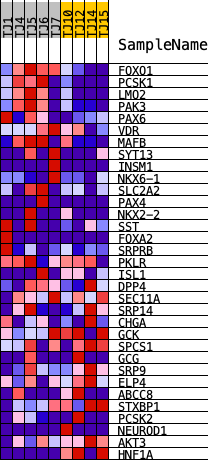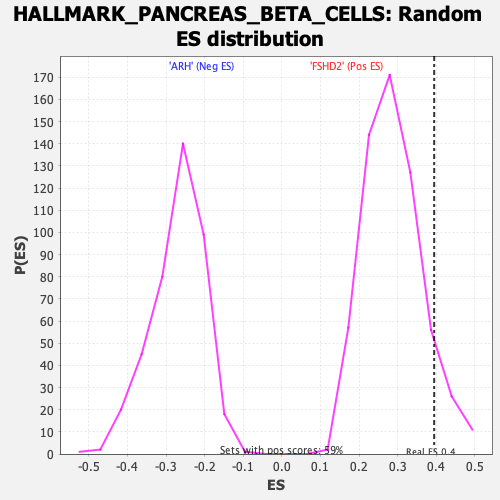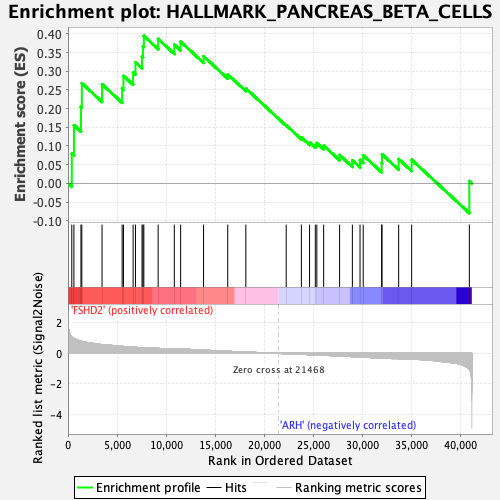
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_PANCREAS_BETA_CELLS |
| Enrichment Score (ES) | 0.39458776 |
| Normalized Enrichment Score (NES) | 1.3713144 |
| Nominal p-value | 0.09259259 |
| FDR q-value | 0.102051325 |
| FWER p-Value | 0.72 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FOXO1 | ENSG00000150907.9 | 386 | 1.072 | 0.0798 | Yes | ||
| 2 | PCSK1 | ENSG00000175426.11 | 603 | 0.976 | 0.1556 | Yes | ||
| 3 | LMO2 | ENSG00000135363.12 | 1320 | 0.799 | 0.2046 | Yes | ||
| 4 | PAK3 | ENSG00000077264.15 | 1408 | 0.788 | 0.2680 | Yes | ||
| 5 | PAX6 | ENSG00000007372.23 | 3471 | 0.569 | 0.2651 | Yes | ||
| 6 | VDR | ENSG00000111424.11 | 5518 | 0.458 | 0.2535 | Yes | ||
| 7 | MAFB | ENSG00000204103.4 | 5655 | 0.450 | 0.2876 | Yes | ||
| 8 | SYT13 | ENSG00000019505.8 | 6637 | 0.402 | 0.2971 | Yes | ||
| 9 | INSM1 | ENSG00000173404.5 | 6877 | 0.394 | 0.3241 | Yes | ||
| 10 | NKX6-1 | ENSG00000163623.9 | 7545 | 0.369 | 0.3385 | Yes | ||
| 11 | SLC2A2 | ENSG00000163581.14 | 7642 | 0.365 | 0.3665 | Yes | ||
| 12 | PAX4 | ENSG00000106331.17 | 7730 | 0.363 | 0.3946 | Yes | ||
| 13 | NKX2-2 | ENSG00000125820.6 | 9187 | 0.328 | 0.3864 | No | ||
| 14 | SST | ENSG00000157005.4 | 10837 | 0.297 | 0.3710 | No | ||
| 15 | FOXA2 | ENSG00000125798.14 | 11481 | 0.285 | 0.3791 | No | ||
| 16 | SRPRB | ENSG00000144867.12 | 13817 | 0.213 | 0.3400 | No | ||
| 17 | PKLR | ENSG00000143627.19 | 16283 | 0.138 | 0.2915 | No | ||
| 18 | ISL1 | ENSG00000016082.15 | 18126 | 0.087 | 0.2540 | No | ||
| 19 | DPP4 | ENSG00000197635.10 | 22241 | -0.019 | 0.1555 | No | ||
| 20 | SEC11A | ENSG00000140612.14 | 23787 | -0.060 | 0.1229 | No | ||
| 21 | SRP14 | ENSG00000140319.11 | 24623 | -0.081 | 0.1093 | No | ||
| 22 | CHGA | ENSG00000100604.13 | 25209 | -0.096 | 0.1031 | No | ||
| 23 | GCK | ENSG00000106633.17 | 25365 | -0.101 | 0.1078 | No | ||
| 24 | SPCS1 | ENSG00000114902.14 | 26059 | -0.123 | 0.1011 | No | ||
| 25 | GCG | ENSG00000115263.15 | 27686 | -0.170 | 0.0757 | No | ||
| 26 | SRP9 | ENSG00000143742.13 | 28986 | -0.212 | 0.0617 | No | ||
| 27 | ELP4 | ENSG00000109911.19 | 29767 | -0.237 | 0.0624 | No | ||
| 28 | ABCC8 | ENSG00000006071.14 | 30090 | -0.247 | 0.0751 | No | ||
| 29 | STXBP1 | ENSG00000136854.21 | 31972 | -0.295 | 0.0539 | No | ||
| 30 | PCSK2 | ENSG00000125851.10 | 32007 | -0.297 | 0.0778 | No | ||
| 31 | NEUROD1 | ENSG00000162992.3 | 33705 | -0.342 | 0.0649 | No | ||
| 32 | AKT3 | ENSG00000117020.18 | 35028 | -0.366 | 0.0632 | No | ||
| 33 | HNF1A | ENSG00000135100.17 | 40909 | -1.033 | 0.0061 | No |