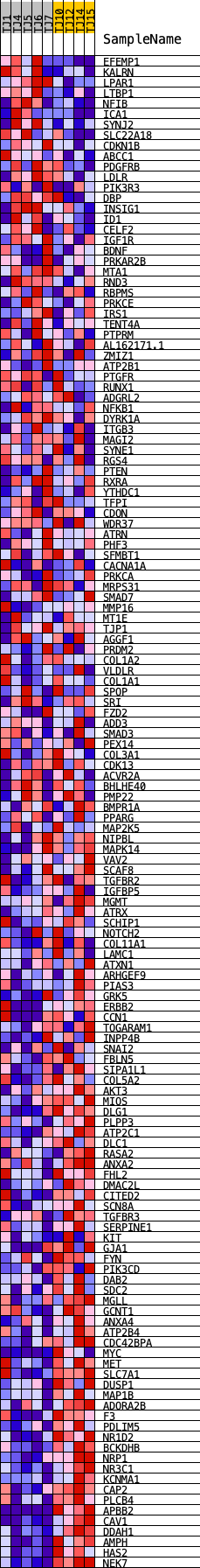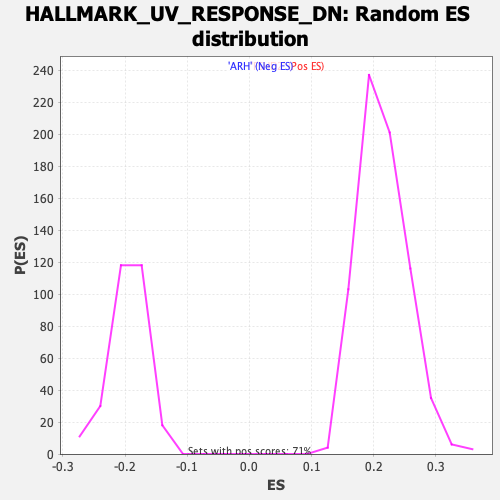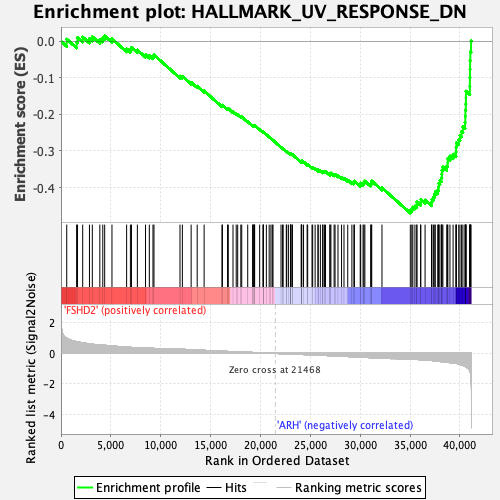
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_UV_RESPONSE_DN |
| Enrichment Score (ES) | -0.47000104 |
| Normalized Enrichment Score (NES) | -2.407154 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EFEMP1 | ENSG00000115380.20 | 575 | 0.988 | 0.0060 | No | ||
| 2 | KALRN | ENSG00000160145.15 | 1565 | 0.764 | -0.0027 | No | ||
| 3 | LPAR1 | ENSG00000198121.13 | 1657 | 0.752 | 0.0104 | No | ||
| 4 | LTBP1 | ENSG00000049323.16 | 2175 | 0.686 | 0.0116 | No | ||
| 5 | NFIB | ENSG00000147862.17 | 2853 | 0.618 | 0.0076 | No | ||
| 6 | ICA1 | ENSG00000003147.19 | 3140 | 0.596 | 0.0127 | No | ||
| 7 | SYNJ2 | ENSG00000078269.15 | 3893 | 0.545 | 0.0054 | No | ||
| 8 | SLC22A18 | ENSG00000110628.15 | 4190 | 0.531 | 0.0090 | No | ||
| 9 | CDKN1B | ENSG00000111276.11 | 4380 | 0.521 | 0.0149 | No | ||
| 10 | ABCC1 | ENSG00000103222.19 | 5110 | 0.483 | 0.0069 | No | ||
| 11 | PDGFRB | ENSG00000113721.14 | 6580 | 0.405 | -0.0207 | No | ||
| 12 | LDLR | ENSG00000130164.13 | 6959 | 0.391 | -0.0220 | No | ||
| 13 | PIK3R3 | ENSG00000117461.15 | 7066 | 0.387 | -0.0168 | No | ||
| 14 | DBP | ENSG00000105516.11 | 7657 | 0.364 | -0.0238 | No | ||
| 15 | INSIG1 | ENSG00000186480.13 | 8475 | 0.352 | -0.0366 | No | ||
| 16 | ID1 | ENSG00000125968.9 | 8857 | 0.341 | -0.0390 | No | ||
| 17 | CELF2 | ENSG00000048740.18 | 9223 | 0.326 | -0.0413 | No | ||
| 18 | IGF1R | ENSG00000140443.15 | 9322 | 0.322 | -0.0371 | No | ||
| 19 | BDNF | ENSG00000176697.19 | 11933 | 0.274 | -0.0952 | No | ||
| 20 | PRKAR2B | ENSG00000005249.13 | 12170 | 0.265 | -0.0956 | No | ||
| 21 | MTA1 | ENSG00000182979.18 | 13050 | 0.236 | -0.1122 | No | ||
| 22 | RND3 | ENSG00000115963.13 | 13657 | 0.217 | -0.1226 | No | ||
| 23 | RBPMS | ENSG00000157110.16 | 14352 | 0.196 | -0.1356 | No | ||
| 24 | PRKCE | ENSG00000171132.14 | 16141 | 0.141 | -0.1763 | No | ||
| 25 | IRS1 | ENSG00000169047.5 | 16185 | 0.140 | -0.1745 | No | ||
| 26 | TENT4A | ENSG00000112941.14 | 16700 | 0.125 | -0.1845 | No | ||
