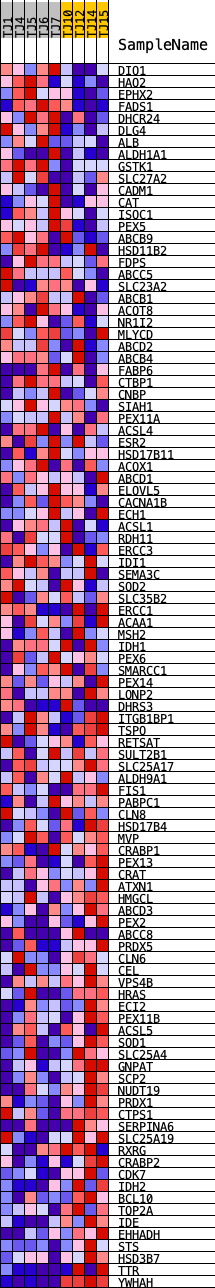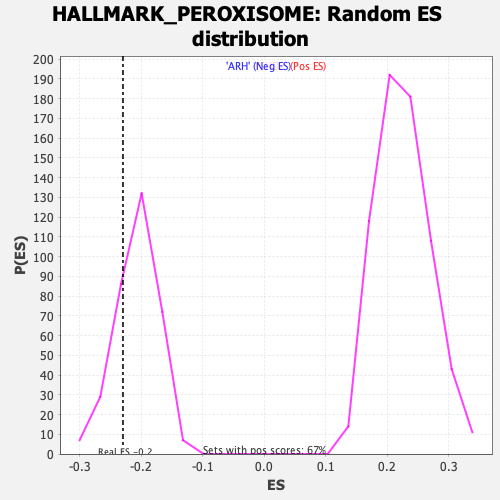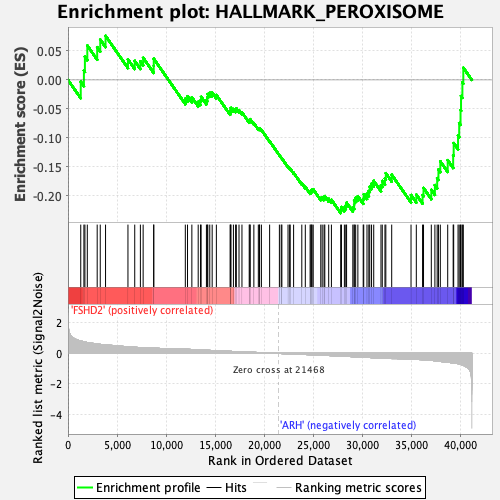
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_PEROXISOME |
| Enrichment Score (ES) | -0.22954817 |
| Normalized Enrichment Score (NES) | -1.1137652 |
| Nominal p-value | 0.21921922 |
| FDR q-value | 0.28918314 |
| FWER p-Value | 0.98 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DIO1 | ENSG00000211452.10 | 1299 | 0.803 | -0.0031 | No | ||
| 2 | HAO2 | ENSG00000116882.14 | 1621 | 0.756 | 0.0160 | No | ||
| 3 | EPHX2 | ENSG00000120915.14 | 1723 | 0.744 | 0.0400 | No | ||
| 4 | FADS1 | ENSG00000149485.19 | 1968 | 0.709 | 0.0592 | No | ||
| 5 | DHCR24 | ENSG00000116133.13 | 2975 | 0.608 | 0.0564 | No | ||
| 6 | DLG4 | ENSG00000132535.19 | 3287 | 0.584 | 0.0696 | No | ||
| 7 | ALB | ENSG00000163631.17 | 3827 | 0.547 | 0.0759 | No | ||
| 8 | ALDH1A1 | ENSG00000165092.13 | 6111 | 0.428 | 0.0355 | No | ||
| 9 | GSTK1 | ENSG00000197448.14 | 6802 | 0.396 | 0.0328 | No | ||
| 10 | SLC27A2 | ENSG00000140284.11 | 7378 | 0.373 | 0.0321 | No | ||
| 11 | CADM1 | ENSG00000182985.17 | 7659 | 0.364 | 0.0382 | No | ||
| 12 | CAT | ENSG00000121691.7 | 8736 | 0.346 | 0.0243 | No | ||
| 13 | ISOC1 | ENSG00000066583.12 | 8741 | 0.346 | 0.0365 | No | ||
| 14 | PEX5 | ENSG00000139197.10 | 11958 | 0.273 | -0.0321 | No | ||
| 15 | ABCB9 | ENSG00000150967.18 | 12183 | 0.265 | -0.0282 | No | ||
| 16 | HSD11B2 | ENSG00000176387.7 | 12625 | 0.251 | -0.0300 | No | ||
| 17 | FDPS | ENSG00000160752.14 | 13271 | 0.229 | -0.0375 | No | ||
| 18 | ABCC5 | ENSG00000114770.17 | 13504 | 0.221 | -0.0353 | No | ||
| 19 | SLC23A2 | ENSG00000089057.15 | 13569 | 0.220 | -0.0291 | No | ||
| 20 | ABCB1 | ENSG00000085563.14 | 14109 | 0.202 | -0.0350 | No | ||
| 21 | ACOT8 | ENSG00000101473.17 | 14200 | 0.199 | -0.0301 | No | ||
| 22 | NR1I2 | ENSG00000144852.17 | 14251 | 0.198 | -0.0243 | No | ||
| 23 | MLYCD | ENSG00000103150.7 | 14450 | 0.192 | -0.0223 | No | ||
| 24 | ABCD2 | ENSG00000173208.4 | 14695 | 0.184 | -0.0217 | No | ||
| 25 | ABCB4 | ENSG00000005471.17 | 15123 | 0.170 | -0.0260 | No | ||
| 26 | FABP6 | ENSG00000170231.15 | 16524 | 0.130 | -0.0555 | No | ||
| 27 | CTBP1 | ENSG00000159692.15 | 16582 | 0.129 | -0.0523 | No | ||
| 28 | CNBP | ENSG00000169714.16 | 16602 | 0.128 | -0.0482 | No | ||
| 29 | SIAH1 | ENSG00000196470.12 | 16861 | 0.121 | -0.0502 | No | ||
| 30 | PEX11A | ENSG00000166821.9 | 17086 | 0.115 | -0.0515 | No | ||
| 31 | ACSL4 | ENSG00000068366.20 | 17156 | 0.113 | -0.0492 | No | ||
