSG00000106624.11 | 36626 | -0.432 | -0.1064 | Yes | ||
| 180 | CSRP3 | ENSG00000129170.10 | 36792 | -0.439 | -0.1019 | Yes | ||
| 181 | MYF6 | ENSG00000111046.4 | 37029 | -0.454 | -0.0989 | Yes | ||
| 182 | SSPN | ENSG00000123096.11 | 37348 | -0.474 | -0.0975 | Yes | ||
| 183 | MYH11 | ENSG00000133392.18 | 37382 | -0.477 | -0.0891 | Yes | ||
| 184 | SGCD | ENSG00000170624.13 | 37487 | -0.484 | -0.0823 | Yes | ||
| 185 | SVIL | ENSG00000197321.14 | 37602 | -0.494 | -0.0755 | Yes | ||
| 186 | ITGB1 | ENSG00000150093.19 | 37687 | -0.501 | -0.0679 | Yes | ||
| 187 | MYOZ1 | ENSG00000177791.12 | 37713 | -0.503 | -0.0588 | Yes | ||
| 188 | ANKRD2 | ENSG00000165887.11 | 38073 | -0.533 | -0.0573 | Yes | ||
| 189 | RIT1 | ENSG00000143622.11 | 38248 | -0.546 | -0.0510 | Yes | ||
| 190 | TAGLN | ENSG00000149591.17 | 38295 | -0.551 | -0.0415 | Yes | ||
| 191 | HBEGF | ENSG00000113070.8 | 38438 | -0.558 | -0.0342 | Yes | ||
| 192 | CKB | ENSG00000166165.13 | 38832 | -0.583 | -0.0325 | Yes | ||
| 193 | CRYAB | ENSG00000109846.9 | 38897 | -0.590 | -0.0227 | Yes | ||
| 194 | CDH13 | ENSG00000140945.17 | 39497 | -0.634 | -0.0250 | Yes | ||
| 195 | MYH9 | ENSG00000100345.21 | 39531 | -0.641 | -0.0135 | Yes | ||
| 196 | GADD45B | ENSG00000099860.9 | 40019 | -0.716 | -0.0115 | Yes | ||
| 197 | ACTC1 | ENSG00000159251.8 | 40334 | -0.785 | -0.0040 | Yes | ||
| 198 | FST | ENSG00000134363.12 | 41059 | -1.252 | 0.0025 | Yes |