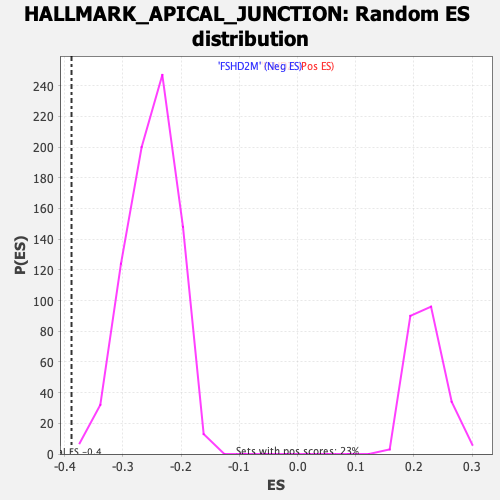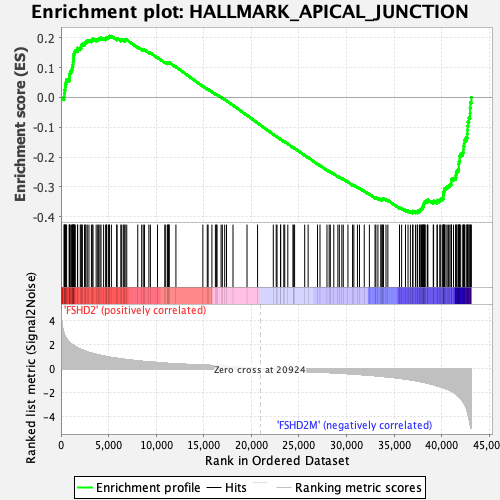
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_APICAL_JUNCTION |
| Enrichment Score (ES) | -0.38823855 |
| Normalized Enrichment Score (NES) | -1.5465555 |
| Nominal p-value | 0.0012970169 |
| FDR q-value | 0.010822162 |
| FWER p-Value | 0.149 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ITGA2 | ENSG00000164171.11 | 327 | 2.899 | 0.0038 | No | ||
| 2 | SGCE | ENSG00000127990.19 | 345 | 2.869 | 0.0147 | No | ||
| 3 | RAC2 | ENSG00000128340.15 | 379 | 2.785 | 0.0249 | No | ||
| 4 | AMIGO1 | ENSG00000181754.7 | 449 | 2.680 | 0.0338 | No | ||
| 5 | SLIT2 | ENSG00000145147.20 | 459 | 2.664 | 0.0441 | No | ||
| 6 | EGFR | ENSG00000146648.18 | 529 | 2.570 | 0.0526 | No | ||
| 7 | COL16A1 | ENSG00000084636.18 | 589 | 2.483 | 0.0610 | No | ||
| 8 | FYB1 | ENSG00000082074.18 | 857 | 2.213 | 0.0635 | No | ||
| 9 | SORBS3 | ENSG00000120896.13 | 884 | 2.183 | 0.0715 | No | ||
| 10 | CERCAM | ENSG00000167123.19 | 910 | 2.164 | 0.0794 | No | ||
| 11 | CLDN11 | ENSG00000013297.11 | 969 | 2.122 | 0.0864 | No | ||
| 12 | FSCN1 | ENSG00000075618.18 | 1089 | 2.055 | 0.0917 | No | ||
| 13 | SRC | ENSG00000197122.11 | 1168 | 2.008 | 0.0978 | No | ||
| 14 | SDC3 | ENSG00000162512.16 | 1208 | 1.985 | 0.1047 | No | ||
| 15 | MAPK13 | ENSG00000156711.17 | 1253 | 1.961 | 0.1114 | No | ||
| 16 | INSIG1 | ENSG00000186480.13 | 1265 | 1.957 | 0.1189 | No | ||
| 17 | EPB41L2 | ENSG00000079819.19 | 1286 | 1.948 | 0.1261 | No | ||
| 18 | RRAS | ENSG00000126458.4 | 1335 | 1.916 | 0.1325 | No | ||
| 19 | TSPAN4 | ENSG00000214063.11 | 1342 | 1.912 | 0.1399 | No | ||
| 20 | TNFRSF11B | ENSG00000164761.9 | 1360 | 1.901 | 0.1470 | No | ||
| 21 | MADCAM1 | ENSG00000099866.15 | 1423 | 1.877 | 0.1529 | No | ||
| 22 | MAP4K2 | ENSG00000168067.12 | 1501 | 1.844 | 0.1584 | No | ||
| 23 | CRAT | ENSG00000095321.17 | 1730 | 1.743 | 0.1599 | No | ||
| 24 | NLGN2 | ENSG00000169992.10 | 1752 | 1.735 | 0.1662 | No | ||
| 25 | MPZL1 | ENSG00000197965.12 | 2024 | 1.621 | 0.1663 | No | ||
| 26 | MYH10 | ENSG00000133026.12 | 2114 | 1.594 | 0.1705 | No | ||
| 27 | MAPK11 | ENSG00000185386.15 | 2130 | 1.590 | 0.1764 | No | ||
| 28 | THY1 | ENSG00000154096.13 | 2244 | 1.552 | 0.1799 | No | ||
| 29 | LAYN | ENSG00000204381.11 | 2467 | 1.482 | 0.1806 | No | ||
| 30 | CLDN7 | ENSG00000181885.18 | 2537 | 1.461 | 0.1847 | No | ||