| 27 | PTPRM | ENSG00000173482.16 | 16775 | 0.123 | -0.1838 | No | ||
| 28 | AL162171.1 | ENSG00000258789.1 | 17248 | 0.111 | -0.1931 | No | ||
| 29 | ZMIZ1 | ENSG00000108175.17 | 17570 | 0.102 | -0.1989 | No | ||
| 30 | ATP2B1 | ENSG00000070961.15 | 17717 | 0.099 | -0.2004 | No | ||
| 31 | PTGFR | ENSG00000122420.10 | 18031 | 0.090 | -0.2062 | No | ||
| 32 | RUNX1 | ENSG00000159216.18 | 18123 | 0.087 | -0.2067 | No | ||
| 33 | ADGRL2 | ENSG00000117114.19 | 18723 | 0.073 | -0.2198 | No | ||
| 34 | NFKB1 | ENSG00000109320.13 | 19218 | 0.059 | -0.2307 | No | ||
| 35 | DYRK1A | ENSG00000157540.21 | 19241 | 0.058 | -0.2300 | No | ||
| 36 | ITGB3 | ENSG00000259207.7 | 19335 | 0.055 | -0.2312 | No | ||
| 37 | MAGI2 | ENSG00000187391.21 | 19355 | 0.055 | -0.2305 | No | ||
| 38 | SYNE1 | ENSG00000131018.24 | 19377 | 0.054 | -0.2299 | No | ||
| 39 | RGS4 | ENSG00000117152.13 | 19418 | 0.053 | -0.2298 | No | ||
| 40 | PTEN | ENSG00000171862.11 | 19922 | 0.040 | -0.2413 | No | ||
| 41 | RXRA | ENSG00000186350.12 | 20264 | 0.032 | -0.2489 | No | ||
| 42 | YTHDC1 | ENSG00000083896.12 | 20287 | 0.032 | -0.2488 | No | ||
| 43 | TFPI | ENSG00000003436.16 | 20290 | 0.032 | -0.2483 | No | ||
| 44 | CDON | ENSG00000064309.14 | 20593 | 0.023 | -0.2552 | No | ||
| 45 | WDR37 | ENSG00000047056.16 | 20874 | 0.016 | -0.2617 | No | ||
| 46 | ATRN | ENSG00000088812.18 | 21050 | 0.010 | -0.2657 | No | ||
| 47 | PHF3 | ENSG00000118482.11 | 21189 | 0.007 | -0.2689 | No | ||
| 48 | SFMBT1 | ENSG00000163935.14 | 21261 | 0.005 | -0.2706 | No | ||
| 49 | CACNA1A | ENSG00000141837.20 | 22062 | -0.016 | -0.2898 | No | ||
| 50 | PRKCA | ENSG00000154229.12 | 22219 | -0.019 | -0.2932 | No | ||
| 51 | MRPS31 | ENSG00000102738.8 | 22274 | -0.020 | -0.2941 | No | ||
| 52 | SMAD7 | ENSG00000101665.9 | 22592 | -0.028 | -0.3013 | No | ||
| 53 | MMP16 | ENSG00000156103.16 | 22632 | -0.029 | -0.3016 | No | ||
| 54 | MT1E | ENSG00000169715.15 | 22802 | -0.033 | -0.3051 | No | ||
| 55 | TJP1 | ENSG00000104067.16 | 23008 | -0.040 | -0.3093 | No | ||
| 56 | AGGF1 | ENSG00000164252.13 | 23018 | -0.040 | -0.3087 | No | ||
