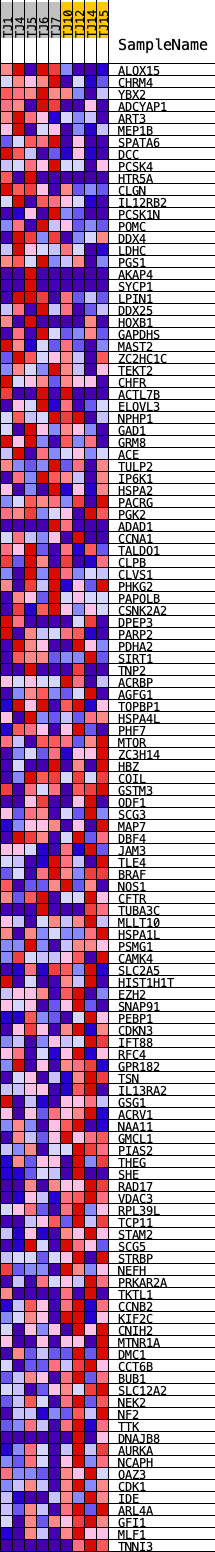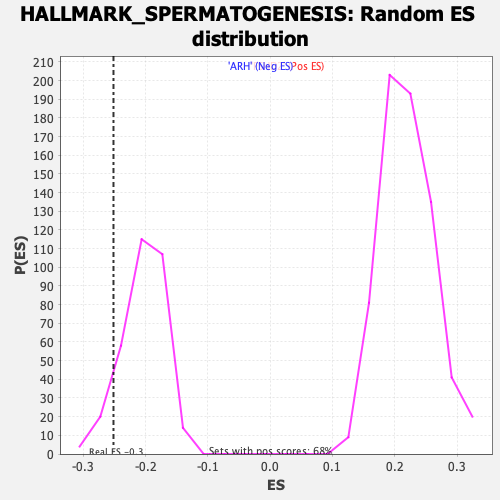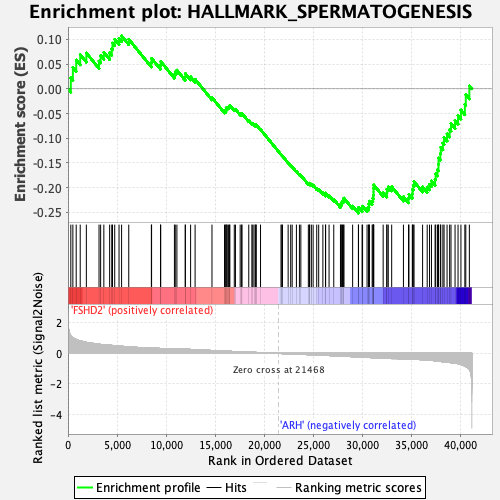
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_SPERMATOGENESIS |
| Enrichment Score (ES) | -0.25144485 |
| Normalized Enrichment Score (NES) | -1.2339151 |
| Nominal p-value | 0.07861635 |
| FDR q-value | 0.14174718 |
| FWER p-Value | 0.773 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ALOX15 | ENSG00000161905.12 | 287 | 1.141 | 0.0223 | No | ||
| 2 | CHRM4 | ENSG00000180720.7 | 496 | 1.017 | 0.0434 | No | ||
| 3 | YBX2 | ENSG00000006047.13 | 836 | 0.902 | 0.0583 | No | ||
| 4 | ADCYAP1 | ENSG00000141433.13 | 1245 | 0.810 | 0.0692 | No | ||
| 5 | ART3 | ENSG00000156219.16 | 1870 | 0.721 | 0.0725 | No | ||
| 6 | MEP1B | ENSG00000141434.12 | 3162 | 0.594 | 0.0563 | No | ||
| 7 | SPATA6 | ENSG00000132122.12 | 3326 | 0.582 | 0.0673 | No | ||
| 8 | DCC | ENSG00000187323.12 | 3653 | 0.561 | 0.0738 | No | ||
| 9 | PCSK4 | ENSG00000115257.15 | 4245 | 0.528 | 0.0729 | No | ||
| 10 | HTR5A | ENSG00000157219.5 | 4461 | 0.518 | 0.0810 | No | ||
| 11 | CLGN | ENSG00000153132.13 | 4537 | 0.516 | 0.0925 | No | ||
| 12 | IL12RB2 | ENSG00000081985.11 | 4762 | 0.507 | 0.1000 | No | ||
| 13 | PCSK1N | ENSG00000102109.9 | 5202 | 0.477 | 0.1016 | No | ||
| 14 | POMC | ENSG00000115138.11 | 5463 | 0.461 | 0.1071 | No | ||
| 15 | DDX4 | ENSG00000152670.19 | 6193 | 0.423 | 0.1002 | No | ||
| 16 | LDHC | ENSG00000166796.12 | 8497 | 0.351 | 0.0531 | No | ||
| 17 | PGS1 | ENSG00000087157.19 | 8520 | 0.350 | 0.0616 | No | ||
| 18 | AKAP4 | ENSG00000147081.15 | 9429 | 0.319 | 0.0477 | No | ||
| 19 | SYCP1 | ENSG00000198765.12 | 9452 | 0.319 | 0.0554 | No | ||
| 20 | LPIN1 | ENSG00000134324.11 | 10847 | 0.297 | 0.0290 | No | ||
| 21 | DDX25 | ENSG00000109832.14 | 10932 | 0.293 | 0.0345 | No | ||
| 22 | HOXB1 | ENSG00000120094.9 | 11100 | 0.287 | 0.0378 | No | ||
| 23 | GAPDHS | ENSG00000105679.9 | 11950 | 0.273 | 0.0241 | No | ||
| 24 | MAST2 | ENSG00000086015.21 | 11962 | 0.272 | 0.0309 | No | ||
| 25 | ZC2HC1C | ENSG00000119703.14 | 12497 | 0.255 | 0.0244 | No | ||
| 26 | TEKT2 | ENSG00000092850.12 | 12955 | 0.239 | 0.0194 | No | ||
| 27 | CHFR | ENSG00000072609.17 | 14676 | 0.185 | -0.0177 | No | ||
| 28 | ACTL7B | ENSG00000148156.8 | 15954 | 0.146 | -0.0451 | No | ||