| 31 | MMP2 | ENSG00000087245.13 | 2616 | 1.437 | 0.1885 | No | ||
| 32 | CDH11 | ENSG00000140937.14 | 2757 | 1.391 | 0.1908 | No | ||
| 33 | CLDN15 | ENSG00000106404.13 | 2934 | 1.348 | 0.1920 | No | ||
| 34 | TMEM8B | ENSG00000137103.20 | 3195 | 1.290 | 0.1910 | No | ||
| 35 | MVD | ENSG00000167508.12 | 3276 | 1.273 | 0.1941 | No | ||
| 36 | LDLRAP1 | ENSG00000157978.11 | 3364 | 1.252 | 0.1970 | No | ||
| 37 | VWF | ENSG00000110799.13 | 3693 | 1.186 | 0.1941 | No | ||
| 38 | WASL | ENSG00000106299.8 | 3858 | 1.154 | 0.1948 | No | ||
| 39 | PBX2 | ENSG00000204304.12 | 3959 | 1.135 | 0.1969 | No | ||
| 40 | ADAM15 | ENSG00000143537.13 | 4091 | 1.114 | 0.1982 | No | ||
| 41 | ICAM1 | ENSG00000090339.9 | 4204 | 1.093 | 0.1999 | No | ||
| 42 | RSU1 | ENSG00000148484.18 | 4475 | 1.048 | 0.1978 | No | ||
| 43 | CLDN18 | ENSG00000066405.13 | 4688 | 1.016 | 0.1968 | No | ||
| 44 | AKT2 | ENSG00000105221.18 | 4775 | 1.001 | 0.1988 | No | ||
| 45 | CX3CL1 | ENSG00000006210.7 | 4796 | 0.999 | 0.2022 | No | ||
| 46 | NECTIN3 | ENSG00000177707.11 | 4974 | 0.972 | 0.2019 | No | ||
| 47 | MDK | ENSG00000110492.15 | 5041 | 0.962 | 0.2042 | No | ||
| 48 | GNAI2 | ENSG00000114353.17 | 5091 | 0.956 | 0.2068 | No | ||
| 49 | CNN2 | ENSG00000064666.15 | 5294 | 0.928 | 0.2058 | No | ||
| 50 | FBN1 | ENSG00000166147.13 | 5850 | 0.858 | 0.1962 | No | ||
| 51 | GNAI1 | ENSG00000127955.17 | 5889 | 0.854 | 0.1987 | No | ||
| 52 | THBS3 | ENSG00000169231.13 | 6284 | 0.808 | 0.1927 | No | ||
| 53 | NLGN3 | ENSG00000196338.12 | 6377 | 0.800 | 0.1937 | No | ||
| 54 | MAP3K20 | ENSG00000091436.17 | 6549 | 0.783 | 0.1928 | No | ||
| 55 | TRAF1 | ENSG00000056558.11 | 6707 | 0.767 | 0.1921 | No | ||
| 56 | SKAP2 | ENSG00000005020.13 | 6720 | 0.765 | 0.1949 | No | ||
| 57 | CD99 | ENSG00000002586.20 | 6907 | 0.746 | 0.1935 | No | ||
| 58 | CLDN4 | ENSG00000189143.9 | 8062 | 0.642 | 0.1691 | No | ||
| 59 | RHOF | ENSG00000139725.8 | 8475 | 0.608 | 0.1619 | No | ||
| 60 | CALB2 | ENSG00000172137.19 | 8638 | 0.596 | 0.1605 | No | ||
| 61 | NFASC | ENSG00000163531.15 | 8774 | 0.585 | 0.1596 | No | ||
| 62 | PDZD3 | ENSG00000172367.16 | 9232 | 0.553 | 0.1512 | No | ||
| 63 | NEGR1 | ENSG00000172260.15 | 9390 | 0.544 | 0.1497 | No | ||
| 64 | ICAM5 | ENSG00000105376.5 | 10136 | 0.501 | 0.1343 | No | ||
