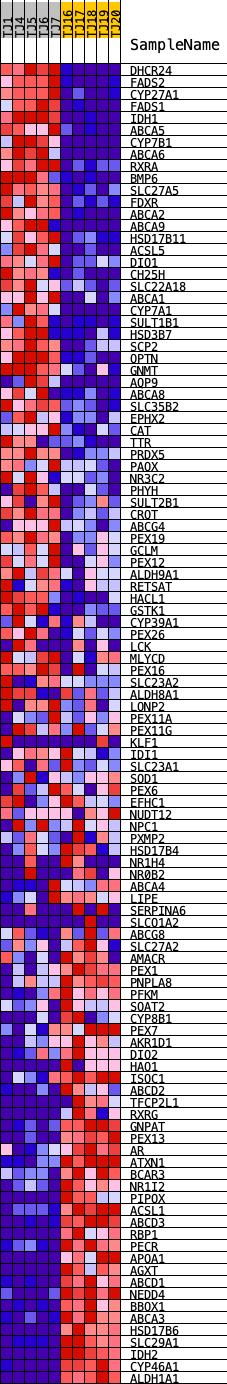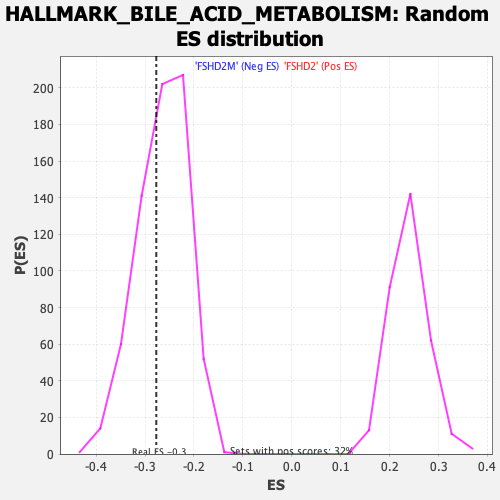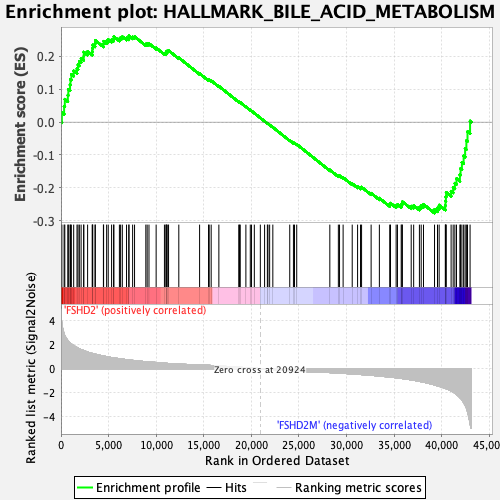
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_BILE_ACID_METABOLISM |
| Enrichment Score (ES) | -0.27693245 |
| Normalized Enrichment Score (NES) | -1.047249 |
| Nominal p-value | 0.3820059 |
| FDR q-value | 0.3674514 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DHCR24 | ENSG00000116133.13 | 91 | 3.768 | 0.0295 | No | ||
| 2 | FADS2 | ENSG00000134824.14 | 326 | 2.899 | 0.0483 | No | ||
| 3 | CYP27A1 | ENSG00000135929.9 | 400 | 2.753 | 0.0697 | No | ||
| 4 | FADS1 | ENSG00000149485.19 | 718 | 2.353 | 0.0820 | No | ||
| 5 | IDH1 | ENSG00000138413.13 | 771 | 2.290 | 0.1000 | No | ||
| 6 | ABCA5 | ENSG00000154265.16 | 950 | 2.133 | 0.1137 | No | ||
| 7 | CYP7B1 | ENSG00000172817.4 | 995 | 2.108 | 0.1304 | No | ||
| 8 | ABCA6 | ENSG00000154262.13 | 1087 | 2.056 | 0.1455 | No | ||
| 9 | RXRA | ENSG00000186350.12 | 1332 | 1.918 | 0.1559 | No | ||
| 10 | BMP6 | ENSG00000153162.9 | 1682 | 1.762 | 0.1625 | No | ||
| 11 | SLC27A5 | ENSG00000083807.10 | 1786 | 1.722 | 0.1746 | No | ||
| 12 | FDXR | ENSG00000161513.12 | 1927 | 1.658 | 0.1852 | No | ||
| 13 | ABCA2 | ENSG00000107331.17 | 2112 | 1.595 | 0.1943 | No | ||
| 14 | ABCA9 | ENSG00000154258.17 | 2386 | 1.509 | 0.2006 | No | ||
| 15 | HSD17B11 | ENSG00000198189.11 | 2389 | 1.508 | 0.2132 | No | ||
| 16 | ACSL5 | ENSG00000197142.10 | 2795 | 1.380 | 0.2153 | No | ||
| 17 | DIO1 | ENSG00000211452.10 | 3281 | 1.272 | 0.2147 | No | ||
| 18 | CH25H | ENSG00000138135.7 | 3303 | 1.268 | 0.2249 | No | ||
| 19 | SLC22A18 | ENSG00000110628.15 | 3336 | 1.257 | 0.2346 | No | ||
| 20 | ABCA1 | ENSG00000165029.16 | 3576 | 1.212 | 0.2392 | No | ||
| 21 | CYP7A1 | ENSG00000167910.4 | 3605 | 1.205 | 0.2487 | No | ||
| 22 | SULT1B1 | ENSG00000173597.9 | 4455 | 1.051 | 0.2378 | No | ||
| 23 | HSD3B7 | ENSG00000099377.14 | 4474 | 1.048 | 0.2461 | No | ||
| 24 | SCP2 | ENSG00000116171.18 | 4794 | 0.999 | 0.2471 | No | ||
