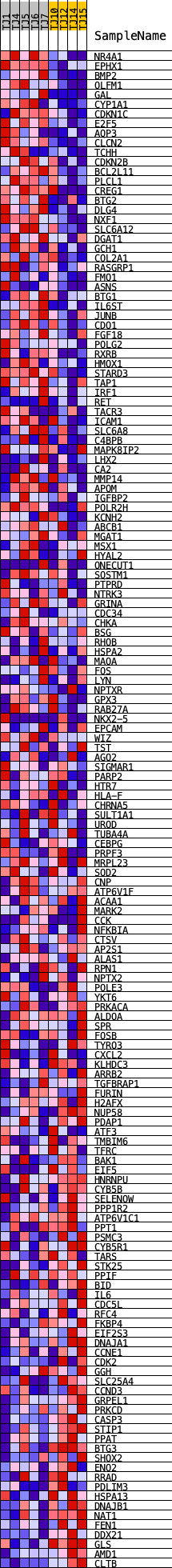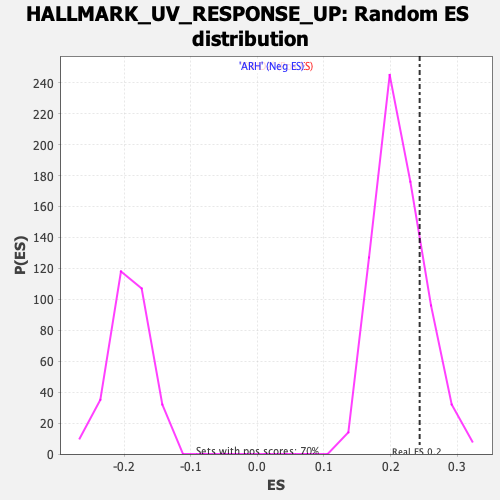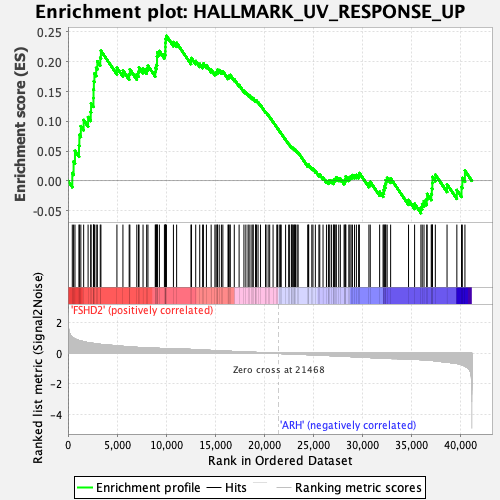
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_UV_RESPONSE_UP |
| Enrichment Score (ES) | 0.24348357 |
| Normalized Enrichment Score (NES) | 1.141802 |
| Nominal p-value | 0.1991404 |
| FDR q-value | 0.26719517 |
| FWER p-Value | 0.999 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NR4A1 | ENSG00000123358.20 | 437 | 1.041 | 0.0131 | Yes | ||
| 2 | EPHX1 | ENSG00000143819.12 | 570 | 0.990 | 0.0324 | Yes | ||
| 3 | BMP2 | ENSG00000125845.7 | 710 | 0.936 | 0.0503 | Yes | ||
| 4 | OLFM1 | ENSG00000130558.19 | 1121 | 0.833 | 0.0593 | Yes | ||
| 5 | GAL | ENSG00000069482.7 | 1157 | 0.825 | 0.0772 | Yes | ||
| 6 | CYP1A1 | ENSG00000140465.14 | 1309 | 0.801 | 0.0918 | Yes | ||
| 7 | CDKN1C | ENSG00000129757.13 | 1596 | 0.759 | 0.1021 | Yes | ||
| 8 | E2F5 | ENSG00000133740.11 | 2048 | 0.699 | 0.1070 | Yes | ||
| 9 | AQP3 | ENSG00000165272.16 | 2318 | 0.670 | 0.1157 | Yes | ||
| 10 | CLCN2 | ENSG00000114859.16 | 2351 | 0.667 | 0.1301 | Yes | ||
| 11 | TCHH | ENSG00000159450.12 | 2593 | 0.644 | 0.1389 | Yes | ||
| 12 | CDKN2B | ENSG00000147883.12 | 2601 | 0.644 | 0.1534 | Yes | ||
| 13 | BCL2L11 | ENSG00000153094.23 | 2637 | 0.640 | 0.1671 | Yes | ||
| 14 | PLCL1 | ENSG00000115896.16 | 2691 | 0.634 | 0.1803 | Yes | ||
| 15 | CREG1 | ENSG00000143162.9 | 2871 | 0.616 | 0.1899 | Yes | ||
| 16 | BTG2 | ENSG00000159388.6 | 2993 | 0.606 | 0.2008 | Yes | ||
| 17 | DLG4 | ENSG00000132535.19 | 3287 | 0.584 | 0.2069 | Yes | ||
| 18 | NXF1 | ENSG00000162231.14 | 3355 | 0.580 | 0.2185 | Yes | ||
| 19 | SLC6A12 | ENSG00000111181.12 | 4987 | 0.492 | 0.1899 | Yes | ||
| 20 | DGAT1 | ENSG00000185000.12 | 5599 | 0.453 | 0.1853 | Yes | ||
| 21 | GCH1 | ENSG00000131979.19 | 6236 | 0.421 | 0.1794 | Yes | ||
| 22 | COL2A1 | ENSG00000139219.19 | 6306 | 0.417 | 0.1872 | Yes | ||
| 23 | RASGRP1 | ENSG00000172575.12 | 7013 | 0.389 | 0.1789 | Yes | ||
| 24 | FMO1 | ENSG00000010932.17 | 7182 | 0.381 | 0.1835 | Yes | ||
| 25 | ASNS | ENSG00000070669.17 | 7241 | 0.378 | 0.1907 | Yes | ||
| 26 | BTG1 | ENSG00000133639.5 | 7649 | 0.365 | 0.1890 | Yes | ||
| 27 | IL6ST | ENSG00000134352.20 | 8019 | 0.360 | 0.1882 | Yes | ||
