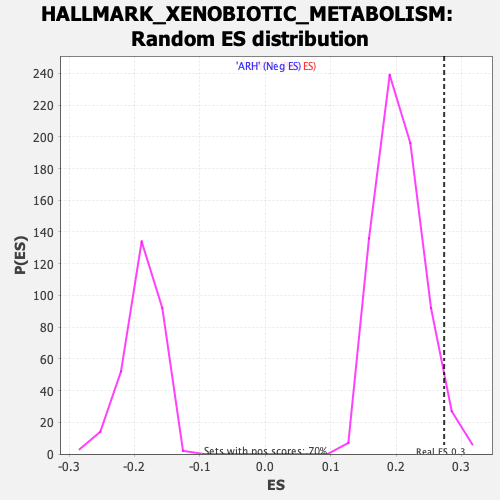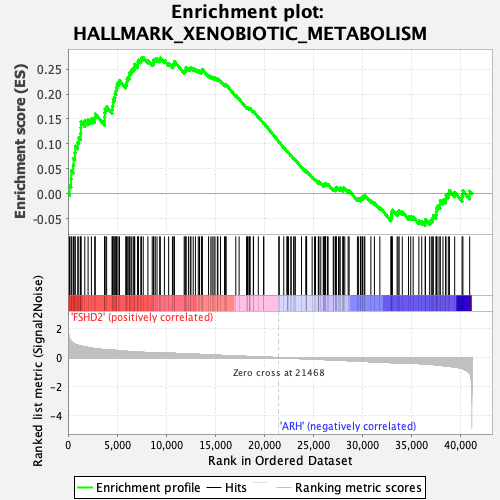
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_XENOBIOTIC_METABOLISM |
| Enrichment Score (ES) | 0.27362055 |
| Normalized Enrichment Score (NES) | 1.3296587 |
| Nominal p-value | 0.038406827 |
| FDR q-value | 0.11556337 |
| FWER p-Value | 0.836 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TGFB2 | ENSG00000092969.12 | 152 | 1.317 | 0.0165 | Yes | ||
| 2 | CYP2E1 | ENSG00000130649.10 | 315 | 1.122 | 0.0297 | Yes | ||
| 3 | GNMT | ENSG00000124713.6 | 333 | 1.100 | 0.0462 | Yes | ||
| 4 | ABCC3 | ENSG00000108846.16 | 520 | 1.008 | 0.0571 | Yes | ||
| 5 | EPHX1 | ENSG00000143819.12 | 570 | 0.990 | 0.0710 | Yes | ||
| 6 | FBLN1 | ENSG00000077942.19 | 705 | 0.938 | 0.0821 | Yes | ||
| 7 | KYNU | ENSG00000115919.15 | 734 | 0.930 | 0.0957 | Yes | ||
| 8 | SLC22A1 | ENSG00000175003.15 | 991 | 0.860 | 0.1026 | Yes | ||
| 9 | DHPS | ENSG00000095059.16 | 1100 | 0.838 | 0.1128 | Yes | ||
| 10 | JUP | ENSG00000173801.17 | 1285 | 0.804 | 0.1207 | Yes | ||
| 11 | CYP1A1 | ENSG00000140465.14 | 1309 | 0.801 | 0.1324 | Yes | ||
| 12 | IL1R1 | ENSG00000115594.12 | 1321 | 0.799 | 0.1443 | Yes | ||
| 13 | SMOX | ENSG00000088826.18 | 1712 | 0.745 | 0.1462 | Yes | ||
| 14 | CYP2S1 | ENSG00000167600.14 | 2047 | 0.699 | 0.1488 | Yes | ||
| 15 | ALDH3A1 | ENSG00000108602.18 | 2386 | 0.664 | 0.1507 | Yes | ||
| 16 | SLC6A6 | ENSG00000131389.17 | 2725 | 0.630 | 0.1521 | Yes | ||
| 17 | SLC46A3 | ENSG00000139508.14 | 2778 | 0.625 | 0.1604 | Yes | ||
| 18 | IGFBP4 | ENSG00000141753.7 | 3723 | 0.556 | 0.1459 | Yes | ||
| 19 | GCKR | ENSG00000084734.9 | 3734 | 0.555 | 0.1542 | Yes | ||
| 20 | PLG | ENSG00000122194.18 | 3746 | 0.554 | 0.1624 | Yes | ||
| 21 | CYP27A1 | ENSG00000135929.9 | 3774 | 0.552 | 0.1702 | Yes | ||
| 22 | MT2A | ENSG00000125148.7 | 3939 | 0.543 | 0.1745 | Yes | ||
| 23 | CYP2C18 | ENSG00000108242.13 | 4489 | 0.518 | 0.1691 | Yes | ||
| 24 | GSTM4 | ENSG00000168765.17 | 4534 | 0.517 | 0.1759 | Yes | ||
| 25 | LEAP2 | ENSG00000164406.8 | 4589 | 0.514 | 0.1825 | Yes | ||
| 26 | HGFAC | ENSG00000109758.8 | 4599 | 0.514 | 0.1901 | Yes | ||
| 27 | PROS1 | ENSG00000184500.15 | 4767 | 0.507 | 0.1938 | Yes | ||
| 28 | CFB | ENSG00000243649.8 | 4800 | 0.504 | 0.2008 | Yes | ||
| 29 | UPB1 | ENSG00000100024.15 | 4894 | 0.498 | 0.2061 | Yes | ||
