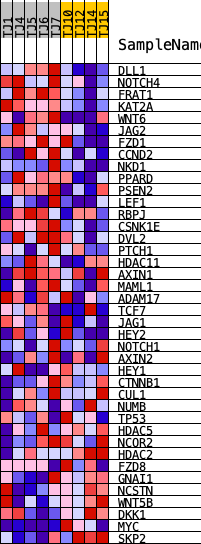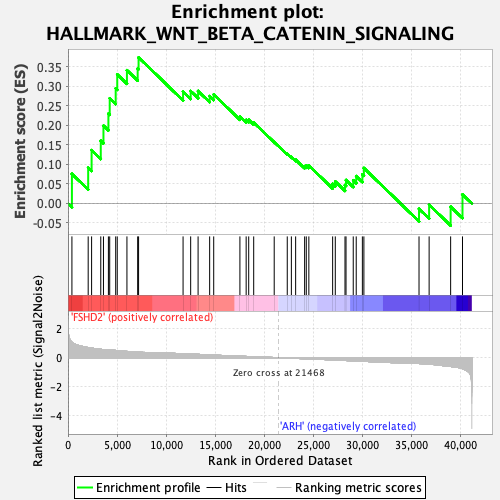
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
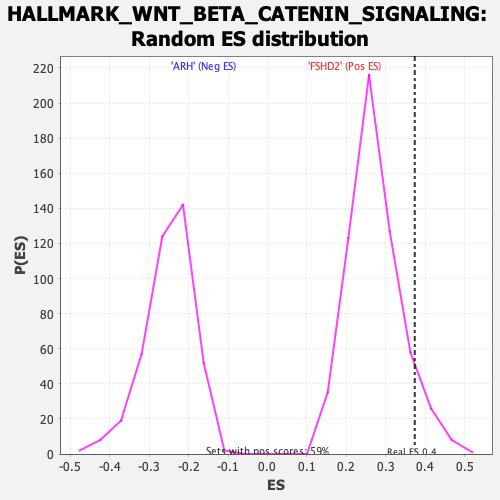
| GeneSet | HALLMARK_WNT_BETA_CATENIN_SIGNALING |
| Enrichment Score (ES) | 0.3735938 |
| Normalized Enrichment Score (NES) | 1.3694108 |
| Nominal p-value | 0.07239057 |
| FDR q-value | 0.09017054 |
| FWER p-Value | 0.723 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DLL1 | ENSG00000198719.9 | 399 | 1.062 | 0.0757 | Yes | ||
| 2 | NOTCH4 | ENSG00000204301.6 | 2057 | 0.698 | 0.0916 | Yes | ||
| 3 | FRAT1 | ENSG00000165879.9 | 2406 | 0.662 | 0.1363 | Yes | ||
| 4 | KAT2A | ENSG00000108773.11 | 3340 | 0.581 | 0.1603 | Yes | ||
| 5 | WNT6 | ENSG00000115596.4 | 3614 | 0.563 | 0.1989 | Yes | ||
| 6 | JAG2 | ENSG00000184916.9 | 4118 | 0.536 | 0.2298 | Yes | ||
| 7 | FZD1 | ENSG00000157240.4 | 4252 | 0.527 | 0.2690 | Yes | ||
| 8 | CCND2 | ENSG00000118971.8 | 4862 | 0.500 | 0.2944 | Yes | ||
| 9 | NKD1 | ENSG00000140807.7 | 5020 | 0.490 | 0.3300 | Yes | ||
| 10 | PPARD | ENSG00000112033.14 | 6007 | 0.433 | 0.3408 | Yes | ||
| 11 | PSEN2 | ENSG00000143801.17 | 7101 | 0.385 | 0.3452 | Yes | ||
| 12 | LEF1 | ENSG00000138795.10 | 7194 | 0.381 | 0.3736 | Yes | ||
| 13 | RBPJ | ENSG00000168214.20 | 11729 | 0.281 | 0.2859 | No | ||
| 14 | CSNK1E | ENSG00000213923.12 | 12504 | 0.254 | 0.2876 | No | ||
| 15 | DVL2 | ENSG00000004975.12 | 13260 | 0.230 | 0.2877 | No | ||
| 16 | PTCH1 | ENSG00000185920.15 | 14439 | 0.193 | 0.2745 | No | ||
| 17 | HDAC11 | ENSG00000163517.15 | 14852 | 0.178 | 0.2789 | No | ||
| 18 | AXIN1 | ENSG00000103126.15 | 17523 | 0.103 | 0.2222 | No | ||
| 19 | MAML1 | ENSG00000161021.13 | 18160 | 0.087 | 0.2138 | No | ||
| 20 | ADAM17 | ENSG00000151694.14 | 18419 | 0.082 | 0.2140 | No | ||
| 21 | TCF7 | ENSG00000081059.20 | 18928 | 0.067 | 0.2071 | No | ||
| 22 | JAG1 | ENSG00000101384.12 | 21019 | 0.012 | 0.1572 | No | ||
| 23 | HEY2 | ENSG00000135547.9 | 22349 | -0.022 | 0.1266 | No | ||
| 24 | NOTCH1 | ENSG00000148400.12 | 22768 | -0.032 | 0.1191 | No | ||
| 25 | AXIN2 | ENSG00000168646.13 | 23204 | -0.045 | 0.1121 | No | ||
| 26 | HEY1 | ENSG00000164683.17 | 24120 | -0.067 | 0.0953 | No | ||
| 27 | CTNNB1 | ENSG00000168036.18 | 24288 | -0.073 | 0.0971 | No | ||
| 28 | CUL1 | ENSG00000055130.17 | 24550 | -0.079 | 0.0970 | No | ||
| 29 | NUMB | ENSG00000133961.20 | 26975 | -0.148 | 0.0500 | No | ||
| 30 | TP53 | ENSG00000141510.17 | 27245 | -0.156 | 0.0561 | No | ||
| 31 | HDAC5 | ENSG00000108840.15 | 28234 | -0.187 | 0.0471 | No | ||
| 32 | NCOR2 | ENSG00000196498.13 | 28339 | -0.191 | 0.0599 | No | ||
| 33 | HDAC2 | ENSG00000196591.12 | 29075 | -0.214 | 0.0593 | No | ||
| 34 | FZD8 | ENSG00000177283.7 | 29379 | -0.224 | 0.0699 | No | ||
| 35 | GNAI1 | ENSG00000127955.17 | 30005 | -0.244 | 0.0743 | No | ||
| 36 | NCSTN | ENSG00000162736.17 | 30160 | -0.249 | 0.0906 | No | ||
| 37 | WNT5B | ENSG00000111186.13 | 35780 | -0.401 | -0.0138 | No | ||
| 38 | DKK1 | ENSG00000107984.10 | 36810 | -0.441 | -0.0034 | No | ||
| 39 | MYC | ENSG00000136997.19 | 39003 | -0.602 | -0.0083 | No | ||
| 40 | SKP2 | ENSG00000145604.16 | 40211 | -0.755 | 0.0231 | No |