| 29 | ELOVL3 | ENSG00000119915.5 | 16089 | 0.143 | -0.0447 | No | ||
| 30 | NPHP1 | ENSG00000144061.12 | 16101 | 0.142 | -0.0413 | No | ||
| 31 | GAD1 | ENSG00000128683.14 | 16125 | 0.142 | -0.0382 | No | ||
| 32 | GRM8 | ENSG00000179603.17 | 16235 | 0.139 | -0.0373 | No | ||
| 33 | ACE | ENSG00000159640.16 | 16370 | 0.135 | -0.0371 | No | ||
| 34 | TULP2 | ENSG00000104804.8 | 16450 | 0.132 | -0.0356 | No | ||
| 35 | IP6K1 | ENSG00000176095.12 | 16499 | 0.131 | -0.0335 | No | ||
| 36 | HSPA2 | ENSG00000126803.9 | 16949 | 0.118 | -0.0414 | No | ||
| 37 | PACRG | ENSG00000112530.11 | 17068 | 0.116 | -0.0413 | No | ||
| 38 | PGK2 | ENSG00000170950.6 | 17549 | 0.102 | -0.0503 | No | ||
| 39 | ADAD1 | ENSG00000164113.11 | 17699 | 0.099 | -0.0514 | No | ||
| 40 | CCNA1 | ENSG00000133101.10 | 17734 | 0.098 | -0.0497 | No | ||
| 41 | TALDO1 | ENSG00000177156.11 | 18425 | 0.081 | -0.0644 | No | ||
| 42 | CLPB | ENSG00000162129.14 | 18742 | 0.072 | -0.0703 | No | ||
| 43 | CLVS1 | ENSG00000177182.11 | 18836 | 0.070 | -0.0708 | No | ||
| 44 | PHKG2 | ENSG00000156873.16 | 18966 | 0.066 | -0.0722 | No | ||
| 45 | PAPOLB | ENSG00000218823.2 | 19127 | 0.062 | -0.0745 | No | ||
| 46 | CSNK2A2 | ENSG00000070770.9 | 19164 | 0.061 | -0.0738 | No | ||
| 47 | DPEP3 | ENSG00000141096.6 | 19167 | 0.061 | -0.0723 | No | ||
| 48 | PARP2 | ENSG00000129484.13 | 19624 | 0.048 | -0.0822 | No | ||
| 49 | PDHA2 | ENSG00000163114.6 | 21711 | -0.006 | -0.1329 | No | ||
| 50 | SIRT1 | ENSG00000096717.12 | 21793 | -0.009 | -0.1346 | No | ||
| 51 | TNP2 | ENSG00000178279.4 | 21864 | -0.011 | -0.1361 | No | ||
| 52 | ACRBP | ENSG00000111644.8 | 22439 | -0.025 | -0.1494 | No | ||
| 53 | AGFG1 | ENSG00000173744.17 | 22700 | -0.031 | -0.1550 | No | ||
| 54 | TOPBP1 | ENSG00000163781.14 | 22850 | -0.035 | -0.1577 | No | ||
| 55 | HSPA4L | ENSG00000164070.12 | 23282 | -0.046 | -0.1670 | No | ||
| 56 | PHF7 | ENSG00000010318.21 | 23590 | -0.055 | -0.1731 | No | ||
| 57 | MTOR | ENSG00000198793.13 | 23734 | -0.059 | -0.1751 | No | ||
| 58 | ZC3H14 | ENSG00000100722.20 | 24498 | -0.077 | -0.1917 | No | ||
| 59 | HBZ | ENSG00000130656.6 | 24610 | -0.080 | -0.1923 | No | ||
| 60 | COIL | ENSG00000121058.5 | 24633 | -0.081 | -0.1908 | No | ||