| 30 | HSD11B1 | ENSG00000117594.10 | 4922 | 0.496 | 0.2131 | Yes | ||
| 31 | SLC6A12 | ENSG00000111181.12 | 4987 | 0.492 | 0.2190 | Yes | ||
| 32 | FBP1 | ENSG00000165140.10 | 5107 | 0.483 | 0.2235 | Yes | ||
| 33 | CYP2J2 | ENSG00000134716.11 | 5250 | 0.474 | 0.2273 | Yes | ||
| 34 | TTPA | ENSG00000137561.5 | 5884 | 0.439 | 0.2186 | Yes | ||
| 35 | ESR1 | ENSG00000091831.23 | 5971 | 0.435 | 0.2232 | Yes | ||
| 36 | PPARD | ENSG00000112033.14 | 6007 | 0.433 | 0.2290 | Yes | ||
| 37 | NQO1 | ENSG00000181019.13 | 6107 | 0.428 | 0.2331 | Yes | ||
| 38 | GCH1 | ENSG00000131979.19 | 6236 | 0.421 | 0.2364 | Yes | ||
| 39 | RBP4 | ENSG00000138207.14 | 6241 | 0.421 | 0.2428 | Yes | ||
| 40 | FAH | ENSG00000103876.12 | 6411 | 0.412 | 0.2450 | Yes | ||
| 41 | UPP1 | ENSG00000183696.14 | 6504 | 0.408 | 0.2490 | Yes | ||
| 42 | CSAD | ENSG00000139631.18 | 6661 | 0.401 | 0.2513 | Yes | ||
| 43 | CCL25 | ENSG00000131142.13 | 6786 | 0.397 | 0.2544 | Yes | ||
| 44 | POR | ENSG00000127948.16 | 6799 | 0.396 | 0.2601 | Yes | ||
| 45 | TNFRSF1A | ENSG00000067182.7 | 7098 | 0.385 | 0.2588 | Yes | ||
| 46 | NPC1 | ENSG00000141458.13 | 7120 | 0.384 | 0.2641 | Yes | ||
| 47 | FMO1 | ENSG00000010932.17 | 7182 | 0.381 | 0.2685 | Yes | ||
| 48 | CYP1A2 | ENSG00000140505.7 | 7424 | 0.372 | 0.2683 | Yes | ||
| 49 | LCAT | ENSG00000213398.8 | 7482 | 0.372 | 0.2726 | Yes | ||
| 50 | GCLC | ENSG00000001084.13 | 7671 | 0.364 | 0.2736 | Yes | ||
| 51 | ACSM1 | ENSG00000166743.9 | 8145 | 0.355 | 0.2675 | No | ||
| 52 | HNF4A | ENSG00000101076.16 | 8585 | 0.348 | 0.2621 | No | ||
| 53 | PEMT | ENSG00000133027.18 | 8729 | 0.346 | 0.2640 | No | ||
| 54 | CAT | ENSG00000121691.7 | 8736 | 0.346 | 0.2691 | No | ||
| 55 | CDO1 | ENSG00000129596.5 | 8894 | 0.340 | 0.2705 | No | ||
| 56 | HMOX1 | ENSG00000100292.17 | 9075 | 0.332 | 0.2712 | No | ||
| 57 | ITIH1 | ENSG00000055957.11 | 9346 | 0.321 | 0.2695 | No | ||
| 58 | MBL2 | ENSG00000165471.6 | 9409 | 0.319 | 0.2729 | No | ||
| 59 | TMEM97 | ENSG00000109084.14 | 9826 | 0.318 | 0.2676 | No | ||
| 60 | CRP | ENSG00000132693.12 | 10259 | 0.305 | 0.2617 | No | ||
| 61 | PTGDS | ENSG00000107317.13 | 10645 | 0.305 | 0.2570 | No | ||
| 62 | AKR1C2 | ENSG00000151632.17 | 10705 | 0.302 | 0.2602 | No | ||
| 63 | APOE | ENSG00000130203.10 | 10826 | 0.298 | 0.2618 | No | ||