| 28 | JUNB | ENSG00000171223.6 | 8148 | 0.355 | 0.1932 | Yes | ||
| 29 | CDO1 | ENSG00000129596.5 | 8894 | 0.340 | 0.1828 | Yes | ||
| 30 | FGF18 | ENSG00000156427.8 | 8917 | 0.339 | 0.1899 | Yes | ||
| 31 | POLG2 | ENSG00000256525.8 | 9023 | 0.334 | 0.1950 | Yes | ||
| 32 | RXRB | ENSG00000204231.10 | 9074 | 0.332 | 0.2013 | Yes | ||
| 33 | HMOX1 | ENSG00000100292.17 | 9075 | 0.332 | 0.2089 | Yes | ||
| 34 | STARD3 | ENSG00000131748.16 | 9106 | 0.331 | 0.2157 | Yes | ||
| 35 | TAP1 | ENSG00000168394.11 | 9314 | 0.323 | 0.2180 | Yes | ||
| 36 | IRF1 | ENSG00000125347.14 | 9828 | 0.318 | 0.2127 | Yes | ||
| 37 | RET | ENSG00000165731.20 | 9916 | 0.315 | 0.2178 | Yes | ||
| 38 | TACR3 | ENSG00000169836.5 | 9919 | 0.315 | 0.2249 | Yes | ||
| 39 | ICAM1 | ENSG00000090339.9 | 9924 | 0.314 | 0.2320 | Yes | ||
| 40 | SLC6A8 | ENSG00000130821.16 | 9951 | 0.313 | 0.2385 | Yes | ||
| 41 | C4BPB | ENSG00000123843.13 | 10036 | 0.310 | 0.2435 | Yes | ||
| 42 | MAPK8IP2 | ENSG00000008735.14 | 10746 | 0.301 | 0.2330 | No | ||
| 43 | LHX2 | ENSG00000106689.11 | 11073 | 0.288 | 0.2317 | No | ||
| 44 | CA2 | ENSG00000104267.10 | 12533 | 0.253 | 0.2018 | No | ||
| 45 | MMP14 | ENSG00000157227.13 | 12590 | 0.252 | 0.2062 | No | ||
| 46 | APOM | ENSG00000204444.11 | 13018 | 0.237 | 0.2012 | No | ||
| 47 | IGFBP2 | ENSG00000115457.10 | 13426 | 0.224 | 0.1964 | No | ||
| 48 | POLR2H | ENSG00000163882.9 | 13722 | 0.215 | 0.1941 | No | ||
| 49 | KCNH2 | ENSG00000055118.15 | 13802 | 0.213 | 0.1970 | No | ||
| 50 | ABCB1 | ENSG00000085563.14 | 14109 | 0.202 | 0.1941 | No | ||
| 51 | MGAT1 | ENSG00000131446.16 | 14591 | 0.187 | 0.1867 | No | ||
| 52 | MSX1 | ENSG00000163132.8 | 14975 | 0.174 | 0.1813 | No | ||
| 53 | HYAL2 | ENSG00000068001.14 | 15095 | 0.171 | 0.1823 | No | ||
| 54 | ONECUT1 | ENSG00000169856.9 | 15209 | 0.168 | 0.1834 | No | ||
| 55 | SQSTM1 | ENSG00000161011.20 | 15233 | 0.167 | 0.1866 | No | ||
| 56 | PTPRD | ENSG00000153707.17 | 15413 | 0.161 | 0.1859 | No | ||
| 57 | NTRK3 | ENSG00000140538.16 | 15638 | 0.155 | 0.1840 | No | ||
| 58 | GRINA | ENSG00000178719.17 | 15785 | 0.151 | 0.1838 | No | ||
| 59 | CDC34 | ENSG00000099804.9 | 16311 | 0.137 | 0.1742 | No | ||
| 60 | CHKA | ENSG00000110721.12 | 16361 | 0.135 | 0.1760 | No | ||
| 61 | BSG | ENSG00000172270.19 | 16494 | 0.131 | 0.1758 | No | ||