| 65 | MAPK14 | ENSG00000112062.10 | 10900 | 0.451 | 0.1183 | No | ||
| 66 | PIK3R3 | ENSG00000117461.15 | 11004 | 0.444 | 0.1176 | No | ||
| 67 | EVL | ENSG00000196405.13 | 11210 | 0.432 | 0.1145 | No | ||
| 68 | SIRPA | ENSG00000198053.11 | 11216 | 0.432 | 0.1161 | No | ||
| 69 | NRTN | ENSG00000171119.2 | 11280 | 0.428 | 0.1163 | No | ||
| 70 | BAIAP2 | ENSG00000175866.15 | 11334 | 0.425 | 0.1168 | No | ||
| 71 | IKBKG | ENSG00000269335.5 | 11365 | 0.423 | 0.1178 | No | ||
| 72 | TIAL1 | ENSG00000151923.17 | 12072 | 0.384 | 0.1028 | No | ||
| 73 | DMP1 | ENSG00000152592.14 | 14902 | 0.287 | 0.0380 | No | ||
| 74 | ADAM9 | ENSG00000168615.12 | 15364 | 0.282 | 0.0284 | No | ||
| 75 | TSC1 | ENSG00000165699.14 | 15451 | 0.278 | 0.0275 | No | ||
| 76 | SHC1 | ENSG00000160691.18 | 15852 | 0.257 | 0.0192 | No | ||
| 77 | CD209 | ENSG00000090659.18 | 16218 | 0.235 | 0.0116 | No | ||
| 78 | PARD6G | ENSG00000178184.16 | 16320 | 0.229 | 0.0102 | No | ||
| 79 | CDH6 | ENSG00000113361.13 | 16393 | 0.226 | 0.0094 | No | ||
| 80 | DSC3 | ENSG00000134762.17 | 16817 | 0.206 | 0.0003 | No | ||
| 81 | ITGB4 | ENSG00000132470.14 | 16947 | 0.199 | -0.0019 | No | ||
| 82 | PARVA | ENSG00000197702.14 | 17194 | 0.188 | -0.0069 | No | ||
| 83 | ICAM2 | ENSG00000108622.11 | 17387 | 0.179 | -0.0107 | No | ||
| 84 | NF1 | ENSG00000196712.17 | 18068 | 0.143 | -0.0259 | No | ||
| 85 | PPP2R2C | ENSG00000074211.14 | 19540 | 0.069 | -0.0599 | No | ||
| 86 | ATP1A3 | ENSG00000105409.19 | 20648 | 0.014 | -0.0857 | No | ||
| 87 | CNTN1 | ENSG00000018236.15 | 22301 | -0.074 | -0.1239 | No | ||
| 88 | PTEN | ENSG00000171862.11 | 22605 | -0.091 | -0.1306 | No | ||
| 89 | CAP1 | ENSG00000131236.17 | 22692 | -0.096 | -0.1322 | No | ||
| 90 | COL9A1 | ENSG00000112280.16 | 23057 | -0.115 | -0.1402 | No | ||
| 91 | STX4 | ENSG00000103496.15 | 23081 | -0.117 | -0.1403 | No | ||
| 92 | ZYX | ENSG00000159840.16 | 23404 | -0.135 | -0.1473 | No | ||
| 93 | PECAM1 | ENSG00000261371.6 | 23502 | -0.141 | -0.1490 | No | ||
| 94 | CDH3 | ENSG00000062038.14 | 23825 | -0.160 | -0.1558 | No | ||
| 95 | ARPC2 | ENSG00000163466.16 | 24381 | -0.192 | -0.1680 | No | ||
| 96 | PKD1 | ENSG00000008710.19 | 24395 | -0.193 | -0.1676 | No | ||
| 97 | TAOK2 | ENSG00000149930.18 | 24538 | -0.201 | -0.1701 | No | ||
| 98 | YWHAH | ENSG00000128245.15 | 25599 | -0.233 | -0.1939 | No | ||
| 99 | CDSN | ENSG00000204539.4 | 25960 | -0.258 | -0.2012 | No | ||