| 64 | PTGES | ENSG00000148344.11 | 10848 | 0.297 | 0.2659 | No | ||
| 65 | ACOX2 | ENSG00000168306.13 | 11842 | 0.277 | 0.2459 | No | ||
| 66 | NINJ1 | ENSG00000131669.10 | 11926 | 0.274 | 0.2480 | No | ||
| 67 | SLC35D1 | ENSG00000116704.8 | 12017 | 0.270 | 0.2500 | No | ||
| 68 | MARCH6 | ENSG00000145495.16 | 12033 | 0.270 | 0.2538 | No | ||
| 69 | PDK4 | ENSG00000004799.8 | 12277 | 0.262 | 0.2518 | No | ||
| 70 | GABARAPL1 | ENSG00000139112.11 | 12462 | 0.255 | 0.2513 | No | ||
| 71 | CA2 | ENSG00000104267.10 | 12533 | 0.253 | 0.2534 | No | ||
| 72 | MPP2 | ENSG00000108852.14 | 12762 | 0.246 | 0.2516 | No | ||
| 73 | SERTAD1 | ENSG00000197019.5 | 12988 | 0.238 | 0.2498 | No | ||
| 74 | VTN | ENSG00000109072.14 | 13279 | 0.229 | 0.2462 | No | ||
| 75 | IGF1 | ENSG00000017427.16 | 13404 | 0.225 | 0.2466 | No | ||
| 76 | ACP2 | ENSG00000134575.12 | 13600 | 0.219 | 0.2452 | No | ||
| 77 | HSD17B2 | ENSG00000086696.11 | 13673 | 0.217 | 0.2468 | No | ||
| 78 | NDRG2 | ENSG00000165795.23 | 13680 | 0.216 | 0.2500 | No | ||
| 79 | GSTT2 | ENSG00000099984.11 | 14325 | 0.197 | 0.2373 | No | ||
| 80 | SHMT2 | ENSG00000182199.11 | 14562 | 0.188 | 0.2344 | No | ||
| 81 | ABCD2 | ENSG00000173208.4 | 14695 | 0.184 | 0.2340 | No | ||
| 82 | CYFIP2 | ENSG00000055163.20 | 14875 | 0.178 | 0.2323 | No | ||
| 83 | TKFC | ENSG00000149476.15 | 14982 | 0.174 | 0.2324 | No | ||
| 84 | ANGPTL3 | ENSG00000132855.5 | 15223 | 0.167 | 0.2291 | No | ||
| 85 | CBR1 | ENSG00000159228.13 | 15279 | 0.165 | 0.2303 | No | ||
| 86 | ALDH2 | ENSG00000111275.13 | 15545 | 0.158 | 0.2263 | No | ||
| 87 | HRG | ENSG00000113905.4 | 15955 | 0.146 | 0.2185 | No | ||
| 88 | DHRS7 | ENSG00000100612.14 | 16041 | 0.144 | 0.2187 | No | ||
| 89 | GAD1 | ENSG00000128683.14 | 16125 | 0.142 | 0.2188 | No | ||
| 90 | CNDP2 | ENSG00000133313.15 | 17103 | 0.115 | 0.1967 | No | ||
| 91 | MAOA | ENSG00000189221.9 | 17441 | 0.106 | 0.1901 | No | ||
| 92 | AKR1C3 | ENSG00000196139.14 | 18197 | 0.085 | 0.1730 | No | ||
| 93 | SLC12A4 | ENSG00000124067.17 | 18300 | 0.084 | 0.1718 | No | ||
| 94 | HES6 | ENSG00000144485.11 | 18342 | 0.083 | 0.1720 | No | ||
| 95 | ACOX1 | ENSG00000161533.12 | 18471 | 0.080 | 0.1701 | No | ||
| 96 | PGD | ENSG00000142657.21 | 18499 | 0.079 | 0.1707 | No | ||
| 97 | DHRS1 | ENSG00000157379.14 | 18520 | 0.078 | 0.1714 | No | ||