| 25 | OPTN | ENSG00000123240.17 | 4961 | 0.974 | 0.2514 | No | ||
| 26 | GNMT | ENSG00000124713.6 | 5314 | 0.925 | 0.2509 | No | ||
| 27 | AQP9 | ENSG00000103569.10 | 5524 | 0.897 | 0.2536 | No | ||
| 28 | ABCA8 | ENSG00000141338.14 | 5557 | 0.893 | 0.2603 | No | ||
| 29 | SLC35B2 | ENSG00000157593.19 | 6134 | 0.825 | 0.2539 | No | ||
| 30 | EPHX2 | ENSG00000120915.14 | 6261 | 0.810 | 0.2577 | No | ||
| 31 | CAT | ENSG00000121691.7 | 6441 | 0.793 | 0.2602 | No | ||
| 32 | TTR | ENSG00000118271.10 | 6884 | 0.749 | 0.2562 | No | ||
| 33 | PRDX5 | ENSG00000126432.14 | 7130 | 0.725 | 0.2566 | No | ||
| 34 | PAOX | ENSG00000148832.16 | 7135 | 0.724 | 0.2626 | No | ||
| 35 | NR3C2 | ENSG00000151623.15 | 7524 | 0.688 | 0.2593 | No | ||
| 36 | PHYH | ENSG00000107537.14 | 7726 | 0.670 | 0.2602 | No | ||
| 37 | SULT2B1 | ENSG00000088002.11 | 8911 | 0.575 | 0.2375 | No | ||
| 38 | CROT | ENSG00000005469.11 | 9050 | 0.567 | 0.2391 | No | ||
| 39 | ABCG4 | ENSG00000172350.10 | 9242 | 0.552 | 0.2393 | No | ||
| 40 | PEX19 | ENSG00000162735.18 | 9994 | 0.512 | 0.2261 | No | ||
| 41 | GCLM | ENSG00000023909.10 | 10865 | 0.454 | 0.2097 | No | ||
| 42 | PEX12 | ENSG00000108733.10 | 11036 | 0.442 | 0.2094 | No | ||
| 43 | ALDH9A1 | ENSG00000143149.12 | 11047 | 0.441 | 0.2129 | No | ||
| 44 | RETSAT | ENSG00000042445.14 | 11072 | 0.439 | 0.2160 | No | ||
| 45 | HACL1 | ENSG00000131373.15 | 11215 | 0.432 | 0.2163 | No | ||
| 46 | GSTK1 | ENSG00000197448.14 | 11298 | 0.427 | 0.2180 | No | ||
| 47 | CYP39A1 | ENSG00000146233.8 | 12375 | 0.371 | 0.1961 | No | ||
| 48 | PEX26 | ENSG00000215193.12 | 14559 | 0.307 | 0.1479 | No | ||
| 49 | LCK | ENSG00000182866.17 | 15496 | 0.275 | 0.1284 | No | ||
| 50 | MLYCD | ENSG00000103150.7 | 15567 | 0.271 | 0.1291 | No | ||
| 51 | PEX16 | ENSG00000121680.16 | 15775 | 0.261 | 0.1264 | No | ||
| 52 | SLC23A2 | ENSG00000089057.15 | 16583 | 0.215 | 0.1095 | No | ||
| 53 | ALDH8A1 | ENSG00000118514.14 | 18672 | 0.114 | 0.0619 | No | ||