| 57 | PRDM2 | ENSG00000116731.22 | 23037 | -0.040 | -0.3083 | No | ||
| 58 | COL1A2 | ENSG00000164692.17 | 23053 | -0.041 | -0.3078 | No | ||
| 59 | VLDLR | ENSG00000147852.16 | 23181 | -0.045 | -0.3100 | No | ||
| 60 | COL1A1 | ENSG00000108821.13 | 23206 | -0.045 | -0.3097 | No | ||
| 61 | SPOP | ENSG00000121067.19 | 24083 | -0.066 | -0.3297 | No | ||
| 62 | SRI | ENSG00000075142.13 | 24123 | -0.067 | -0.3293 | No | ||
| 63 | FZD2 | ENSG00000180340.7 | 24126 | -0.067 | -0.3280 | No | ||
| 64 | ADD3 | ENSG00000148700.15 | 24136 | -0.068 | -0.3268 | No | ||
| 65 | SMAD3 | ENSG00000166949.16 | 24321 | -0.074 | -0.3298 | No | ||
| 66 | PEX14 | ENSG00000142655.13 | 24680 | -0.082 | -0.3369 | No | ||
| 67 | COL3A1 | ENSG00000168542.14 | 24737 | -0.084 | -0.3365 | No | ||
| 68 | CDK13 | ENSG00000065883.16 | 25171 | -0.095 | -0.3452 | No | ||
| 69 | ACVR2A | ENSG00000121989.15 | 25237 | -0.097 | -0.3448 | No | ||
| 70 | BHLHE40 | ENSG00000134107.5 | 25483 | -0.105 | -0.3487 | No | ||
| 71 | PMP22 | ENSG00000109099.15 | 25773 | -0.113 | -0.3534 | No | ||
| 72 | BMPR1A | ENSG00000107779.13 | 25782 | -0.113 | -0.3513 | No | ||
| 73 | PPARG | ENSG00000132170.21 | 26008 | -0.121 | -0.3544 | No | ||
| 74 | MAP2K5 | ENSG00000137764.20 | 26238 | -0.127 | -0.3574 | No | ||
| 75 | NIPBL | ENSG00000164190.19 | 26266 | -0.128 | -0.3554 | No | ||
| 76 | MAPK14 | ENSG00000112062.10 | 26416 | -0.133 | -0.3564 | No | ||
| 77 | VAV2 | ENSG00000160293.17 | 26522 | -0.136 | -0.3562 | No | ||
| 78 | SCAF8 | ENSG00000213079.9 | 26968 | -0.148 | -0.3640 | No | ||
| 79 | TGFBR2 | ENSG00000163513.18 | 26973 | -0.148 | -0.3611 | No | ||
| 80 | IGFBP5 | ENSG00000115461.5 | 27066 | -0.151 | -0.3603 | No | ||
| 81 | MGMT | ENSG00000170430.10 | 27385 | -0.160 | -0.3648 | No | ||
| 82 | ATRX | ENSG00000085224.22 | 27481 | -0.164 | -0.3638 | No | ||
| 83 | SCHIP1 | ENSG00000151967.18 | 27775 | -0.173 | -0.3675 | No | ||
| 84 | NOTCH2 | ENSG00000134250.20 | 28133 | -0.185 | -0.3724 | No | ||
| 85 | COL11A1 | ENSG00000060718.22 | 28377 | -0.192 | -0.3745 | No | ||
| 86 | LAMC1 | ENSG00000135862.6 | 28753 | -0.203 | -0.3795 | No | ||
