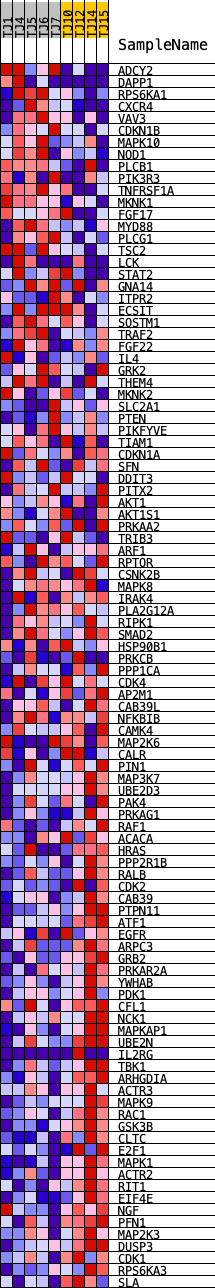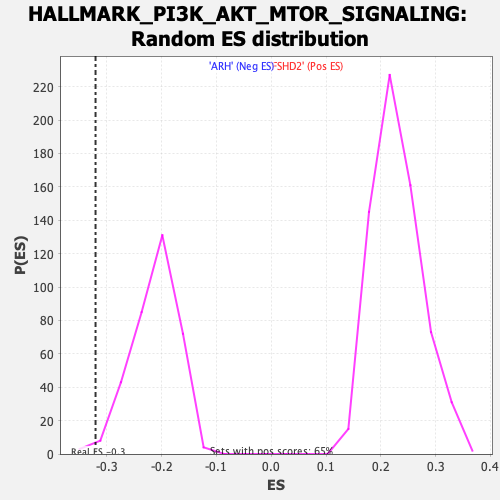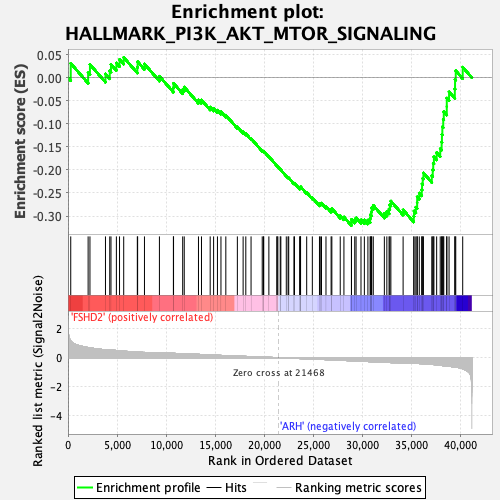
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_PI3K_AKT_MTOR_SIGNALING |
| Enrichment Score (ES) | -0.32066926 |
| Normalized Enrichment Score (NES) | -1.508644 |
| Nominal p-value | 0.011560693 |
| FDR q-value | 0.02174344 |
| FWER p-Value | 0.138 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADCY2 | ENSG00000078295.17 | 279 | 1.147 | 0.0309 | No | ||
| 2 | DAPP1 | ENSG00000070190.13 | 2051 | 0.699 | 0.0106 | No | ||
| 3 | RPS6KA1 | ENSG00000117676.14 | 2237 | 0.680 | 0.0285 | No | ||
| 4 | CXCR4 | ENSG00000121966.6 | 3819 | 0.548 | 0.0079 | No | ||
| 5 | VAV3 | ENSG00000134215.16 | 4256 | 0.527 | 0.0146 | No | ||
| 6 | CDKN1B | ENSG00000111276.11 | 4380 | 0.521 | 0.0287 | No | ||
| 7 | MAPK10 | ENSG00000109339.23 | 4931 | 0.496 | 0.0316 | No | ||
| 8 | NOD1 | ENSG00000106100.11 | 5252 | 0.474 | 0.0393 | No | ||
| 9 | PLCB1 | ENSG00000182621.18 | 5678 | 0.449 | 0.0437 | No | ||
| 10 | PIK3R3 | ENSG00000117461.15 | 7066 | 0.387 | 0.0226 | No | ||
| 11 | TNFRSF1A | ENSG00000067182.7 | 7098 | 0.385 | 0.0345 | No | ||
| 12 | MKNK1 | ENSG00000079277.21 | 7794 | 0.363 | 0.0295 | No | ||
| 13 | FGF17 | ENSG00000158815.11 | 9318 | 0.322 | 0.0029 | No | ||
| 14 | MYD88 | ENSG00000172936.14 | 10743 | 0.301 | -0.0219 | No | ||
| 15 | PLCG1 | ENSG00000124181.14 | 10758 | 0.300 | -0.0124 | No | ||
| 16 | TSC2 | ENSG00000103197.18 | 11691 | 0.283 | -0.0258 | No | ||
| 17 | LCK | ENSG00000182866.17 | 11848 | 0.276 | -0.0205 | No | ||
| 18 | STAT2 | ENSG00000170581.14 | 13299 | 0.229 | -0.0483 | No | ||
| 19 | GNA14 | ENSG00000156049.7 | 13606 | 0.219 | -0.0486 | No | ||
| 20 | ITPR2 | ENSG00000123104.12 | 14493 | 0.191 | -0.0639 | No | ||
| 21 | ECSIT | ENSG00000130159.14 | 14842 | 0.179 | -0.0665 | No | ||
| 22 | SQSTM1 | ENSG00000161011.20 | 15233 | 0.167 | -0.0706 | No | ||
| 23 | TRAF2 | ENSG00000127191.18 | 15591 | 0.157 | -0.0741 | No | ||
| 24 | FGF22 | ENSG00000070388.11 | 16087 | 0.143 | -0.0815 | No | ||
| 25 | IL4 | ENSG00000113520.11 | 17263 | 0.110 | -0.1065 | No | ||
| 26 | GRK2 | ENSG00000173020.11 | 17850 | 0.095 | -0.1177 | No | ||
| 27 | THEM4 | ENSG00000159445.13 | 18124 | 0.087 | -0.1214 | No | ||
| 28 | MKNK2 | ENSG00000099875.14 | 18659 | 0.075 | -0.1320 | No | ||
| 29 | SLC2A1 | ENSG00000117394.23 | 19798 | 0.043 | -0.1583 | No | ||
| 30 | PTEN | ENSG00000171862.11 | 19922 | 0.040 | -0.1600 | No | ||
| 31 | PIKFYVE | ENSG00000115020.17 | 19952 | 0.040 | -0.1594 | No | ||