| 100 | MPZL2 | ENSG00000149573.9 | 26944 | -0.275 | -0.2230 | No | ||
| 101 | AKT3 | ENSG00000117020.18 | 27205 | -0.288 | -0.2280 | No | ||
| 102 | CLDN19 | ENSG00000164007.11 | 27939 | -0.305 | -0.2438 | No | ||
| 103 | CLDN6 | ENSG00000184697.7 | 28170 | -0.321 | -0.2479 | No | ||
| 104 | CD276 | ENSG00000103855.18 | 28292 | -0.329 | -0.2495 | No | ||
| 105 | CRB3 | ENSG00000130545.16 | 28656 | -0.341 | -0.2566 | No | ||
| 106 | WNK4 | ENSG00000126562.17 | 29085 | -0.353 | -0.2652 | No | ||
| 107 | CD86 | ENSG00000114013.16 | 29238 | -0.360 | -0.2673 | No | ||
| 108 | PLCG1 | ENSG00000124181.14 | 29500 | -0.374 | -0.2719 | No | ||
| 109 | NECTIN1 | ENSG00000110400.11 | 29652 | -0.385 | -0.2739 | No | ||
| 110 | PIK3CB | ENSG00000051382.8 | 30141 | -0.417 | -0.2836 | No | ||
| 111 | ACTG1 | ENSG00000184009.12 | 30626 | -0.445 | -0.2932 | No | ||
| 112 | GTF2F1 | ENSG00000125651.14 | 30762 | -0.451 | -0.2945 | No | ||
| 113 | MMP9 | ENSG00000100985.7 | 31139 | -0.471 | -0.3014 | No | ||
| 114 | CDK8 | ENSG00000132964.12 | 31354 | -0.482 | -0.3045 | No | ||
| 115 | JUP | ENSG00000173801.17 | 31855 | -0.505 | -0.3142 | No | ||
| 116 | EXOC4 | ENSG00000131558.15 | 32395 | -0.538 | -0.3246 | No | ||
| 117 | LAMA3 | ENSG00000053747.16 | 32988 | -0.582 | -0.3361 | No | ||
| 118 | ADAMTS5 | ENSG00000154736.6 | 33077 | -0.588 | -0.3359 | No | ||
| 119 | ITGA9 | ENSG00000144668.12 | 33291 | -0.604 | -0.3384 | No | ||
| 120 | ADAM23 | ENSG00000114948.13 | 33607 | -0.630 | -0.3433 | No | ||
| 121 | SYMPK | ENSG00000125755.19 | 33623 | -0.631 | -0.3412 | No | ||
| 122 | COL17A1 | ENSG00000065618.21 | 33743 | -0.641 | -0.3414 | No | ||
| 123 | DHX16 | ENSG00000204560.10 | 33810 | -0.646 | -0.3404 | No | ||
| 124 | GRB7 | ENSG00000141738.14 | 33880 | -0.652 | -0.3394 | No | ||
| 125 | PTK2 | ENSG00000169398.19 | 34139 | -0.671 | -0.3428 | No | ||
| 126 | LIMA1 | ENSG00000050405.13 | 34334 | -0.686 | -0.3446 | No | ||
| 127 | SHROOM2 | ENSG00000146950.13 | 35559 | -0.793 | -0.3700 | No | ||
| 128 | IRS1 | ENSG00000169047.5 | 35781 | -0.816 | -0.3720 | No | ||
| 129 | VCAN | ENSG00000038427.16 | 36187 | -0.866 | -0.3780 | No | ||
| 130 | ITGB1 | ENSG00000150093.19 | 36454 | -0.895 | -0.3807 | No | ||
| 131 | CADM3 | ENSG00000162706.13 | 36703 | -0.930 | -0.3828 | No | ||
| 132 | ADRA1B | ENSG00000170214.5 | 36938 | -0.957 | -0.3845 | Yes | ||