| 87 | ATXN1 | ENSG00000124788.18 | 29169 | -0.217 | -0.3852 | No | ||
| 88 | ARHGEF9 | ENSG00000131089.16 | 29371 | -0.224 | -0.3856 | No | ||
| 89 | PIAS3 | ENSG00000131788.16 | 29423 | -0.225 | -0.3823 | No | ||
| 90 | GRK5 | ENSG00000198873.12 | 30006 | -0.244 | -0.3916 | No | ||
| 91 | ERBB2 | ENSG00000141736.13 | 30053 | -0.246 | -0.3877 | No | ||
| 92 | CCN1 | ENSG00000142871.17 | 30290 | -0.253 | -0.3883 | No | ||
| 93 | TOGARAM1 | ENSG00000198718.13 | 30375 | -0.256 | -0.3852 | No | ||
| 94 | INPP4B | ENSG00000109452.12 | 30469 | -0.259 | -0.3822 | No | ||
| 95 | SNAI2 | ENSG00000019549.13 | 31053 | -0.279 | -0.3908 | No | ||
| 96 | FBLN5 | ENSG00000140092.14 | 31116 | -0.281 | -0.3866 | No | ||
| 97 | SIPA1L1 | ENSG00000197555.9 | 31158 | -0.283 | -0.3819 | No | ||
| 98 | COL5A2 | ENSG00000204262.14 | 32186 | -0.304 | -0.4007 | No | ||
| 99 | AKT3 | ENSG00000117020.18 | 35028 | -0.366 | -0.4626 | Yes | ||
| 100 | MIOS | ENSG00000164654.16 | 35177 | -0.373 | -0.4587 | Yes | ||
| 101 | DLG1 | ENSG00000075711.21 | 35280 | -0.377 | -0.4535 | Yes | ||
| 102 | PLPP3 | ENSG00000162407.9 | 35490 | -0.388 | -0.4507 | Yes | ||
| 103 | ATP2C1 | ENSG00000017260.19 | 35680 | -0.397 | -0.4473 | Yes | ||
| 104 | DLC1 | ENSG00000164741.15 | 35687 | -0.397 | -0.4394 | Yes | ||
| 105 | RASA2 | ENSG00000155903.12 | 36064 | -0.407 | -0.4404 | Yes | ||
| 106 | ANXA2 | ENSG00000182718.16 | 36074 | -0.407 | -0.4323 | Yes | ||
| 107 | FHL2 | ENSG00000115641.19 | 36508 | -0.427 | -0.4342 | Yes | ||
| 108 | DMAC2L | ENSG00000125375.15 | 37153 | -0.461 | -0.4406 | Yes | ||
| 109 | CITED2 | ENSG00000164442.10 | 37189 | -0.463 | -0.4321 | Yes | ||
| 110 | SCN8A | ENSG00000196876.17 | 37337 | -0.474 | -0.4261 | Yes | ||
| 111 | TGFBR3 | ENSG00000069702.11 | 37426 | -0.479 | -0.4185 | Yes | ||
| 112 | SERPINE1 | ENSG00000106366.8 | 37533 | -0.487 | -0.4113 | Yes | ||
| 113 | KIT | ENSG00000157404.15 | 37779 | -0.511 | -0.4069 | Yes | ||
| 114 | GJA1 | ENSG00000152661.9 | 37853 | -0.517 | -0.3982 | Yes | ||
| 115 | FYN | ENSG00000010810.17 | 37878 | -0.518 | -0.3883 | Yes | ||