| 98 | ELOVL5 | ENSG00000012660.14 | 18561 | 0.077 | 0.1716 | No | ||
| 99 | RAP1GAP | ENSG00000076864.19 | 18890 | 0.068 | 0.1647 | No | ||
| 100 | GCNT2 | ENSG00000111846.17 | 18915 | 0.068 | 0.1651 | No | ||
| 101 | ECH1 | ENSG00000104823.9 | 19400 | 0.054 | 0.1541 | No | ||
| 102 | PSMB10 | ENSG00000205220.12 | 19937 | 0.040 | 0.1416 | No | ||
| 103 | CROT | ENSG00000005469.11 | 19961 | 0.039 | 0.1417 | No | ||
| 104 | PGRMC1 | ENSG00000101856.10 | 21489 | -0.001 | 0.1044 | No | ||
| 105 | ACO2 | ENSG00000100412.16 | 21529 | -0.002 | 0.1035 | No | ||
| 106 | BPHL | ENSG00000137274.13 | 21993 | -0.014 | 0.0924 | No | ||
| 107 | PYCR1 | ENSG00000183010.17 | 22333 | -0.022 | 0.0845 | No | ||
| 108 | AOX1 | ENSG00000138356.14 | 22339 | -0.022 | 0.0847 | No | ||
| 109 | DDAH2 | ENSG00000213722.9 | 22360 | -0.022 | 0.0845 | No | ||
| 110 | ATP2A2 | ENSG00000174437.17 | 22480 | -0.026 | 0.0820 | No | ||
| 111 | ETS2 | ENSG00000157557.13 | 22722 | -0.031 | 0.0766 | No | ||
| 112 | AP4B1 | ENSG00000134262.13 | 22742 | -0.032 | 0.0766 | No | ||
| 113 | IDH1 | ENSG00000138413.13 | 22996 | -0.039 | 0.0711 | No | ||
| 114 | ALAS1 | ENSG00000023330.15 | 23130 | -0.043 | 0.0685 | No | ||
| 115 | ITIH4 | ENSG00000055955.17 | 23165 | -0.044 | 0.0683 | No | ||
| 116 | ABCC2 | ENSG00000023839.12 | 23803 | -0.060 | 0.0537 | No | ||
| 117 | TPST1 | ENSG00000169902.15 | 24255 | -0.072 | 0.0438 | No | ||
| 118 | ARG1 | ENSG00000118520.15 | 24273 | -0.072 | 0.0445 | No | ||
| 119 | SLC35B1 | ENSG00000121073.15 | 24312 | -0.073 | 0.0447 | No | ||
| 120 | ACP1 | ENSG00000143727.16 | 24324 | -0.074 | 0.0456 | No | ||
| 121 | DCXR | ENSG00000169738.8 | 24886 | -0.088 | 0.0332 | No | ||
| 122 | TMEM176B | ENSG00000106565.18 | 25133 | -0.094 | 0.0286 | No | ||
| 123 | F10 | ENSG00000126218.12 | 25229 | -0.097 | 0.0278 | No | ||
| 124 | CYB5A | ENSG00000166347.19 | 25515 | -0.106 | 0.0225 | No | ||
| 125 | PMM1 | ENSG00000100417.12 | 25550 | -0.107 | 0.0233 | No | ||
| 126 | KARS | ENSG00000065427.14 | 25576 | -0.107 | 0.0243 | No | ||
| 127 | RETSAT | ENSG00000042445.14 | 25766 | -0.113 | 0.0214 | No | ||
| 128 | LONP1 | ENSG00000196365.12 | 26016 | -0.121 | 0.0172 | No | ||
| 129 | COMT | ENSG00000093010.13 | 26064 | -0.123 | 0.0179 | No | ||
| 130 | ALDH9A1 | ENSG00000143149.12 | 26176 | -0.125 | 0.0171 | No | ||