| 62 | RHOB | ENSG00000143878.10 | 16535 | 0.130 | 0.1778 | No | ||
| 63 | HSPA2 | ENSG00000126803.9 | 16949 | 0.118 | 0.1704 | No | ||
| 64 | MAOA | ENSG00000189221.9 | 17441 | 0.106 | 0.1608 | No | ||
| 65 | FOS | ENSG00000170345.10 | 17933 | 0.092 | 0.1510 | No | ||
| 66 | LYN | ENSG00000254087.8 | 18130 | 0.087 | 0.1482 | No | ||
| 67 | NPTXR | ENSG00000221890.4 | 18353 | 0.082 | 0.1446 | No | ||
| 68 | GPX3 | ENSG00000211445.12 | 18462 | 0.080 | 0.1438 | No | ||
| 69 | RAB27A | ENSG00000069974.16 | 18661 | 0.075 | 0.1407 | No | ||
| 70 | NKX2-5 | ENSG00000183072.10 | 18847 | 0.069 | 0.1378 | No | ||
| 71 | EPCAM | ENSG00000119888.10 | 18870 | 0.069 | 0.1388 | No | ||
| 72 | WIZ | ENSG00000011451.19 | 19118 | 0.062 | 0.1342 | No | ||
| 73 | TST | ENSG00000128311.14 | 19138 | 0.061 | 0.1351 | No | ||
| 74 | AGO2 | ENSG00000123908.12 | 19222 | 0.059 | 0.1344 | No | ||
| 75 | SIGMAR1 | ENSG00000147955.17 | 19374 | 0.054 | 0.1320 | No | ||
| 76 | PARP2 | ENSG00000129484.13 | 19624 | 0.048 | 0.1270 | No | ||
| 77 | HTR7 | ENSG00000148680.16 | 20138 | 0.036 | 0.1153 | No | ||
| 78 | HLA-F | ENSG00000204642.14 | 20184 | 0.034 | 0.1150 | No | ||
| 79 | CHRNA5 | ENSG00000169684.13 | 20402 | 0.028 | 0.1103 | No | ||
| 80 | SULT1A1 | ENSG00000196502.11 | 20558 | 0.023 | 0.1071 | No | ||
| 81 | UROD | ENSG00000126088.14 | 20903 | 0.015 | 0.0990 | No | ||
| 82 | TUBA4A | ENSG00000127824.14 | 21284 | 0.004 | 0.0899 | No | ||
| 83 | CEBPG | ENSG00000153879.9 | 21361 | 0.002 | 0.0881 | No | ||
| 84 | PRPF3 | ENSG00000117360.13 | 21532 | -0.002 | 0.0840 | No | ||
| 85 | MRPL23 | ENSG00000214026.11 | 21654 | -0.005 | 0.0811 | No | ||
| 86 | SOD2 | ENSG00000112096.18 | 21731 | -0.007 | 0.0794 | No | ||
| 87 | CNP | ENSG00000173786.17 | 22180 | -0.018 | 0.0689 | No | ||
| 88 | ATP6V1F | ENSG00000128524.5 | 22492 | -0.026 | 0.0619 | No | ||
| 89 | ACAA1 | ENSG00000060971.18 | 22593 | -0.028 | 0.0601 | No | ||
| 90 | MARK2 | ENSG00000072518.20 | 22797 | -0.033 | 0.0559 | No | ||
| 91 | CCK | ENSG00000187094.12 | 22824 | -0.034 | 0.0561 | No | ||
| 92 | NFKBIA | ENSG00000100906.10 | 22848 | -0.035 | 0.0563 | No | ||
| 93 | CTSV | ENSG00000136943.11 | 22972 | -0.039 | 0.0542 | No | ||
| 94 | AP2S1 | ENSG00000042753.11 | 23077 | -0.042 | 0.0526 | No | ||
