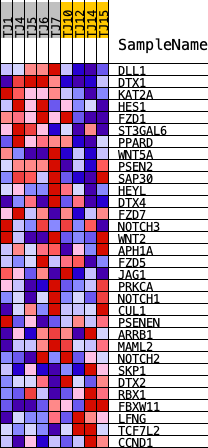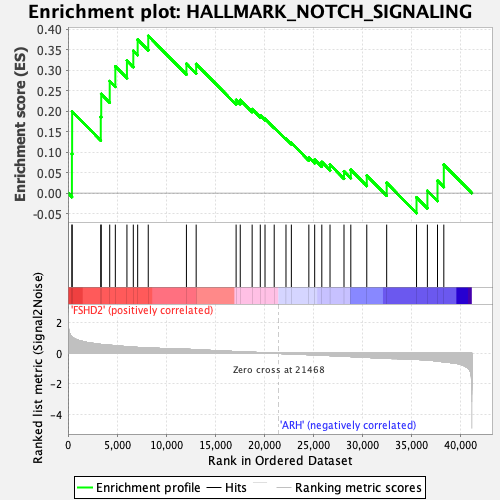
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
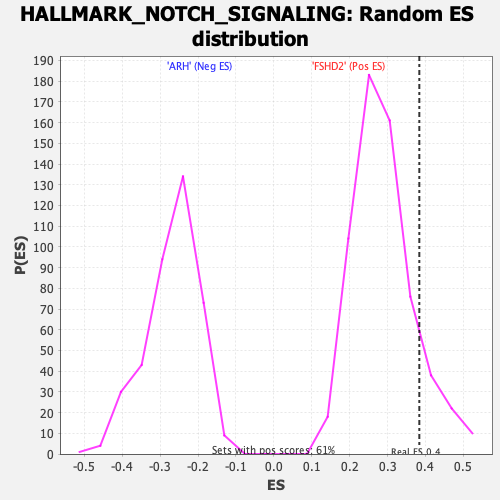
| GeneSet | HALLMARK_NOTCH_SIGNALING |
| Enrichment Score (ES) | 0.38387233 |
| Normalized Enrichment Score (NES) | 1.3279853 |
| Nominal p-value | 0.12418301 |
| FDR q-value | 0.10529469 |
| FWER p-Value | 0.839 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DLL1 | ENSG00000198719.9 | 399 | 1.062 | 0.0957 | Yes | ||
| 2 | DTX1 | ENSG00000135144.7 | 421 | 1.050 | 0.1993 | Yes | ||
| 3 | KAT2A | ENSG00000108773.11 | 3340 | 0.581 | 0.1860 | Yes | ||
| 4 | HES1 | ENSG00000114315.4 | 3386 | 0.577 | 0.2421 | Yes | ||
| 5 | FZD1 | ENSG00000157240.4 | 4252 | 0.527 | 0.2733 | Yes | ||
| 6 | ST3GAL6 | ENSG00000064225.12 | 4825 | 0.502 | 0.3093 | Yes | ||
| 7 | PPARD | ENSG00000112033.14 | 6007 | 0.433 | 0.3235 | Yes | ||
| 8 | WNT5A | ENSG00000114251.14 | 6657 | 0.401 | 0.3475 | Yes | ||
| 9 | PSEN2 | ENSG00000143801.17 | 7101 | 0.385 | 0.3750 | Yes | ||
| 10 | SAP30 | ENSG00000164105.4 | 8180 | 0.354 | 0.3839 | Yes | ||
| 11 | HEYL | ENSG00000163909.8 | 12071 | 0.268 | 0.3159 | No | ||
| 12 | DTX4 | ENSG00000110042.8 | 13064 | 0.236 | 0.3151 | No | ||
| 13 | FZD7 | ENSG00000155760.2 | 17136 | 0.114 | 0.2275 | No | ||
| 14 | NOTCH3 | ENSG00000074181.9 | 17558 | 0.102 | 0.2273 | No | ||
| 15 | WNT2 | ENSG00000105989.10 | 18770 | 0.071 | 0.2050 | No | ||
| 16 | APH1A | ENSG00000117362.13 | 19600 | 0.049 | 0.1897 | No | ||
| 17 | FZD5 | ENSG00000163251.4 | 20082 | 0.037 | 0.1817 | No | ||
| 18 | JAG1 | ENSG00000101384.12 | 21019 | 0.012 | 0.1600 | No | ||
| 19 | PRKCA | ENSG00000154229.12 | 22219 | -0.019 | 0.1327 | No | ||
| 20 | NOTCH1 | ENSG00000148400.12 | 22768 | -0.032 | 0.1226 | No | ||
| 21 | CUL1 | ENSG00000055130.17 | 24550 | -0.079 | 0.0871 | No | ||
| 22 | PSENEN | ENSG00000205155.8 | 25144 | -0.094 | 0.0820 | No | ||
| 23 | ARRB1 | ENSG00000137486.17 | 25871 | -0.116 | 0.0759 | No | ||
| 24 | MAML2 | ENSG00000184384.14 | 26708 | -0.141 | 0.0696 | No | ||
| 25 | NOTCH2 | ENSG00000134250.20 | 28133 | -0.185 | 0.0533 | No | ||
| 26 | SKP1 | ENSG00000113558.18 | 28827 | -0.206 | 0.0568 | No | ||
| 27 | DTX2 | ENSG00000091073.19 | 30454 | -0.258 | 0.0429 | No | ||
| 28 | RBX1 | ENSG00000100387.9 | 32483 | -0.316 | 0.0250 | No | ||
| 29 | FBXW11 | ENSG00000072803.17 | 35526 | -0.390 | -0.0103 | No | ||
| 30 | LFNG | ENSG00000106003.13 | 36633 | -0.433 | 0.0058 | No | ||
| 31 | TCF7L2 | ENSG00000148737.17 | 37664 | -0.499 | 0.0302 | No | ||
| 32 | CCND1 | ENSG00000110092.3 | 38307 | -0.552 | 0.0694 | No |