| 131 | PINK1 | ENSG00000158828.8 | 26178 | -0.125 | 0.0190 | No | ||
| 132 | AQP9 | ENSG00000103569.10 | 26201 | -0.126 | 0.0204 | No | ||
| 133 | NFS1 | ENSG00000244005.13 | 26256 | -0.127 | 0.0211 | No | ||
| 134 | BCAT1 | ENSG00000060982.15 | 26443 | -0.133 | 0.0186 | No | ||
| 135 | TDO2 | ENSG00000151790.9 | 26514 | -0.136 | 0.0189 | No | ||
| 136 | ATOH8 | ENSG00000168874.13 | 27038 | -0.151 | 0.0085 | No | ||
| 137 | TMBIM6 | ENSG00000139644.13 | 27100 | -0.152 | 0.0093 | No | ||
| 138 | PTS | ENSG00000150787.8 | 27261 | -0.157 | 0.0078 | No | ||
| 139 | PAPSS2 | ENSG00000198682.13 | 27309 | -0.158 | 0.0091 | No | ||
| 140 | CASP6 | ENSG00000138794.10 | 27327 | -0.159 | 0.0111 | No | ||
| 141 | BLVRB | ENSG00000090013.11 | 27336 | -0.159 | 0.0133 | No | ||
| 142 | BCAR1 | ENSG00000050820.17 | 27576 | -0.167 | 0.0101 | No | ||
| 143 | UGDH | ENSG00000109814.12 | 27716 | -0.171 | 0.0093 | No | ||
| 144 | ENPEP | ENSG00000138792.10 | 27731 | -0.171 | 0.0116 | No | ||
| 145 | SLC1A5 | ENSG00000105281.12 | 28017 | -0.181 | 0.0074 | No | ||
| 146 | SAR1B | ENSG00000152700.14 | 28046 | -0.182 | 0.0095 | No | ||
| 147 | CES1 | ENSG00000198848.12 | 28060 | -0.183 | 0.0120 | No | ||
| 148 | GSR | ENSG00000104687.14 | 28212 | -0.187 | 0.0112 | No | ||
| 149 | IGFBP1 | ENSG00000146678.10 | 28557 | -0.198 | 0.0058 | No | ||
| 150 | NMT1 | ENSG00000136448.12 | 28656 | -0.201 | 0.0065 | No | ||
| 151 | ARG2 | ENSG00000081181.8 | 29501 | -0.227 | -0.0106 | No | ||
| 152 | GSS | ENSG00000100983.12 | 29622 | -0.231 | -0.0100 | No | ||
| 153 | CYP17A1 | ENSG00000148795.7 | 29819 | -0.239 | -0.0111 | No | ||
| 154 | CDA | ENSG00000158825.6 | 29864 | -0.240 | -0.0085 | No | ||
| 155 | PC | ENSG00000173599.15 | 30008 | -0.244 | -0.0083 | No | ||
| 156 | GART | ENSG00000159131.17 | 30040 | -0.245 | -0.0053 | No | ||
| 157 | PTGES3 | ENSG00000110958.16 | 30197 | -0.250 | -0.0053 | No | ||
| 158 | SPINT2 | ENSG00000167642.13 | 30256 | -0.252 | -0.0028 | No | ||
| 159 | ETFDH | ENSG00000171503.12 | 30873 | -0.272 | -0.0137 | No | ||
| 160 | F11 | ENSG00000088926.13 | 31232 | -0.285 | -0.0180 | No | ||
| 161 | ADH1C | ENSG00000248144.6 | 31787 | -0.289 | -0.0271 | No | ||
| 162 | ABHD6 | ENSG00000163686.15 | 32914 | -0.332 | -0.0495 | No | ||
| 163 | ENTPD5 | ENSG00000187097.12 | 32955 | -0.334 | -0.0454 | No | ||