| 133 | TUBG1 | ENSG00000131462.8 | 36973 | -0.963 | -0.3815 | Yes | ||
| 134 | CDH1 | ENSG00000039068.19 | 37235 | -0.997 | -0.3836 | Yes | ||
| 135 | LAMC2 | ENSG00000058085.15 | 37414 | -1.022 | -0.3838 | Yes | ||
| 136 | PCDH1 | ENSG00000156453.13 | 37463 | -1.029 | -0.3808 | Yes | ||
| 137 | VCL | ENSG00000035403.17 | 37662 | -1.061 | -0.3813 | Yes | ||
| 138 | VAV2 | ENSG00000160293.17 | 37686 | -1.065 | -0.3776 | Yes | ||
| 139 | MYL12B | ENSG00000118680.13 | 37797 | -1.083 | -0.3759 | Yes | ||
| 140 | TJP1 | ENSG00000104067.16 | 37865 | -1.093 | -0.3732 | Yes | ||
| 141 | VASP | ENSG00000125753.14 | 37949 | -1.108 | -0.3707 | Yes | ||
| 142 | CLDN9 | ENSG00000213937.4 | 37970 | -1.113 | -0.3668 | Yes | ||
| 143 | SYK | ENSG00000165025.15 | 38062 | -1.127 | -0.3645 | Yes | ||
| 144 | SPEG | ENSG00000072195.15 | 38065 | -1.128 | -0.3601 | Yes | ||
| 145 | CD34 | ENSG00000174059.17 | 38113 | -1.137 | -0.3567 | Yes | ||
| 146 | NF2 | ENSG00000186575.18 | 38185 | -1.151 | -0.3539 | Yes | ||
| 147 | HRAS | ENSG00000174775.17 | 38198 | -1.153 | -0.3496 | Yes | ||
| 148 | GAMT | ENSG00000130005.13 | 38317 | -1.174 | -0.3477 | Yes | ||
| 149 | ACTB | ENSG00000075624.16 | 38506 | -1.208 | -0.3474 | Yes | ||
| 150 | CADM2 | ENSG00000175161.13 | 38535 | -1.213 | -0.3432 | Yes | ||
| 151 | ACTN3 | ENSG00000248746.6 | 39105 | -1.332 | -0.3512 | Yes | ||
| 152 | TRO | ENSG00000067445.21 | 39139 | -1.338 | -0.3467 | Yes | ||
| 153 | BMP1 | ENSG00000168487.19 | 39505 | -1.423 | -0.3497 | Yes | ||
| 154 | ITGA10 | ENSG00000143127.13 | 39526 | -1.426 | -0.3445 | Yes | ||
| 155 | B4GALT1 | ENSG00000086062.12 | 39786 | -1.489 | -0.3447 | Yes | ||
| 156 | ALOX15B | ENSG00000179593.16 | 39887 | -1.519 | -0.3410 | Yes | ||
| 157 | PFN1 | ENSG00000108518.7 | 40055 | -1.560 | -0.3388 | Yes | ||
| 158 | LAMB3 | ENSG00000196878.15 | 40157 | -1.589 | -0.3349 | Yes | ||
| 159 | NRAP | ENSG00000197893.13 | 40158 | -1.589 | -0.3286 | Yes | ||
| 160 | MYL9 | ENSG00000101335.10 | 40187 | -1.596 | -0.3230 | Yes | ||
| 161 | JAM3 | ENSG00000166086.13 | 40198 | -1.599 | -0.3169 | Yes | ||
| 162 | DLG1 | ENSG00000075711.21 | 40257 | -1.615 | -0.3119 | Yes | ||
| 163 | CLDN14 | ENSG00000159261.12 | 40281 | -1.620 | -0.3061 | Yes | ||
| 164 | HADH | ENSG00000138796.16 | 40435 | -1.667 | -0.3031 | Yes | ||