| 54 | LONP2 | ENSG00000102910.14 | 18749 | 0.109 | 0.0610 | No | ||
| 55 | PEX11A | ENSG00000166821.9 | 18829 | 0.105 | 0.0601 | No | ||
| 56 | PEX11G | ENSG00000104883.8 | 19421 | 0.076 | 0.0470 | No | ||
| 57 | KLF1 | ENSG00000105610.5 | 19884 | 0.052 | 0.0367 | No | ||
| 58 | IDI1 | ENSG00000067064.11 | 20014 | 0.048 | 0.0341 | No | ||
| 59 | SLC23A1 | ENSG00000170482.17 | 20308 | 0.031 | 0.0275 | No | ||
| 60 | SOD1 | ENSG00000142168.14 | 20939 | -0.001 | 0.0129 | No | ||
| 61 | PEX6 | ENSG00000124587.14 | 21386 | -0.025 | 0.0027 | No | ||
| 62 | EFHC1 | ENSG00000096093.16 | 21678 | -0.041 | -0.0037 | No | ||
| 63 | NUDT12 | ENSG00000112874.10 | 21740 | -0.044 | -0.0047 | No | ||
| 64 | NPC1 | ENSG00000141458.13 | 21917 | -0.052 | -0.0084 | No | ||
| 65 | PXMP2 | ENSG00000176894.10 | 22252 | -0.071 | -0.0156 | No | ||
| 66 | HSD17B4 | ENSG00000133835.16 | 24029 | -0.173 | -0.0554 | No | ||
| 67 | NR1H4 | ENSG00000012504.15 | 24416 | -0.194 | -0.0628 | No | ||
| 68 | NR0B2 | ENSG00000131910.5 | 24529 | -0.200 | -0.0637 | No | ||
| 69 | ABCA4 | ENSG00000198691.13 | 24762 | -0.216 | -0.0673 | No | ||
| 70 | LIPE | ENSG00000079435.10 | 28239 | -0.325 | -0.1454 | No | ||
| 71 | SERPINA6 | ENSG00000170099.6 | 29135 | -0.356 | -0.1632 | No | ||
| 72 | SLCO1A2 | ENSG00000084453.16 | 29259 | -0.360 | -0.1630 | No | ||
| 73 | ABCG8 | ENSG00000143921.9 | 29635 | -0.384 | -0.1685 | No | ||
| 74 | SLC27A2 | ENSG00000140284.11 | 30591 | -0.441 | -0.1870 | No | ||
| 75 | AMACR | ENSG00000242110.8 | 31155 | -0.472 | -0.1962 | No | ||
| 76 | PEX1 | ENSG00000127980.16 | 31464 | -0.488 | -0.1992 | No | ||
| 77 | PNPLA8 | ENSG00000135241.17 | 31574 | -0.494 | -0.1976 | No | ||
| 78 | PFKM | ENSG00000152556.17 | 32577 | -0.549 | -0.2163 | No | ||
| 79 | SOAT2 | ENSG00000167780.12 | 33443 | -0.616 | -0.2313 | No | ||
| 80 | CYP8B1 | ENSG00000180432.6 | 34531 | -0.704 | -0.2507 | No | ||
| 81 | PEX7 | ENSG00000112357.13 | 34613 | -0.712 | -0.2466 | No | ||
| 82 | AKR1D1 | ENSG00000122787.15 | 35205 | -0.760 | -0.2540 | No | ||
