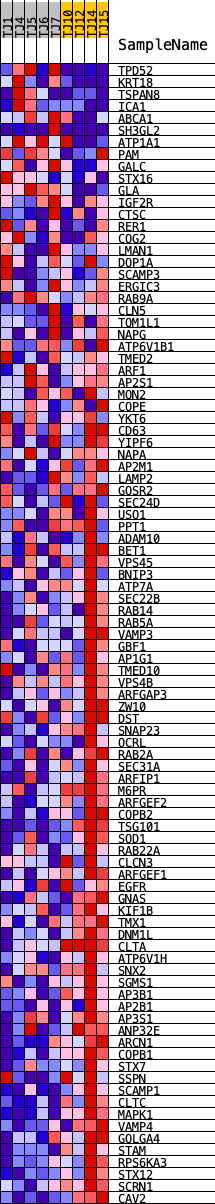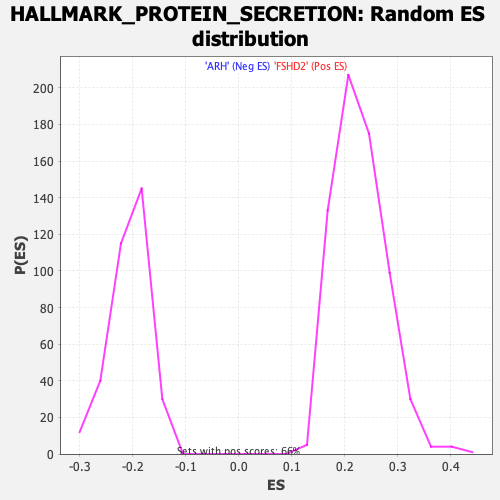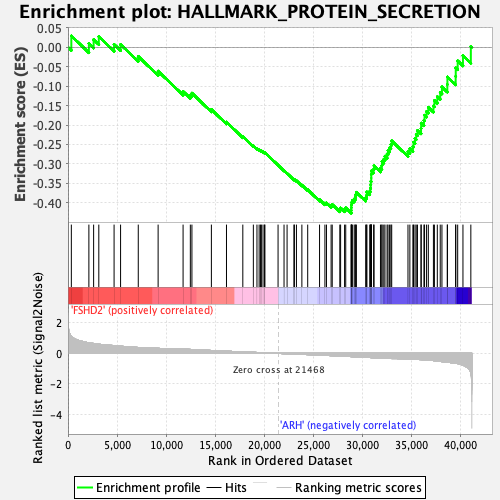
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_PROTEIN_SECRETION |
| Enrichment Score (ES) | -0.42682368 |
| Normalized Enrichment Score (NES) | -2.0630693 |
| Nominal p-value | 0.0 |
| FDR q-value | 2.5210084E-4 |
| FWER p-Value | 0.001 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TPD52 | ENSG00000076554.15 | 340 | 1.096 | 0.0295 | No | ||
| 2 | KRT18 | ENSG00000111057.11 | 2128 | 0.690 | 0.0097 | No | ||
| 3 | TSPAN8 | ENSG00000127324.9 | 2612 | 0.642 | 0.0201 | No | ||
| 4 | ICA1 | ENSG00000003147.19 | 3140 | 0.596 | 0.0278 | No | ||
| 5 | ABCA1 | ENSG00000165029.16 | 4697 | 0.511 | 0.0075 | No | ||
| 6 | SH3GL2 | ENSG00000107295.10 | 5359 | 0.468 | 0.0075 | No | ||
| 7 | ATP1A1 | ENSG00000163399.16 | 7162 | 0.382 | -0.0232 | No | ||
| 8 | PAM | ENSG00000145730.20 | 9186 | 0.328 | -0.0612 | No | ||
| 9 | GALC | ENSG00000054983.17 | 11730 | 0.281 | -0.1135 | No | ||
| 10 | STX16 | ENSG00000124222.22 | 12470 | 0.255 | -0.1227 | No | ||
| 11 | GLA | ENSG00000102393.10 | 12631 | 0.250 | -0.1179 | No | ||
| 12 | IGF2R | ENSG00000197081.14 | 14615 | 0.187 | -0.1598 | No | ||
| 13 | CTSC | ENSG00000109861.16 | 16154 | 0.141 | -0.1924 | No | ||
| 14 | RER1 | ENSG00000157916.20 | 17819 | 0.096 | -0.2296 | No | ||
| 15 | COG2 | ENSG00000135775.14 | 18893 | 0.068 | -0.2534 | No | ||
| 16 | LMAN1 | ENSG00000074695.6 | 19261 | 0.058 | -0.2603 | No | ||
| 17 | DOP1A | ENSG00000083097.14 | 19484 | 0.052 | -0.2640 | No | ||
| 18 | SCAMP3 | ENSG00000116521.11 | 19626 | 0.048 | -0.2657 | No | ||
| 19 | ERGIC3 | ENSG00000125991.19 | 19711 | 0.046 | -0.2662 | No | ||
| 20 | RAB9A | ENSG00000123595.8 | 19879 | 0.041 | -0.2689 | No | ||
| 21 | CLN5 | ENSG00000102805.15 | 20037 | 0.039 | -0.2714 | No | ||
| 22 | TOM1L1 | ENSG00000141198.16 | 20068 | 0.038 | -0.2708 | No | ||
| 23 | NAPG | ENSG00000134265.13 | 21412 | 0.001 | -0.3034 | No | ||
| 24 | ATP6V1B1 | ENSG00000116039.13 | 22021 | -0.015 | -0.3177 | No | ||
| 25 | TMED2 | ENSG00000086598.11 | 22331 | -0.022 | -0.3245 | No | ||
| 26 | ARF1 | ENSG00000143761.16 | 23029 | -0.040 | -0.3401 | No | ||
| 27 | AP2S1 | ENSG00000042753.11 | 23077 | -0.042 | -0.3398 | No | ||
| 28 | MON2 | ENSG00000061987.16 | 23285 | -0.046 | -0.3433 | No | ||