| 95 | ALAS1 | ENSG00000023330.15 | 23130 | -0.043 | 0.0523 | No | ||
| 96 | RPN1 | ENSG00000163902.12 | 23194 | -0.045 | 0.0518 | No | ||
| 97 | NPTX2 | ENSG00000106236.4 | 23349 | -0.048 | 0.0491 | No | ||
| 98 | POLE3 | ENSG00000148229.13 | 23456 | -0.051 | 0.0477 | No | ||
| 99 | YKT6 | ENSG00000106636.8 | 24433 | -0.075 | 0.0256 | No | ||
| 100 | PRKACA | ENSG00000072062.14 | 24521 | -0.077 | 0.0253 | No | ||
| 101 | ALDOA | ENSG00000149925.20 | 24530 | -0.078 | 0.0268 | No | ||
| 102 | SPR | ENSG00000116096.6 | 24852 | -0.087 | 0.0210 | No | ||
| 103 | FOSB | ENSG00000125740.14 | 24969 | -0.091 | 0.0202 | No | ||
| 104 | TYRO3 | ENSG00000092445.12 | 25213 | -0.096 | 0.0165 | No | ||
| 105 | CXCL2 | ENSG00000081041.9 | 25575 | -0.107 | 0.0101 | No | ||
| 106 | KLHDC3 | ENSG00000124702.18 | 25664 | -0.110 | 0.0105 | No | ||
| 107 | ARRB2 | ENSG00000141480.18 | 25996 | -0.120 | 0.0052 | No | ||
| 108 | TGFBRAP1 | ENSG00000135966.13 | 26344 | -0.131 | -0.0003 | No | ||
| 109 | FURIN | ENSG00000140564.12 | 26542 | -0.136 | -0.0020 | No | ||
| 110 | H2AFX | ENSG00000188486.3 | 26590 | -0.138 | -0.0000 | No | ||
| 111 | NUP58 | ENSG00000139496.16 | 26686 | -0.140 | 0.0008 | No | ||
| 112 | PDAP1 | ENSG00000106244.13 | 26869 | -0.145 | -0.0003 | No | ||
| 113 | ATF3 | ENSG00000162772.17 | 27076 | -0.151 | -0.0019 | No | ||
| 114 | TMBIM6 | ENSG00000139644.13 | 27100 | -0.152 | 0.0010 | No | ||
| 115 | TFRC | ENSG00000072274.13 | 27176 | -0.154 | 0.0027 | No | ||
| 116 | BAK1 | ENSG00000030110.13 | 27307 | -0.158 | 0.0031 | No | ||
| 117 | EIF5 | ENSG00000100664.11 | 27344 | -0.159 | 0.0059 | No | ||
| 118 | HNRNPU | ENSG00000153187.20 | 27597 | -0.167 | 0.0036 | No | ||
| 119 | CYB5B | ENSG00000103018.16 | 27762 | -0.172 | 0.0035 | No | ||
| 120 | SELENOW | ENSG00000178980.15 | 28142 | -0.185 | -0.0015 | No | ||
| 121 | PPP1R2 | ENSG00000184203.8 | 28214 | -0.187 | 0.0010 | No | ||
| 122 | ATP6V1C1 | ENSG00000155097.12 | 28302 | -0.190 | 0.0032 | No | ||
| 123 | PPT1 | ENSG00000131238.17 | 28310 | -0.190 | 0.0073 | No | ||
| 124 | PSMC3 | ENSG00000165916.8 | 28625 | -0.200 | 0.0042 | No | ||
| 125 | CYB5R1 | ENSG00000159348.13 | 28731 | -0.203 | 0.0063 | No | ||
| 126 | TARS | ENSG00000113407.13 | 28859 | -0.207 | 0.0079 | No | ||
