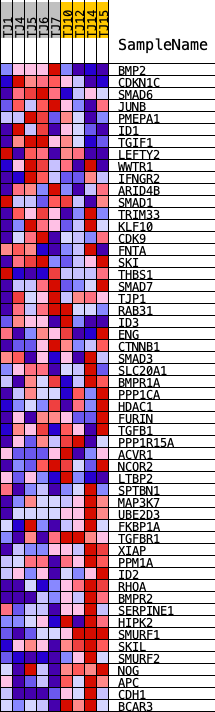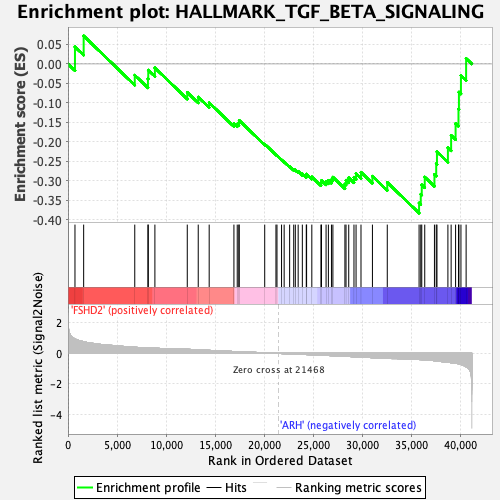
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_TGF_BETA_SIGNALING |
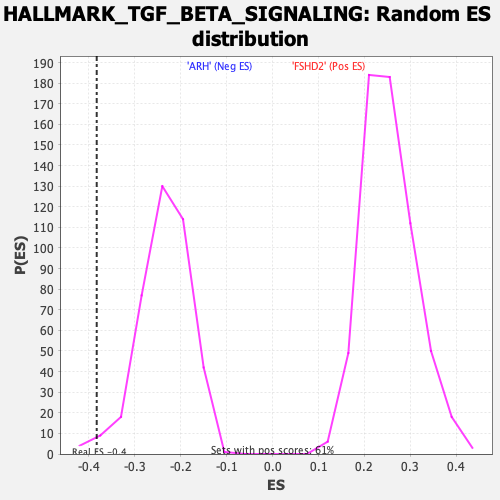
| Enrichment Score (ES) | -0.38293803 |
| Normalized Enrichment Score (NES) | -1.6288116 |
| Nominal p-value | 0.015189873 |
| FDR q-value | 0.010251815 |
| FWER p-Value | 0.054 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | BMP2 | ENSG00000125845.7 | 710 | 0.936 | 0.0439 | No | ||
| 2 | CDKN1C | ENSG00000129757.13 | 1596 | 0.759 | 0.0720 | No | ||
| 3 | SMAD6 | ENSG00000137834.15 | 6805 | 0.396 | -0.0288 | No | ||
| 4 | JUNB | ENSG00000171223.6 | 8148 | 0.355 | -0.0382 | No | ||
| 5 | PMEPA1 | ENSG00000124225.16 | 8181 | 0.354 | -0.0158 | No | ||
| 6 | ID1 | ENSG00000125968.9 | 8857 | 0.341 | -0.0100 | No | ||
| 7 | TGIF1 | ENSG00000177426.20 | 12157 | 0.266 | -0.0728 | No | ||
| 8 | LEFTY2 | ENSG00000143768.13 | 13275 | 0.229 | -0.0850 | No | ||
| 9 | WWTR1 | ENSG00000018408.14 | 14391 | 0.194 | -0.0995 | No | ||
| 10 | IFNGR2 | ENSG00000159128.14 | 16912 | 0.119 | -0.1530 | No | ||
| 11 | ARID4B | ENSG00000054267.22 | 17250 | 0.111 | -0.1539 | No | ||
| 12 | SMAD1 | ENSG00000170365.10 | 17415 | 0.107 | -0.1510 | No | ||
| 13 | TRIM33 | ENSG00000197323.12 | 17449 | 0.105 | -0.1449 | No | ||
| 14 | KLF10 | ENSG00000155090.15 | 20051 | 0.038 | -0.2057 | No | ||
| 15 | CDK9 | ENSG00000136807.14 | 21197 | 0.007 | -0.2331 | No | ||
| 16 | FNTA | ENSG00000168522.13 | 21301 | 0.004 | -0.2353 | No | ||
| 17 | SKI | ENSG00000157933.10 | 21767 | -0.008 | -0.2461 | No | ||
| 18 | THBS1 | ENSG00000137801.11 | 22033 | -0.015 | -0.2515 | No | ||
| 19 | SMAD7 | ENSG00000101665.9 | 22592 | -0.028 | -0.2633 | No | ||
| 20 | TJP1 | ENSG00000104067.16 | 23008 | -0.040 | -0.2708 | No | ||
| 21 | RAB31 | ENSG00000168461.13 | 23170 | -0.044 | -0.2718 | No | ||
| 22 | ID3 | ENSG00000117318.9 | 23461 | -0.051 | -0.2755 | No | ||
| 23 | ENG | ENSG00000106991.13 | 23886 | -0.063 | -0.2817 | No | ||
| 24 | CTNNB1 | ENSG00000168036.18 | 24288 | -0.073 | -0.2867 | No | ||
