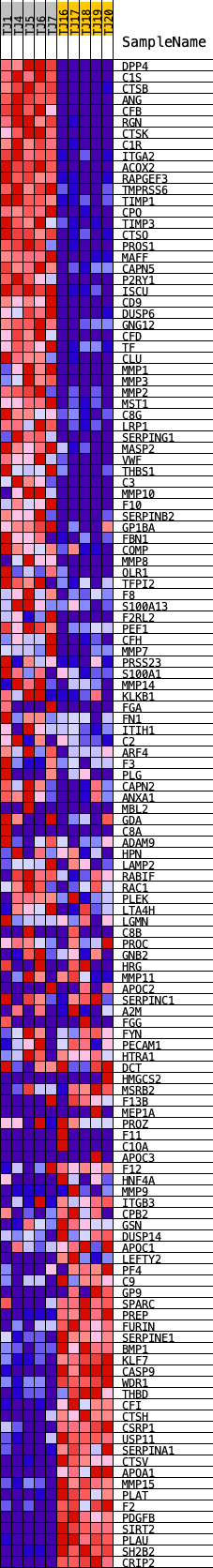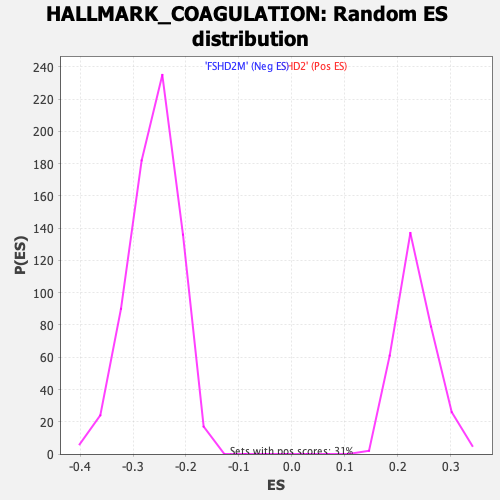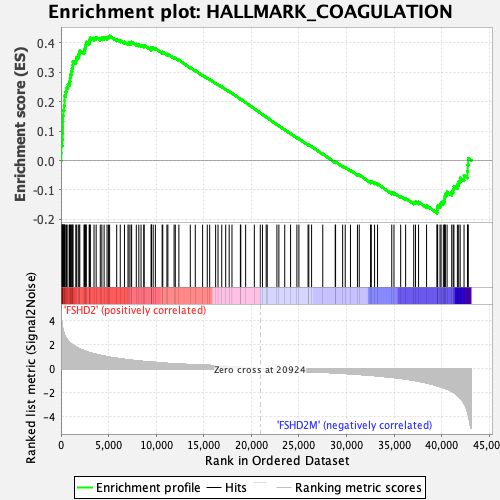
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_COAGULATION |
| Enrichment Score (ES) | 0.4239292 |
| Normalized Enrichment Score (NES) | 1.804934 |
| Nominal p-value | 0.0 |
| FDR q-value | 8.7152555E-4 |
| FWER p-Value | 0.004 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DPP4 | ENSG00000197635.10 | 30 | 4.288 | 0.0259 | Yes | ||
| 2 | C1S | ENSG00000182326.15 | 34 | 4.267 | 0.0523 | Yes | ||
| 3 | CTSB | ENSG00000164733.21 | 145 | 3.504 | 0.0715 | Yes | ||
| 4 | ANG | ENSG00000214274.9 | 174 | 3.361 | 0.0917 | Yes | ||
| 5 | CFB | ENSG00000243649.8 | 196 | 3.306 | 0.1118 | Yes | ||
| 6 | RGN | ENSG00000130988.13 | 200 | 3.302 | 0.1322 | Yes | ||
| 7 | CTSK | ENSG00000143387.13 | 210 | 3.273 | 0.1523 | Yes | ||
| 8 | C1R | ENSG00000159403.18 | 251 | 3.121 | 0.1708 | Yes | ||
| 9 | ITGA2 | ENSG00000164171.11 | 327 | 2.899 | 0.1870 | Yes | ||
| 10 | ACOX2 | ENSG00000168306.13 | 370 | 2.812 | 0.2035 | Yes | ||
| 11 | RAPGEF3 | ENSG00000079337.16 | 372 | 2.805 | 0.2209 | Yes | ||
| 12 | TMPRSS6 | ENSG00000187045.18 | 499 | 2.600 | 0.2341 | Yes | ||
| 13 | TIMP1 | ENSG00000102265.12 | 582 | 2.491 | 0.2476 | Yes | ||
| 14 | CPQ | ENSG00000104324.16 | 721 | 2.349 | 0.2590 | Yes | ||
| 15 | TIMP3 | ENSG00000100234.11 | 892 | 2.178 | 0.2686 | Yes | ||
| 16 | CTSO | ENSG00000256043.3 | 953 | 2.131 | 0.2804 | Yes | ||
| 17 | PROS1 | ENSG00000184500.15 | 997 | 2.107 | 0.2925 | Yes | ||
| 18 | MAFF | ENSG00000185022.12 | 1086 | 2.057 | 0.3032 | Yes | ||
| 19 | CAPN5 | ENSG00000149260.18 | 1156 | 2.013 | 0.3141 | Yes | ||
| 20 | P2RY1 | ENSG00000169860.7 | 1233 | 1.973 | 0.3246 | Yes | ||
| 21 | ISCU | ENSG00000136003.15 | 1254 | 1.961 | 0.3363 | Yes | ||
| 22 | CD9 | ENSG00000010278.14 | 1562 | 1.814 | 0.3404 | Yes | ||
| 23 | DUSP6 | ENSG00000139318.8 | 1611 | 1.795 | 0.3504 | Yes | ||
| 24 | GNG12 | ENSG00000172380.6 | 1804 | 1.714 | 0.3566 | Yes | ||
