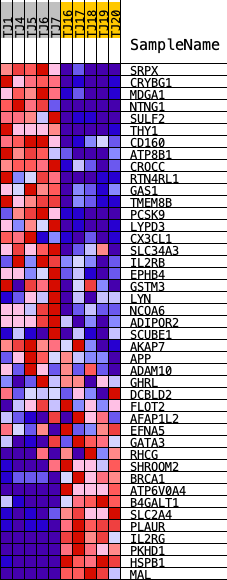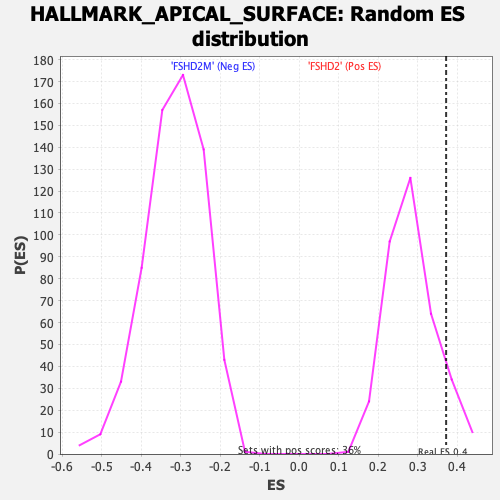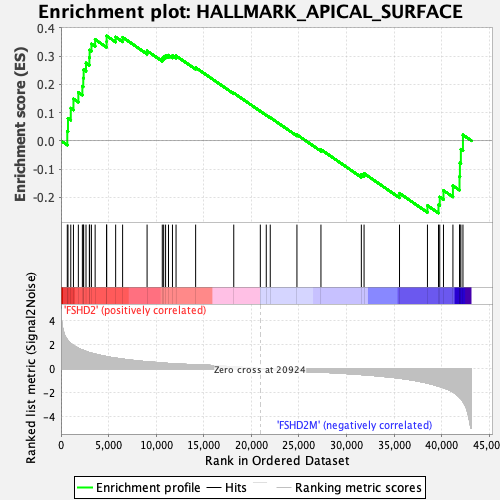
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_APICAL_SURFACE |
| Enrichment Score (ES) | 0.37158316 |
| Normalized Enrichment Score (NES) | 1.3101124 |
| Nominal p-value | 0.10674157 |
| FDR q-value | 0.06640269 |
| FWER p-Value | 0.478 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SRPX | ENSG00000101955.15 | 660 | 2.402 | 0.0337 | Yes | ||
| 2 | CRYBG1 | ENSG00000112297.15 | 734 | 2.336 | 0.0797 | Yes | ||
| 3 | MDGA1 | ENSG00000112139.16 | 1027 | 2.093 | 0.1156 | Yes | ||
| 4 | NTNG1 | ENSG00000162631.18 | 1308 | 1.933 | 0.1485 | Yes | ||
| 5 | SULF2 | ENSG00000196562.14 | 1823 | 1.703 | 0.1714 | Yes | ||
| 6 | THY1 | ENSG00000154096.13 | 2244 | 1.552 | 0.1933 | Yes | ||
| 7 | CD160 | ENSG00000117281.16 | 2339 | 1.522 | 0.2222 | Yes | ||
| 8 | ATP8B1 | ENSG00000081923.14 | 2380 | 1.510 | 0.2521 | Yes | ||
| 9 | CROCC | ENSG00000058453.17 | 2627 | 1.433 | 0.2756 | Yes | ||
| 10 | RTN4RL1 | ENSG00000185924.7 | 2979 | 1.337 | 0.2947 | Yes | ||
| 11 | GAS1 | ENSG00000180447.7 | 3007 | 1.331 | 0.3213 | Yes | ||
| 12 | TMEM8B | ENSG00000137103.20 | 3195 | 1.290 | 0.3432 | Yes | ||
| 13 | PCSK9 | ENSG00000169174.10 | 3586 | 1.208 | 0.3588 | Yes | ||
| 14 | LYPD3 | ENSG00000124466.9 | 4788 | 1.000 | 0.3514 | Yes | ||
| 15 | CX3CL1 | ENSG00000006210.7 | 4796 | 0.999 | 0.3716 | Yes | ||
| 16 | SLC34A3 | ENSG00000198569.9 | 5741 | 0.871 | 0.3674 | No | ||
| 17 | IL2RB | ENSG00000100385.14 | 6472 | 0.790 | 0.3666 | No | ||
| 18 | EPHB4 | ENSG00000196411.10 | 9043 | 0.567 | 0.3185 | No | ||
| 19 | GSTM3 | ENSG00000134202.11 | 10640 | 0.469 | 0.2911 | No | ||
| 20 | LYN | ENSG00000254087.8 | 10784 | 0.460 | 0.2971 | No | ||
| 21 | NCOA6 | ENSG00000198646.14 | 10986 | 0.446 | 0.3015 | No | ||
| 22 | ADIPOR2 | ENSG00000006831.10 | 11292 | 0.427 | 0.3032 | No | ||
| 23 | SCUBE1 | ENSG00000159307.19 | 11709 | 0.401 | 0.3017 | No | ||
| 24 | AKAP7 | ENSG00000118507.17 | 12085 | 0.383 | 0.3008 | No | ||
| 25 | APP | ENSG00000142192.21 | 14143 | 0.308 | 0.2594 | No | ||
| 26 | ADAM10 | ENSG00000137845.15 | 18147 | 0.139 | 0.1693 | No | ||
| 27 | GHRL | ENSG00000157017.16 | 20938 | -0.001 | 0.1045 | No | ||
| 28 | DCBLD2 | ENSG00000057019.16 | 21563 | -0.035 | 0.0908 | No | ||
| 29 | FLOT2 | ENSG00000132589.16 | 21981 | -0.056 | 0.0822 | No | ||
| 30 | AFAP1L2 | ENSG00000169129.15 | 24784 | -0.217 | 0.0216 | No | ||
| 31 | EFNA5 | ENSG00000184349.13 | 27306 | -0.296 | -0.0309 | No | ||
| 32 | GATA3 | ENSG00000107485.18 | 31550 | -0.493 | -0.1193 | No | ||
| 33 | RHCG | ENSG00000140519.14 | 31836 | -0.504 | -0.1157 | No | ||
| 34 | SHROOM2 | ENSG00000146950.13 | 35559 | -0.793 | -0.1859 | No | ||
| 35 | BRCA1 | ENSG00000012048.22 | 38494 | -1.205 | -0.2294 | No | ||
| 36 | ATP6V0A4 | ENSG00000105929.16 | 39655 | -1.455 | -0.2266 | No | ||
| 37 | B4GALT1 | ENSG00000086062.12 | 39786 | -1.489 | -0.1993 | No | ||
| 38 | SLC2A4 | ENSG00000181856.15 | 40178 | -1.594 | -0.1758 | No | ||
| 39 | PLAUR | ENSG00000011422.12 | 41170 | -1.945 | -0.1592 | No | ||
| 40 | IL2RG | ENSG00000147168.12 | 41862 | -2.374 | -0.1267 | No | ||
| 41 | PKHD1 | ENSG00000170927.15 | 41904 | -2.406 | -0.0786 | No | ||
| 42 | HSPB1 | ENSG00000106211.9 | 42005 | -2.494 | -0.0300 | No | ||
| 43 | MAL | ENSG00000172005.11 | 42224 | -2.738 | 0.0208 | No |