| 61 | GSTM3 | ENSG00000134202.11 | 24806 | -0.086 | -0.1928 | No | ||
| 62 | ODF1 | ENSG00000155087.4 | 24987 | -0.091 | -0.1948 | No | ||
| 63 | SCG3 | ENSG00000104112.9 | 25353 | -0.101 | -0.2011 | No | ||
| 64 | MAP7 | ENSG00000135525.18 | 25553 | -0.107 | -0.2032 | No | ||
| 65 | DBF4 | ENSG00000006634.8 | 25980 | -0.120 | -0.2105 | No | ||
| 66 | JAM3 | ENSG00000166086.13 | 26245 | -0.127 | -0.2137 | No | ||
| 67 | TLE4 | ENSG00000106829.19 | 26262 | -0.128 | -0.2108 | No | ||
| 68 | BRAF | ENSG00000157764.13 | 26610 | -0.138 | -0.2157 | No | ||
| 69 | NOS1 | ENSG00000089250.19 | 27089 | -0.152 | -0.2235 | No | ||
| 70 | CFTR | ENSG00000001626.15 | 27772 | -0.173 | -0.2356 | No | ||
| 71 | TUBA3C | ENSG00000198033.12 | 27829 | -0.175 | -0.2325 | No | ||
| 72 | MLLT10 | ENSG00000078403.17 | 27899 | -0.176 | -0.2297 | No | ||
| 73 | HSPA1L | ENSG00000204390.10 | 27981 | -0.180 | -0.2270 | No | ||
| 74 | PSMG1 | ENSG00000183527.12 | 28095 | -0.184 | -0.2250 | No | ||
| 75 | CAMK4 | ENSG00000152495.11 | 28125 | -0.184 | -0.2210 | No | ||
| 76 | SLC2A5 | ENSG00000142583.18 | 29013 | -0.212 | -0.2372 | No | ||
| 77 | HIST1H1T | ENSG00000187475.5 | 29600 | -0.230 | -0.2455 | Yes | ||
| 78 | EZH2 | ENSG00000106462.10 | 29613 | -0.231 | -0.2399 | Yes | ||
| 79 | SNAP91 | ENSG00000065609.14 | 29977 | -0.243 | -0.2425 | Yes | ||
| 80 | PEBP1 | ENSG00000089220.5 | 30027 | -0.245 | -0.2374 | Yes | ||
| 81 | CDKN3 | ENSG00000100526.20 | 30484 | -0.259 | -0.2418 | Yes | ||
| 82 | IFT88 | ENSG00000032742.17 | 30640 | -0.264 | -0.2388 | Yes | ||
| 83 | RFC4 | ENSG00000163918.10 | 30672 | -0.265 | -0.2328 | Yes | ||
| 84 | GPR182 | ENSG00000166856.3 | 30724 | -0.267 | -0.2272 | Yes | ||
| 85 | TSN | ENSG00000211460.12 | 31004 | -0.277 | -0.2268 | Yes | ||
| 86 | IL13RA2 | ENSG00000123496.8 | 31074 | -0.280 | -0.2213 | Yes | ||
| 87 | GSG1 | ENSG00000111305.19 | 31123 | -0.282 | -0.2153 | Yes | ||
| 88 | ACRV1 | ENSG00000134940.14 | 31144 | -0.283 | -0.2085 | Yes | ||
| 89 | NAA11 | ENSG00000156269.5 | 31145 | -0.283 | -0.2012 | Yes | ||
| 90 | GMCL1 | ENSG00000087338.5 | 31165 | -0.283 | -0.1944 | Yes | ||
| 91 | PIAS2 | ENSG00000078043.16 | 32124 | -0.301 | -0.2100 | Yes | ||
| 92 | THEG | ENSG00000105549.11 | 32498 | -0.317 | -0.2110 | Yes | ||