| 29 | COPE | ENSG00000105669.14 | 23839 | -0.062 | -0.3546 | No | ||
| 30 | YKT6 | ENSG00000106636.8 | 24433 | -0.075 | -0.3664 | No | ||
| 31 | CD63 | ENSG00000135404.11 | 25641 | -0.110 | -0.3921 | No | ||
| 32 | YIPF6 | ENSG00000181704.12 | 26187 | -0.125 | -0.4010 | No | ||
| 33 | NAPA | ENSG00000105402.8 | 26346 | -0.131 | -0.4004 | No | ||
| 34 | AP2M1 | ENSG00000161203.13 | 26819 | -0.144 | -0.4069 | No | ||
| 35 | LAMP2 | ENSG00000005893.15 | 26933 | -0.147 | -0.4046 | No | ||
| 36 | GOSR2 | ENSG00000108433.16 | 27707 | -0.171 | -0.4175 | No | ||
| 37 | SEC24D | ENSG00000150961.15 | 27788 | -0.173 | -0.4135 | No | ||
| 38 | USO1 | ENSG00000138768.14 | 28192 | -0.186 | -0.4169 | No | ||
| 39 | PPT1 | ENSG00000131238.17 | 28310 | -0.190 | -0.4132 | No | ||
| 40 | ADAM10 | ENSG00000137845.15 | 28870 | -0.207 | -0.4197 | Yes | ||
| 41 | BET1 | ENSG00000105829.13 | 28876 | -0.207 | -0.4127 | Yes | ||
| 42 | VPS45 | ENSG00000136631.15 | 28882 | -0.208 | -0.4056 | Yes | ||
| 43 | BNIP3 | ENSG00000176171.11 | 28912 | -0.209 | -0.3991 | Yes | ||
| 44 | ATP7A | ENSG00000165240.20 | 28987 | -0.212 | -0.3937 | Yes | ||
| 45 | SEC22B | ENSG00000265808.4 | 29171 | -0.217 | -0.3907 | Yes | ||
| 46 | RAB14 | ENSG00000119396.11 | 29274 | -0.220 | -0.3855 | Yes | ||
| 47 | RAB5A | ENSG00000144566.11 | 29326 | -0.222 | -0.3791 | Yes | ||
| 48 | VAMP3 | ENSG00000049245.13 | 29395 | -0.225 | -0.3731 | Yes | ||
| 49 | GBF1 | ENSG00000107862.4 | 30343 | -0.255 | -0.3873 | Yes | ||
| 50 | AP1G1 | ENSG00000166747.12 | 30442 | -0.258 | -0.3808 | Yes | ||
| 51 | TMED10 | ENSG00000170348.9 | 30459 | -0.259 | -0.3723 | Yes | ||
| 52 | VPS4B | ENSG00000119541.10 | 30759 | -0.268 | -0.3704 | Yes | ||
| 53 | ARFGAP3 | ENSG00000242247.11 | 30822 | -0.270 | -0.3626 | Yes | ||
| 54 | ZW10 | ENSG00000086827.9 | 30856 | -0.271 | -0.3540 | Yes | ||
| 55 | DST | ENSG00000151914.20 | 30866 | -0.272 | -0.3449 | Yes | ||
| 56 | SNAP23 | ENSG00000092531.10 | 30911 | -0.274 | -0.3365 | Yes | ||
| 57 | OCRL | ENSG00000122126.17 | 30912 | -0.274 | -0.3271 | Yes | ||
| 58 | RAB2A | ENSG00000104388.15 | 30923 | -0.274 | -0.3179 | Yes | ||
| 59 | SEC31A | ENSG00000138674.17 | 31154 | -0.283 | -0.3137 | Yes | ||
| 60 | ARFIP1 | ENSG00000164144.16 | 31197 | -0.285 | -0.3049 | Yes | ||
| 61 | M6PR | ENSG00000003056.8 | 31850 | -0.291 | -0.3108 | Yes | ||
| 62 | ARFGEF2 | ENSG00000124198.9 | 31991 | -0.296 | -0.3040 | Yes | ||