| 32 | TIAM1 | ENSG00000156299.13 | 20475 | 0.025 | -0.1713 | No | ||
| 33 | CDKN1A | ENSG00000124762.13 | 21297 | 0.004 | -0.1911 | No | ||
| 34 | SFN | ENSG00000175793.12 | 21306 | 0.004 | -0.1912 | No | ||
| 35 | DDIT3 | ENSG00000175197.12 | 21390 | 0.001 | -0.1932 | No | ||
| 36 | PITX2 | ENSG00000164093.17 | 21632 | -0.005 | -0.1989 | No | ||
| 37 | AKT1 | ENSG00000142208.16 | 21695 | -0.006 | -0.2002 | No | ||
| 38 | AKT1S1 | ENSG00000204673.10 | 22250 | -0.019 | -0.2131 | No | ||
| 39 | PRKAA2 | ENSG00000162409.11 | 22430 | -0.024 | -0.2166 | No | ||
| 40 | TRIB3 | ENSG00000101255.11 | 22498 | -0.027 | -0.2174 | No | ||
| 41 | ARF1 | ENSG00000143761.16 | 23029 | -0.040 | -0.2290 | No | ||
| 42 | RPTOR | ENSG00000141564.15 | 23086 | -0.042 | -0.2289 | No | ||
| 43 | CSNK2B | ENSG00000204435.13 | 23621 | -0.056 | -0.2401 | No | ||
| 44 | MAPK8 | ENSG00000107643.16 | 23655 | -0.057 | -0.2391 | No | ||
| 45 | IRAK4 | ENSG00000198001.13 | 23681 | -0.057 | -0.2378 | No | ||
| 46 | PLA2G12A | ENSG00000123739.11 | 23706 | -0.058 | -0.2365 | No | ||
| 47 | RIPK1 | ENSG00000137275.14 | 24330 | -0.074 | -0.2492 | No | ||
| 48 | SMAD2 | ENSG00000175387.16 | 24905 | -0.088 | -0.2603 | No | ||
| 49 | HSP90B1 | ENSG00000166598.15 | 25632 | -0.109 | -0.2744 | No | ||
| 50 | PRKCB | ENSG00000166501.14 | 25710 | -0.111 | -0.2726 | No | ||
| 51 | PPP1CA | ENSG00000172531.15 | 25840 | -0.115 | -0.2720 | No | ||
| 52 | CDK4 | ENSG00000135446.17 | 26296 | -0.129 | -0.2788 | No | ||
| 53 | AP2M1 | ENSG00000161203.13 | 26819 | -0.144 | -0.2868 | No | ||
| 54 | CAB39L | ENSG00000102547.19 | 26895 | -0.146 | -0.2839 | No | ||
| 55 | NFKBIB | ENSG00000104825.17 | 27749 | -0.172 | -0.2990 | No | ||
| 56 | CAMK4 | ENSG00000152495.11 | 28125 | -0.184 | -0.3021 | No | ||
| 57 | MAP2K6 | ENSG00000108984.15 | 28890 | -0.208 | -0.3138 | Yes | ||
| 58 | CALR | ENSG00000179218.13 | 28898 | -0.208 | -0.3072 | Yes | ||
| 59 | PIN1 | ENSG00000127445.13 | 29228 | -0.218 | -0.3080 | Yes | ||
| 60 | MAP3K7 | ENSG00000135341.18 | 29362 | -0.223 | -0.3040 | Yes | ||
| 61 | UBE2D3 | ENSG00000109332.20 | 29865 | -0.240 | -0.3083 | Yes | ||
| 62 | PAK4 | ENSG00000130669.17 | 30208 | -0.250 | -0.3084 | Yes | ||
| 63 | PRKAG1 | ENSG00000181929.13 | 30570 | -0.262 | -0.3086 | Yes | ||
| 64 | RAF1 | ENSG00000132155.11 | 30802 | -0.270 | -0.3054 | Yes | ||
| 65 | ACACA | ENSG00000278540.5 | 30823 | -0.270 | -0.2970 | Yes | ||
| 66 | HRAS | ENSG00000174775.17 | 30941 | -0.275 | -0.2908 | Yes | ||