| 32 | ESR2 | ENSG00000140009.18 | 17431 | 0.106 | -0.0521 | No | ||
| 33 | HSD17B11 | ENSG00000198189.11 | 17731 | 0.098 | -0.0559 | No | ||
| 34 | ACOX1 | ENSG00000161533.12 | 18471 | 0.080 | -0.0710 | No | ||
| 35 | ABCD1 | ENSG00000101986.12 | 18545 | 0.078 | -0.0700 | No | ||
| 36 | ELOVL5 | ENSG00000012660.14 | 18561 | 0.077 | -0.0677 | No | ||
| 37 | CACNA1B | ENSG00000148408.13 | 18944 | 0.067 | -0.0746 | No | ||
| 38 | ECH1 | ENSG00000104823.9 | 19400 | 0.054 | -0.0838 | No | ||
| 39 | ACSL1 | ENSG00000151726.14 | 19506 | 0.051 | -0.0845 | No | ||
| 40 | RDH11 | ENSG00000072042.13 | 19531 | 0.051 | -0.0833 | No | ||
| 41 | ERCC3 | ENSG00000163161.13 | 19714 | 0.046 | -0.0861 | No | ||
| 42 | IDI1 | ENSG00000067064.11 | 20552 | 0.023 | -0.1056 | No | ||
| 43 | SEMA3C | ENSG00000075223.14 | 21547 | -0.002 | -0.1298 | No | ||
| 44 | SOD2 | ENSG00000112096.18 | 21731 | -0.007 | -0.1340 | No | ||
| 45 | SLC35B2 | ENSG00000157593.19 | 21800 | -0.009 | -0.1353 | No | ||
| 46 | ERCC1 | ENSG00000012061.15 | 22428 | -0.024 | -0.1497 | No | ||
| 47 | ACAA1 | ENSG00000060971.18 | 22593 | -0.028 | -0.1527 | No | ||
| 48 | MSH2 | ENSG00000095002.14 | 22645 | -0.030 | -0.1529 | No | ||
| 49 | IDH1 | ENSG00000138413.13 | 22996 | -0.039 | -0.1600 | No | ||
| 50 | PEX6 | ENSG00000124587.14 | 23832 | -0.061 | -0.1782 | No | ||
| 51 | SMARCC1 | ENSG00000173473.11 | 24189 | -0.069 | -0.1844 | No | ||
| 52 | PEX14 | ENSG00000142655.13 | 24680 | -0.082 | -0.1934 | No | ||
| 53 | LONP2 | ENSG00000102910.14 | 24767 | -0.085 | -0.1925 | No | ||
| 54 | DHRS3 | ENSG00000162496.9 | 24804 | -0.086 | -0.1903 | No | ||
| 55 | ITGB1BP1 | ENSG00000119185.12 | 24879 | -0.088 | -0.1890 | No | ||
| 56 | TSPO | ENSG00000100300.18 | 24994 | -0.091 | -0.1885 | No | ||
| 57 | RETSAT | ENSG00000042445.14 | 25766 | -0.113 | -0.2033 | No | ||
| 58 | SULT2B1 | ENSG00000088002.11 | 25907 | -0.117 | -0.2025 | No | ||
| 59 | SLC25A17 | ENSG00000100372.15 | 26097 | -0.123 | -0.2027 | No | ||
| 60 | ALDH9A1 | ENSG00000143149.12 | 26176 | -0.125 | -0.2002 | No | ||
| 61 | FIS1 | ENSG00000214253.9 | 26573 | -0.137 | -0.2050 | No | ||
| 62 | PABPC1 | ENSG00000070756.16 | 26850 | -0.145 | -0.2065 | No | ||
| 63 | CLN8 | ENSG00000182372.9 | 27796 | -0.174 | -0.2234 | Yes | ||
| 64 | HSD17B4 | ENSG00000133835.16 | 27875 | -0.175 | -0.2190 | Yes | ||
| 65 | MVP | ENSG00000013364.19 | 28173 | -0.185 | -0.2197 | Yes | ||
| 66 | CRABP1 | ENSG00000166426.8 | 28315 | -0.190 | -0.2164 | Yes | ||
| 67 | PEX13 | ENSG00000162928.9 | 28396 | -0.193 | -0.2115 | Yes | ||