| 127 | STK25 | ENSG00000115694.15 | 28992 | -0.212 | 0.0095 | No | ||
| 128 | PPIF | ENSG00000108179.14 | 29206 | -0.218 | 0.0093 | No | ||
| 129 | BID | ENSG00000015475.18 | 29396 | -0.225 | 0.0098 | No | ||
| 130 | IL6 | ENSG00000136244.12 | 29624 | -0.231 | 0.0095 | No | ||
| 131 | CDC5L | ENSG00000096401.8 | 29701 | -0.235 | 0.0130 | No | ||
| 132 | RFC4 | ENSG00000163918.10 | 30672 | -0.265 | -0.0046 | No | ||
| 133 | FKBP4 | ENSG00000004478.8 | 30809 | -0.270 | -0.0018 | No | ||
| 134 | EIF2S3 | ENSG00000130741.11 | 31765 | -0.288 | -0.0185 | No | ||
| 135 | DNAJA1 | ENSG00000086061.16 | 32115 | -0.301 | -0.0202 | No | ||
| 136 | CCNE1 | ENSG00000105173.14 | 32197 | -0.305 | -0.0152 | No | ||
| 137 | CDK2 | ENSG00000123374.11 | 32246 | -0.307 | -0.0094 | No | ||
| 138 | GGH | ENSG00000137563.12 | 32348 | -0.311 | -0.0048 | No | ||
| 139 | SLC25A4 | ENSG00000151729.11 | 32397 | -0.314 | 0.0012 | No | ||
| 140 | CCND3 | ENSG00000112576.12 | 32532 | -0.317 | 0.0051 | No | ||
| 141 | GRPEL1 | ENSG00000109519.13 | 32886 | -0.331 | 0.0041 | No | ||
| 142 | PRKCD | ENSG00000163932.15 | 34711 | -0.356 | -0.0323 | No | ||
| 143 | CASP3 | ENSG00000164305.19 | 35329 | -0.380 | -0.0387 | No | ||
| 144 | STIP1 | ENSG00000168439.16 | 35973 | -0.404 | -0.0452 | No | ||
| 145 | PPAT | ENSG00000128059.8 | 36104 | -0.409 | -0.0390 | No | ||
| 146 | BTG3 | ENSG00000154640.14 | 36289 | -0.416 | -0.0340 | No | ||
| 147 | SHOX2 | ENSG00000168779.19 | 36566 | -0.430 | -0.0310 | No | ||
| 148 | ENO2 | ENSG00000111674.9 | 36606 | -0.431 | -0.0221 | No | ||
| 149 | RRAD | ENSG00000166592.12 | 37027 | -0.454 | -0.0220 | No | ||
| 150 | PDLIM3 | ENSG00000154553.15 | 37084 | -0.457 | -0.0130 | No | ||
| 151 | HSPA13 | ENSG00000155304.6 | 37137 | -0.459 | -0.0038 | No | ||
| 152 | DNAJB1 | ENSG00000132002.8 | 37147 | -0.460 | 0.0065 | No | ||
| 153 | NAT1 | ENSG00000171428.14 | 37443 | -0.481 | 0.0102 | No | ||
| 154 | FEN1 | ENSG00000168496.4 | 38636 | -0.565 | -0.0060 | No | ||
| 155 | DDX21 | ENSG00000165732.13 | 39636 | -0.651 | -0.0155 | No | ||
| 156 | GLS | ENSG00000115419.12 | 40112 | -0.734 | -0.0104 | No | ||
| 157 | AMD1 | ENSG00000123505.17 | 40193 | -0.751 | 0.0048 | No | ||
| 158 | CLTB | ENSG00000175416.13 | 40460 | -0.827 | 0.0171 | No |