| 83 | DIO2 | ENSG00000211448.12 | 35343 | -0.772 | -0.2507 | No | ||
| 84 | HAO1 | ENSG00000101323.5 | 35737 | -0.812 | -0.2530 | No | ||
| 85 | ISOC1 | ENSG00000066583.12 | 35810 | -0.819 | -0.2478 | No | ||
| 86 | ABCD2 | ENSG00000173208.4 | 35860 | -0.826 | -0.2420 | No | ||
| 87 | TFCP2L1 | ENSG00000115112.8 | 36787 | -0.939 | -0.2557 | No | ||
| 88 | RXRG | ENSG00000143171.13 | 37046 | -0.974 | -0.2535 | No | ||
| 89 | GNPAT | ENSG00000116906.13 | 37649 | -1.059 | -0.2587 | No | ||
| 90 | PEX13 | ENSG00000162928.9 | 37837 | -1.089 | -0.2539 | No | ||
| 91 | AR | ENSG00000169083.17 | 38078 | -1.131 | -0.2500 | No | ||
| 92 | ATXN1 | ENSG00000124788.18 | 39238 | -1.362 | -0.2655 | Yes | ||
| 93 | BCAR3 | ENSG00000137936.18 | 39551 | -1.431 | -0.2608 | Yes | ||
| 94 | NR1I2 | ENSG00000144852.17 | 39723 | -1.472 | -0.2524 | Yes | ||
| 95 | PIPOX | ENSG00000179761.12 | 40370 | -1.647 | -0.2536 | Yes | ||
| 96 | ACSL1 | ENSG00000151726.14 | 40389 | -1.654 | -0.2402 | Yes | ||
| 97 | ABCD3 | ENSG00000117528.14 | 40428 | -1.663 | -0.2271 | Yes | ||
| 98 | RBP1 | ENSG00000114115.10 | 40461 | -1.676 | -0.2138 | Yes | ||
| 99 | PECR | ENSG00000115425.14 | 40979 | -1.853 | -0.2103 | Yes | ||
| 100 | APOA1 | ENSG00000118137.9 | 41195 | -1.956 | -0.1989 | Yes | ||
| 101 | AGXT | ENSG00000172482.5 | 41374 | -2.047 | -0.1859 | Yes | ||
| 102 | ABCD1 | ENSG00000101986.12 | 41544 | -2.159 | -0.1718 | Yes | ||
| 103 | NEDD4 | ENSG00000069869.16 | 41902 | -2.405 | -0.1599 | Yes | ||
| 104 | BBOX1 | ENSG00000129151.9 | 41973 | -2.470 | -0.1408 | Yes | ||
| 105 | ABCA3 | ENSG00000167972.14 | 42113 | -2.590 | -0.1224 | Yes | ||
| 106 | HSD17B6 | ENSG00000025423.11 | 42295 | -2.840 | -0.1028 | Yes | ||
| 107 | SLC29A1 | ENSG00000112759.18 | 42451 | -3.060 | -0.0807 | Yes | ||
| 108 | IDH2 | ENSG00000182054.9 | 42583 | -3.291 | -0.0562 | Yes | ||
| 109 | CYP46A1 | ENSG00000036530.9 | 42702 | -3.599 | -0.0288 | Yes | ||
| 110 | ALDH1A1 | ENSG00000165092.13 | 42972 | -4.592 | 0.0034 | Yes |