| 25 | CFD | ENSG00000197766.8 | 1895 | 1.671 | 0.3649 | Yes | ||
| 26 | TF | ENSG00000091513.16 | 1973 | 1.641 | 0.3733 | Yes | ||
| 27 | CLU | ENSG00000120885.22 | 2415 | 1.500 | 0.3724 | Yes | ||
| 28 | MMP1 | ENSG00000196611.5 | 2473 | 1.481 | 0.3802 | Yes | ||
| 29 | MMP3 | ENSG00000149968.12 | 2562 | 1.453 | 0.3872 | Yes | ||
| 30 | MMP2 | ENSG00000087245.13 | 2616 | 1.437 | 0.3949 | Yes | ||
| 31 | MST1 | ENSG00000173531.15 | 2650 | 1.426 | 0.4030 | Yes | ||
| 32 | C8G | ENSG00000176919.13 | 2959 | 1.343 | 0.4041 | Yes | ||
| 33 | LRP1 | ENSG00000123384.14 | 3000 | 1.333 | 0.4115 | Yes | ||
| 34 | SERPING1 | ENSG00000149131.15 | 3087 | 1.313 | 0.4176 | Yes | ||
| 35 | MASP2 | ENSG00000009724.17 | 3469 | 1.231 | 0.4164 | Yes | ||
| 36 | VWF | ENSG00000110799.13 | 3693 | 1.186 | 0.4186 | Yes | ||
| 37 | THBS1 | ENSG00000137801.11 | 4127 | 1.107 | 0.4154 | Yes | ||
| 38 | C3 | ENSG00000125730.17 | 4292 | 1.078 | 0.4183 | Yes | ||
| 39 | MMP10 | ENSG00000166670.10 | 4529 | 1.040 | 0.4192 | Yes | ||
| 40 | F10 | ENSG00000126218.12 | 4837 | 0.991 | 0.4182 | Yes | ||
| 41 | SERPINB2 | ENSG00000197632.9 | 4983 | 0.970 | 0.4209 | Yes | ||
| 42 | GP1BA | ENSG00000185245.8 | 5108 | 0.953 | 0.4239 | Yes | ||
| 43 | FBN1 | ENSG00000166147.13 | 5850 | 0.858 | 0.4120 | No | ||
| 44 | COMP | ENSG00000105664.11 | 6225 | 0.814 | 0.4084 | No | ||
| 45 | MMP8 | ENSG00000118113.12 | 6665 | 0.771 | 0.4029 | No | ||
| 46 | OLR1 | ENSG00000173391.9 | 7038 | 0.734 | 0.3988 | No | ||
| 47 | TFPI2 | ENSG00000105825.14 | 7157 | 0.723 | 0.4006 | No | ||
| 48 | F8 | ENSG00000185010.15 | 7356 | 0.705 | 0.4004 | No | ||
| 49 | S100A13 | ENSG00000189171.14 | 7420 | 0.698 | 0.4032 | No | ||
| 50 | F2RL2 | ENSG00000164220.7 | 7910 | 0.654 | 0.3959 | No | ||
| 51 | PEF1 | ENSG00000162517.13 | 8174 | 0.631 | 0.3937 | No | ||
| 52 | CFH | ENSG00000000971.15 | 8408 | 0.612 | 0.3921 | No | ||
| 53 | MMP7 | ENSG00000137673.9 | 8697 | 0.591 | 0.3891 | No | ||
| 54 | PRSS23 | ENSG00000150687.12 | 8732 | 0.588 | 0.3919 | No | ||
| 55 | S100A1 | ENSG00000160678.11 | 9481 | 0.539 | 0.3779 | No | ||