| 116 | PIK3CD | ENSG00000171608.16 | 37991 | -0.527 | -0.3804 | Yes | ||
| 117 | DAB2 | ENSG00000153071.15 | 38148 | -0.538 | -0.3733 | Yes | ||
| 118 | SDC2 | ENSG00000169439.12 | 38187 | -0.541 | -0.3632 | Yes | ||
| 119 | MGLL | ENSG00000074416.14 | 38196 | -0.542 | -0.3525 | Yes | ||
| 120 | GCNT1 | ENSG00000187210.14 | 38293 | -0.551 | -0.3437 | Yes | ||
| 121 | ANXA4 | ENSG00000196975.16 | 38680 | -0.569 | -0.3416 | Yes | ||
| 122 | ATP2B4 | ENSG00000058668.14 | 38776 | -0.578 | -0.3322 | Yes | ||
| 123 | CDC42BPA | ENSG00000143776.18 | 38777 | -0.578 | -0.3205 | Yes | ||
| 124 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.3138 | Yes | ||
| 125 | MET | ENSG00000105976.15 | 39317 | -0.624 | -0.3088 | Yes | ||
| 126 | SLC7A1 | ENSG00000139514.13 | 39608 | -0.647 | -0.3027 | Yes | ||
| 127 | DUSP1 | ENSG00000120129.6 | 39618 | -0.648 | -0.2898 | Yes | ||
| 128 | MAP1B | ENSG00000131711.15 | 39648 | -0.653 | -0.2773 | Yes | ||
| 129 | ADORA2B | ENSG00000170425.3 | 39882 | -0.692 | -0.2690 | Yes | ||
| 130 | F3 | ENSG00000117525.14 | 40005 | -0.714 | -0.2575 | Yes | ||
| 131 | PDLIM5 | ENSG00000163110.15 | 40169 | -0.746 | -0.2464 | Yes | ||
| 132 | NR1D2 | ENSG00000174738.13 | 40302 | -0.778 | -0.2339 | Yes | ||
| 133 | BCKDHB | ENSG00000083123.15 | 40514 | -0.843 | -0.2219 | Yes | ||
| 134 | NRP1 | ENSG00000099250.18 | 40541 | -0.854 | -0.2053 | Yes | ||
| 135 | NR3C1 | ENSG00000113580.15 | 40550 | -0.857 | -0.1881 | Yes | ||
| 136 | KCNMA1 | ENSG00000156113.23 | 40594 | -0.873 | -0.1715 | Yes | ||
| 137 | CAP2 | ENSG00000112186.12 | 40604 | -0.877 | -0.1540 | Yes | ||
| 138 | PLCB4 | ENSG00000101333.16 | 40607 | -0.879 | -0.1362 | Yes | ||
| 139 | APBB2 | ENSG00000163697.17 | 40996 | -1.135 | -0.1227 | Yes | ||
| 140 | CAV1 | ENSG00000105974.12 | 41003 | -1.149 | -0.0996 | Yes | ||
| 141 | DDAH1 | ENSG00000153904.20 | 41014 | -1.158 | -0.0764 | Yes | ||
| 142 | AMPH | ENSG00000078053.17 | 41023 | -1.183 | -0.0526 | Yes | ||
| 143 | HAS2 | ENSG00000170961.7 | 41046 | -1.217 | -0.0285 | Yes | ||
| 144 | NEK7 | ENSG00000151414.15 | 41118 | -1.546 | 0.0010 | Yes |