| 68 | CRAT | ENSG00000095321.17 | 29033 | -0.213 | -0.2194 | Yes | ||
| 69 | ATXN1 | ENSG00000124788.18 | 29169 | -0.217 | -0.2150 | Yes | ||
| 70 | HMGCL | ENSG00000117305.15 | 29172 | -0.217 | -0.2073 | Yes | ||
| 71 | ABCD3 | ENSG00000117528.14 | 29322 | -0.222 | -0.2030 | Yes | ||
| 72 | PEX2 | ENSG00000164751.15 | 29547 | -0.228 | -0.2004 | Yes | ||
| 73 | ABCC8 | ENSG00000006071.14 | 30090 | -0.247 | -0.2048 | Yes | ||
| 74 | PRDX5 | ENSG00000126432.14 | 30144 | -0.249 | -0.1972 | Yes | ||
| 75 | CLN6 | ENSG00000128973.13 | 30476 | -0.259 | -0.1961 | Yes | ||
| 76 | CEL | ENSG00000170835.14 | 30665 | -0.265 | -0.1912 | Yes | ||
| 77 | VPS4B | ENSG00000119541.10 | 30759 | -0.268 | -0.1840 | Yes | ||
| 78 | HRAS | ENSG00000174775.17 | 30941 | -0.275 | -0.1786 | Yes | ||
| 79 | ECI2 | ENSG00000198721.12 | 31163 | -0.283 | -0.1739 | Yes | ||
| 80 | PEX11B | ENSG00000131779.11 | 31903 | -0.293 | -0.1815 | Yes | ||
| 81 | ACSL5 | ENSG00000197142.10 | 32034 | -0.298 | -0.1741 | Yes | ||
| 82 | SOD1 | ENSG00000142168.14 | 32307 | -0.310 | -0.1697 | Yes | ||
| 83 | SLC25A4 | ENSG00000151729.11 | 32397 | -0.314 | -0.1607 | Yes | ||
| 84 | GNPAT | ENSG00000116906.13 | 32995 | -0.336 | -0.1633 | Yes | ||
| 85 | SCP2 | ENSG00000116171.18 | 34965 | -0.364 | -0.1984 | Yes | ||
| 86 | NUDT19 | ENSG00000213965.3 | 35504 | -0.389 | -0.1976 | Yes | ||
| 87 | PRDX1 | ENSG00000117450.14 | 36141 | -0.411 | -0.1985 | Yes | ||
| 88 | CTPS1 | ENSG00000171793.16 | 36239 | -0.413 | -0.1862 | Yes | ||
| 89 | SERPINA6 | ENSG00000170099.6 | 37032 | -0.454 | -0.1893 | Yes | ||
| 90 | SLC25A19 | ENSG00000125454.12 | 37398 | -0.478 | -0.1812 | Yes | ||
| 91 | RXRG | ENSG00000143171.13 | 37631 | -0.496 | -0.1692 | Yes | ||
| 92 | CRABP2 | ENSG00000143320.9 | 37760 | -0.508 | -0.1543 | Yes | ||
| 93 | CDK7 | ENSG00000134058.12 | 37950 | -0.524 | -0.1403 | Yes | ||
| 94 | IDH2 | ENSG00000182054.9 | 38704 | -0.571 | -0.1383 | Yes | ||
| 95 | BCL10 | ENSG00000142867.14 | 39261 | -0.619 | -0.1298 | Yes | ||
| 96 | TOP2A | ENSG00000131747.15 | 39313 | -0.624 | -0.1089 | Yes | ||
| 97 | IDE | ENSG00000119912.17 | 39758 | -0.668 | -0.0959 | Yes | ||
| 98 | EHHADH | ENSG00000113790.11 | 39876 | -0.690 | -0.0743 | Yes | ||
| 99 | STS | ENSG00000101846.7 | 40012 | -0.715 | -0.0521 | Yes | ||
| 100 | HSD3B7 | ENSG00000099377.14 | 40063 | -0.724 | -0.0276 | Yes | ||
| 101 | TTR | ENSG00000118271.10 | 40201 | -0.753 | -0.0042 | Yes | ||
| 102 | YWHAH | ENSG00000128245.15 | 40295 | -0.774 | 0.0211 | Yes |