| 25 | SMAD3 | ENSG00000166949.16 | 24321 | -0.074 | -0.2827 | No | ||
| 26 | SLC20A1 | ENSG00000144136.11 | 24853 | -0.087 | -0.2899 | No | ||
| 27 | BMPR1A | ENSG00000107779.13 | 25782 | -0.113 | -0.3051 | No | ||
| 28 | PPP1CA | ENSG00000172531.15 | 25840 | -0.115 | -0.2990 | No | ||
| 29 | HDAC1 | ENSG00000116478.12 | 26306 | -0.129 | -0.3019 | No | ||
| 30 | FURIN | ENSG00000140564.12 | 26542 | -0.136 | -0.2987 | No | ||
| 31 | TGFB1 | ENSG00000105329.10 | 26855 | -0.145 | -0.2968 | No | ||
| 32 | PPP1R15A | ENSG00000087074.8 | 27000 | -0.149 | -0.2905 | No | ||
| 33 | ACVR1 | ENSG00000115170.14 | 28226 | -0.187 | -0.3081 | No | ||
| 34 | NCOR2 | ENSG00000196498.13 | 28339 | -0.191 | -0.2984 | No | ||
| 35 | LTBP2 | ENSG00000119681.12 | 28615 | -0.200 | -0.2920 | No | ||
| 36 | SPTBN1 | ENSG00000115306.16 | 29149 | -0.216 | -0.2908 | No | ||
| 37 | MAP3K7 | ENSG00000135341.18 | 29362 | -0.223 | -0.2814 | No | ||
| 38 | UBE2D3 | ENSG00000109332.20 | 29865 | -0.240 | -0.2779 | No | ||
| 39 | FKBP1A | ENSG00000088832.17 | 31031 | -0.278 | -0.2881 | No | ||
| 40 | TGFBR1 | ENSG00000106799.13 | 32544 | -0.318 | -0.3041 | No | ||
| 41 | XIAP | ENSG00000101966.12 | 35786 | -0.401 | -0.3567 | Yes | ||
| 42 | PPM1A | ENSG00000100614.18 | 35958 | -0.404 | -0.3345 | Yes | ||
| 43 | ID2 | ENSG00000115738.10 | 36059 | -0.406 | -0.3103 | Yes | ||
| 44 | RHOA | ENSG00000067560.12 | 36363 | -0.419 | -0.2903 | Yes | ||
| 45 | BMPR2 | ENSG00000204217.15 | 37352 | -0.475 | -0.2833 | Yes | ||
| 46 | SERPINE1 | ENSG00000106366.8 | 37533 | -0.487 | -0.2559 | Yes | ||
| 47 | HIPK2 | ENSG00000064393.16 | 37609 | -0.494 | -0.2254 | Yes | ||
| 48 | SMURF1 | ENSG00000198742.9 | 38724 | -0.573 | -0.2150 | Yes | ||
| 49 | SKIL | ENSG00000136603.14 | 39053 | -0.607 | -0.1833 | Yes | ||
| 50 | SMURF2 | ENSG00000108854.16 | 39508 | -0.636 | -0.1528 | Yes | ||
| 51 | NOG | ENSG00000183691.5 | 39803 | -0.675 | -0.1159 | Yes | ||
| 52 | APC | ENSG00000134982.17 | 39848 | -0.685 | -0.0722 | Yes | ||
| 53 | CDH1 | ENSG00000039068.19 | 40053 | -0.722 | -0.0299 | Yes | ||
| 54 | BCAR3 | ENSG00000137936.18 | 40581 | -0.870 | 0.0141 | Yes |