| 63 | COPB2 | ENSG00000184432.10 | 32009 | -0.297 | -0.2942 | Yes | ||
| 64 | TSG101 | ENSG00000074319.13 | 32189 | -0.304 | -0.2881 | Yes | ||
| 65 | SOD1 | ENSG00000142168.14 | 32307 | -0.310 | -0.2803 | Yes | ||
| 66 | RAB22A | ENSG00000124209.4 | 32525 | -0.317 | -0.2746 | Yes | ||
| 67 | CLCN3 | ENSG00000109572.14 | 32620 | -0.321 | -0.2658 | Yes | ||
| 68 | ARFGEF1 | ENSG00000066777.9 | 32780 | -0.327 | -0.2584 | Yes | ||
| 69 | EGFR | ENSG00000146648.18 | 32910 | -0.332 | -0.2501 | Yes | ||
| 70 | GNAS | ENSG00000087460.25 | 32996 | -0.336 | -0.2407 | Yes | ||
| 71 | KIF1B | ENSG00000054523.17 | 34654 | -0.354 | -0.2688 | Yes | ||
| 72 | TMX1 | ENSG00000139921.13 | 34847 | -0.363 | -0.2610 | Yes | ||
| 73 | DNM1L | ENSG00000087470.18 | 35155 | -0.372 | -0.2557 | Yes | ||
| 74 | CLTA | ENSG00000122705.17 | 35217 | -0.375 | -0.2442 | Yes | ||
| 75 | ATP6V1H | ENSG00000047249.18 | 35368 | -0.382 | -0.2347 | Yes | ||
| 76 | SNX2 | ENSG00000205302.7 | 35495 | -0.388 | -0.2244 | Yes | ||
| 77 | SGMS1 | ENSG00000198964.14 | 35622 | -0.395 | -0.2139 | Yes | ||
| 78 | AP3B1 | ENSG00000132842.13 | 35986 | -0.405 | -0.2088 | Yes | ||
| 79 | AP2B1 | ENSG00000006125.18 | 35998 | -0.406 | -0.1951 | Yes | ||
| 80 | AP3S1 | ENSG00000177879.16 | 36282 | -0.415 | -0.1877 | Yes | ||
| 81 | ANP32E | ENSG00000143401.15 | 36344 | -0.418 | -0.1748 | Yes | ||
| 82 | ARCN1 | ENSG00000095139.14 | 36550 | -0.430 | -0.1649 | Yes | ||
| 83 | COPB1 | ENSG00000129083.12 | 36728 | -0.437 | -0.1542 | Yes | ||
| 84 | STX7 | ENSG00000079950.14 | 37247 | -0.467 | -0.1507 | Yes | ||
| 85 | SSPN | ENSG00000123096.11 | 37348 | -0.474 | -0.1368 | Yes | ||
| 86 | SCAMP1 | ENSG00000085365.18 | 37644 | -0.497 | -0.1269 | Yes | ||
| 87 | CLTC | ENSG00000141367.12 | 37942 | -0.523 | -0.1161 | Yes | ||
| 88 | MAPK1 | ENSG00000100030.14 | 38116 | -0.536 | -0.1019 | Yes | ||
| 89 | VAMP4 | ENSG00000117533.15 | 38664 | -0.567 | -0.0956 | Yes | ||
| 90 | GOLGA4 | ENSG00000144674.16 | 38674 | -0.568 | -0.0763 | Yes | ||
| 91 | STAM | ENSG00000136738.15 | 39514 | -0.637 | -0.0748 | Yes | ||
| 92 | RPS6KA3 | ENSG00000177189.14 | 39524 | -0.639 | -0.0530 | Yes | ||
| 93 | STX12 | ENSG00000117758.14 | 39721 | -0.664 | -0.0349 | Yes | ||
| 94 | SCRN1 | ENSG00000136193.17 | 40258 | -0.767 | -0.0215 | Yes | ||
| 95 | CAV2 | ENSG00000105971.15 | 41062 | -1.262 | 0.0024 | Yes |