| 164 | PTGR1 | ENSG00000106853.20 | 32967 | -0.335 | -0.0405 | No | ||
| 165 | CD36 | ENSG00000135218.19 | 32994 | -0.336 | -0.0360 | No | ||
| 166 | G6PC | ENSG00000131482.10 | 33064 | -0.336 | -0.0325 | No | ||
| 167 | MAN1A1 | ENSG00000111885.7 | 33550 | -0.338 | -0.0392 | No | ||
| 168 | DDT | ENSG00000099977.14 | 33641 | -0.342 | -0.0362 | No | ||
| 169 | TYR | ENSG00000077498.9 | 33753 | -0.342 | -0.0336 | No | ||
| 170 | FAS | ENSG00000026103.22 | 34075 | -0.344 | -0.0362 | No | ||
| 171 | MCCC2 | ENSG00000131844.16 | 34708 | -0.356 | -0.0462 | No | ||
| 172 | CYP26A1 | ENSG00000095596.12 | 34937 | -0.363 | -0.0462 | No | ||
| 173 | AHCY | ENSG00000101444.13 | 35184 | -0.373 | -0.0464 | No | ||
| 174 | GSTO1 | ENSG00000148834.13 | 35765 | -0.400 | -0.0545 | No | ||
| 175 | ID2 | ENSG00000115738.10 | 36059 | -0.406 | -0.0554 | No | ||
| 176 | HACL1 | ENSG00000131373.15 | 36411 | -0.421 | -0.0575 | No | ||
| 177 | VNN1 | ENSG00000112299.8 | 36430 | -0.423 | -0.0515 | No | ||
| 178 | HPRT1 | ENSG00000165704.15 | 36862 | -0.444 | -0.0552 | No | ||
| 179 | SERPINA6 | ENSG00000170099.6 | 37032 | -0.454 | -0.0524 | No | ||
| 180 | DDC | ENSG00000132437.18 | 37176 | -0.462 | -0.0488 | No | ||
| 181 | SSR3 | ENSG00000114850.6 | 37238 | -0.466 | -0.0431 | No | ||
| 182 | FMO3 | ENSG00000007933.13 | 37515 | -0.486 | -0.0424 | No | ||
| 183 | SERPINE1 | ENSG00000106366.8 | 37533 | -0.487 | -0.0353 | No | ||
| 184 | ASL | ENSG00000126522.17 | 37561 | -0.490 | -0.0285 | No | ||
| 185 | ADH5 | ENSG00000197894.11 | 37712 | -0.503 | -0.0245 | No | ||
| 186 | LPIN2 | ENSG00000101577.9 | 37920 | -0.522 | -0.0215 | No | ||
| 187 | ADH7 | ENSG00000196344.11 | 37933 | -0.523 | -0.0138 | No | ||
| 188 | TAT | ENSG00000198650.11 | 38222 | -0.544 | -0.0125 | No | ||
| 189 | CYP4F2 | ENSG00000186115.13 | 38494 | -0.558 | -0.0105 | No | ||
| 190 | EPHA2 | ENSG00000142627.13 | 38526 | -0.559 | -0.0027 | No | ||
| 191 | XDH | ENSG00000158125.10 | 38767 | -0.577 | 0.0003 | No | ||
| 192 | ARPP19 | ENSG00000128989.10 | 38877 | -0.588 | 0.0066 | No | ||
| 193 | ACOX3 | ENSG00000087008.16 | 39414 | -0.628 | 0.0031 | No | ||
| 194 | PDLIM5 | ENSG00000163110.15 | 40169 | -0.746 | -0.0038 | No | ||
| 195 | MTHFD1 | ENSG00000100714.17 | 40252 | -0.766 | 0.0059 | No | ||
| 196 | FETUB | ENSG00000090512.12 | 40944 | -1.061 | 0.0053 | No |