| 93 | SHE | ENSG00000169291.10 | 32504 | -0.317 | -0.2029 | Yes | ||
| 94 | RAD17 | ENSG00000152942.19 | 32645 | -0.322 | -0.1981 | Yes | ||
| 95 | VDAC3 | ENSG00000078668.14 | 32993 | -0.336 | -0.1979 | Yes | ||
| 96 | RPL39L | ENSG00000163923.10 | 34191 | -0.349 | -0.2181 | Yes | ||
| 97 | TCP11 | ENSG00000124678.18 | 34710 | -0.356 | -0.2216 | Yes | ||
| 98 | STAM2 | ENSG00000115145.10 | 34753 | -0.359 | -0.2134 | Yes | ||
| 99 | SCG5 | ENSG00000166922.8 | 35094 | -0.368 | -0.2122 | Yes | ||
| 100 | STRBP | ENSG00000165209.19 | 35121 | -0.370 | -0.2033 | Yes | ||
| 101 | NEFH | ENSG00000100285.10 | 35194 | -0.374 | -0.1955 | Yes | ||
| 102 | PRKAR2A | ENSG00000114302.16 | 35272 | -0.377 | -0.1877 | Yes | ||
| 103 | TKTL1 | ENSG00000007350.17 | 36144 | -0.411 | -0.1983 | Yes | ||
| 104 | CCNB2 | ENSG00000157456.8 | 36613 | -0.432 | -0.1986 | Yes | ||
| 105 | KIF2C | ENSG00000142945.13 | 36852 | -0.443 | -0.1931 | Yes | ||
| 106 | CNIH2 | ENSG00000174871.11 | 37049 | -0.455 | -0.1861 | Yes | ||
| 107 | MTNR1A | ENSG00000168412.6 | 37417 | -0.479 | -0.1828 | Yes | ||
| 108 | DMC1 | ENSG00000100206.10 | 37480 | -0.483 | -0.1719 | Yes | ||
| 109 | CCT6B | ENSG00000132141.14 | 37660 | -0.499 | -0.1634 | Yes | ||
| 110 | BUB1 | ENSG00000169679.15 | 37763 | -0.508 | -0.1528 | Yes | ||
| 111 | SLC12A2 | ENSG00000064651.14 | 37771 | -0.510 | -0.1399 | Yes | ||
| 112 | NEK2 | ENSG00000117650.13 | 37958 | -0.525 | -0.1310 | Yes | ||
| 113 | NF2 | ENSG00000186575.18 | 37996 | -0.527 | -0.1183 | Yes | ||
| 114 | TTK | ENSG00000112742.10 | 38213 | -0.544 | -0.1096 | Yes | ||
| 115 | DNAJB8 | ENSG00000179407.3 | 38332 | -0.555 | -0.0982 | Yes | ||
| 116 | AURKA | ENSG00000087586.17 | 38638 | -0.565 | -0.0911 | Yes | ||
| 117 | NCAPH | ENSG00000121152.10 | 38902 | -0.590 | -0.0824 | Yes | ||
| 118 | OAZ3 | ENSG00000143450.16 | 39039 | -0.605 | -0.0701 | Yes | ||
| 119 | CDK1 | ENSG00000170312.16 | 39453 | -0.632 | -0.0640 | Yes | ||
| 120 | IDE | ENSG00000119912.17 | 39758 | -0.668 | -0.0542 | Yes | ||
| 121 | ARL4A | ENSG00000122644.13 | 40051 | -0.722 | -0.0428 | Yes | ||
| 122 | GFI1 | ENSG00000162676.12 | 40452 | -0.825 | -0.0313 | Yes | ||
| 123 | MLF1 | ENSG00000178053.19 | 40547 | -0.856 | -0.0116 | Yes | ||
| 124 | TNNI3 | ENSG00000129991.13 | 40911 | -1.033 | 0.0061 | Yes |