| 165 | MSN | ENSG00000147065.17 | 40565 | -1.706 | -0.2994 | Yes | ||
| 166 | TGFBI | ENSG00000120708.17 | 40715 | -1.752 | -0.2960 | Yes | ||
| 167 | ACTN1 | ENSG00000072110.13 | 40848 | -1.801 | -0.2919 | Yes | ||
| 168 | CDH4 | ENSG00000179242.16 | 40997 | -1.866 | -0.2881 | Yes | ||
| 169 | CD274 | ENSG00000120217.14 | 40999 | -1.867 | -0.2807 | Yes | ||
| 170 | CTNND1 | ENSG00000198561.13 | 41012 | -1.873 | -0.2736 | Yes | ||
| 171 | VCAM1 | ENSG00000162692.12 | 41228 | -1.969 | -0.2709 | Yes | ||
| 172 | MPP5 | ENSG00000072415.9 | 41456 | -2.104 | -0.2679 | Yes | ||
| 173 | CTNNA1 | ENSG00000044115.21 | 41488 | -2.125 | -0.2603 | Yes | ||
| 174 | NRXN2 | ENSG00000110076.18 | 41510 | -2.135 | -0.2523 | Yes | ||
| 175 | MYH9 | ENSG00000100345.21 | 41599 | -2.193 | -0.2458 | Yes | ||
| 176 | ACTG2 | ENSG00000163017.14 | 41741 | -2.285 | -0.2401 | Yes | ||
| 177 | RASA1 | ENSG00000145715.15 | 41787 | -2.315 | -0.2320 | Yes | ||
| 178 | ACTN4 | ENSG00000130402.12 | 41788 | -2.315 | -0.2229 | Yes | ||
| 179 | SLC30A3 | ENSG00000115194.11 | 41805 | -2.332 | -0.2141 | Yes | ||
| 180 | NECTIN2 | ENSG00000130202.10 | 41868 | -2.379 | -0.2062 | Yes | ||
| 181 | NECTIN4 | ENSG00000143217.9 | 41869 | -2.381 | -0.1968 | Yes | ||
| 182 | KCNH2 | ENSG00000055118.15 | 41985 | -2.477 | -0.1897 | Yes | ||
| 183 | PTPRC | ENSG00000081237.20 | 42203 | -2.706 | -0.1841 | Yes | ||
| 184 | AMIGO2 | ENSG00000139211.6 | 42276 | -2.821 | -0.1747 | Yes | ||
| 185 | AMH | ENSG00000104899.8 | 42282 | -2.829 | -0.1637 | Yes | ||
| 186 | INPPL1 | ENSG00000165458.14 | 42351 | -2.918 | -0.1538 | Yes | ||
| 187 | ARHGEF6 | ENSG00000129675.16 | 42382 | -2.958 | -0.1428 | Yes | ||
| 188 | ITGA3 | ENSG00000005884.18 | 42556 | -3.222 | -0.1342 | Yes | ||
| 189 | CDH8 | ENSG00000150394.14 | 42668 | -3.494 | -0.1230 | Yes | ||
| 190 | KRT31 | ENSG00000094796.5 | 42678 | -3.521 | -0.1094 | Yes | ||
| 191 | ACTA1 | ENSG00000143632.14 | 42721 | -3.653 | -0.0960 | Yes | ||
| 192 | CLDN5 | ENSG00000184113.9 | 42752 | -3.728 | -0.0820 | Yes | ||
| 193 | FLNC | ENSG00000128591.15 | 42852 | -4.113 | -0.0681 | Yes | ||
| 194 | ACTN2 | ENSG00000077522.13 | 42966 | -4.553 | -0.0528 | Yes | ||
| 195 | NEXN | ENSG00000162614.18 | 42973 | -4.596 | -0.0349 | Yes | ||
| 196 | CDH15 | ENSG00000129910.8 | 43003 | -4.739 | -0.0169 | Yes | ||
| 197 | ACTC1 | ENSG00000159251.8 | 43108 | -4.973 | 0.0003 | Yes |