| 67 | PPP2R1B | ENSG00000137713.16 | 30954 | -0.276 | -0.2821 | Yes | ||
| 68 | RALB | ENSG00000144118.14 | 31117 | -0.282 | -0.2768 | Yes | ||
| 69 | CDK2 | ENSG00000123374.11 | 32246 | -0.307 | -0.2942 | Yes | ||
| 70 | CAB39 | ENSG00000135932.11 | 32500 | -0.317 | -0.2900 | Yes | ||
| 71 | PTPN11 | ENSG00000179295.17 | 32726 | -0.325 | -0.2848 | Yes | ||
| 72 | ATF1 | ENSG00000123268.9 | 32800 | -0.328 | -0.2758 | Yes | ||
| 73 | EGFR | ENSG00000146648.18 | 32910 | -0.332 | -0.2676 | Yes | ||
| 74 | ARPC3 | ENSG00000111229.16 | 34158 | -0.348 | -0.2865 | Yes | ||
| 75 | GRB2 | ENSG00000177885.15 | 35242 | -0.375 | -0.3006 | Yes | ||
| 76 | PRKAR2A | ENSG00000114302.16 | 35272 | -0.377 | -0.2889 | Yes | ||
| 77 | YWHAB | ENSG00000166913.13 | 35457 | -0.386 | -0.2808 | Yes | ||
| 78 | PDK1 | ENSG00000152256.13 | 35589 | -0.393 | -0.2711 | Yes | ||
| 79 | CFL1 | ENSG00000172757.13 | 35593 | -0.393 | -0.2582 | Yes | ||
| 80 | NCK1 | ENSG00000158092.7 | 35806 | -0.402 | -0.2502 | Yes | ||
| 81 | MAPKAP1 | ENSG00000119487.17 | 36057 | -0.406 | -0.2429 | Yes | ||
| 82 | UBE2N | ENSG00000177889.10 | 36090 | -0.408 | -0.2303 | Yes | ||
| 83 | IL2RG | ENSG00000147168.12 | 36157 | -0.412 | -0.2184 | Yes | ||
| 84 | TBK1 | ENSG00000183735.10 | 36229 | -0.412 | -0.2066 | Yes | ||
| 85 | ARHGDIA | ENSG00000141522.12 | 37082 | -0.457 | -0.2124 | Yes | ||
| 86 | ACTR3 | ENSG00000115091.12 | 37178 | -0.462 | -0.1995 | Yes | ||
| 87 | MAPK9 | ENSG00000050748.17 | 37228 | -0.465 | -0.1855 | Yes | ||
| 88 | RAC1 | ENSG00000136238.18 | 37298 | -0.471 | -0.1717 | Yes | ||
| 89 | GSK3B | ENSG00000082701.15 | 37577 | -0.491 | -0.1623 | Yes | ||
| 90 | CLTC | ENSG00000141367.12 | 37942 | -0.523 | -0.1540 | Yes | ||
| 91 | E2F1 | ENSG00000101412.13 | 38085 | -0.533 | -0.1400 | Yes | ||
| 92 | MAPK1 | ENSG00000100030.14 | 38116 | -0.536 | -0.1232 | Yes | ||
| 93 | ACTR2 | ENSG00000138071.13 | 38158 | -0.539 | -0.1065 | Yes | ||
| 94 | RIT1 | ENSG00000143622.11 | 38248 | -0.546 | -0.0907 | Yes | ||
| 95 | EIF4E | ENSG00000151247.12 | 38303 | -0.552 | -0.0739 | Yes | ||
| 96 | NGF | ENSG00000134259.3 | 38608 | -0.563 | -0.0629 | Yes | ||
| 97 | PFN1 | ENSG00000108518.7 | 38610 | -0.563 | -0.0444 | Yes | ||
| 98 | MAP2K3 | ENSG00000034152.19 | 38842 | -0.584 | -0.0309 | Yes | ||
| 99 | DUSP3 | ENSG00000108861.9 | 39429 | -0.629 | -0.0245 | Yes | ||
| 100 | CDK1 | ENSG00000170312.16 | 39453 | -0.632 | -0.0044 | Yes | ||
| 101 | RPS6KA3 | ENSG00000177189.14 | 39524 | -0.639 | 0.0149 | Yes | ||
| 102 | SLA | ENSG00000155926.14 | 40233 | -0.760 | 0.0226 | Yes |