| 56 | MMP14 | ENSG00000157227.13 | 9489 | 0.539 | 0.3811 | No | ||
| 57 | KLKB1 | ENSG00000164344.16 | 9496 | 0.538 | 0.3843 | No | ||
| 58 | FGA | ENSG00000171560.16 | 9683 | 0.524 | 0.3832 | No | ||
| 59 | FN1 | ENSG00000115414.19 | 9913 | 0.516 | 0.3811 | No | ||
| 60 | ITIH1 | ENSG00000055957.11 | 10626 | 0.470 | 0.3674 | No | ||
| 61 | C2 | ENSG00000166278.15 | 10689 | 0.466 | 0.3689 | No | ||
| 62 | ARF4 | ENSG00000168374.11 | 11127 | 0.437 | 0.3614 | No | ||
| 63 | F3 | ENSG00000117525.14 | 11227 | 0.431 | 0.3618 | No | ||
| 64 | PLG | ENSG00000122194.18 | 11880 | 0.394 | 0.3491 | No | ||
| 65 | CAPN2 | ENSG00000162909.18 | 12007 | 0.388 | 0.3485 | No | ||
| 66 | ANXA1 | ENSG00000135046.14 | 12384 | 0.370 | 0.3421 | No | ||
| 67 | MBL2 | ENSG00000165471.6 | 13582 | 0.322 | 0.3162 | No | ||
| 68 | GDA | ENSG00000119125.16 | 14105 | 0.311 | 0.3060 | No | ||
| 69 | C8A | ENSG00000157131.10 | 14876 | 0.287 | 0.2899 | No | ||
| 70 | ADAM9 | ENSG00000168615.12 | 15364 | 0.282 | 0.2803 | No | ||
| 71 | HPN | ENSG00000105707.14 | 15619 | 0.269 | 0.2761 | No | ||
| 72 | LAMP2 | ENSG00000005893.15 | 16244 | 0.233 | 0.2630 | No | ||
| 73 | RABIF | ENSG00000183155.5 | 16494 | 0.220 | 0.2586 | No | ||
| 74 | RAC1 | ENSG00000136238.18 | 16886 | 0.203 | 0.2507 | No | ||
| 75 | PLEK | ENSG00000115956.10 | 17299 | 0.183 | 0.2423 | No | ||
| 76 | LTA4H | ENSG00000111144.10 | 17667 | 0.165 | 0.2348 | No | ||
| 77 | LGMN | ENSG00000100600.15 | 17960 | 0.148 | 0.2289 | No | ||
| 78 | C8B | ENSG00000021852.13 | 18843 | 0.105 | 0.2090 | No | ||
| 79 | PROC | ENSG00000115718.17 | 18885 | 0.104 | 0.2087 | No | ||
| 80 | GNB2 | ENSG00000172354.10 | 19396 | 0.077 | 0.1973 | No | ||
| 81 | HRG | ENSG00000113905.4 | 20309 | 0.031 | 0.1763 | No | ||
| 82 | MMP11 | ENSG00000099953.10 | 20933 | -0.001 | 0.1618 | No | ||
| 83 | APOC2 | ENSG00000234906.10 | 21160 | -0.014 | 0.1567 | No | ||
| 84 | SERPINC1 | ENSG00000117601.13 | 21552 | -0.034 | 0.1478 | No | ||
| 85 | A2M | ENSG00000175899.14 | 21664 | -0.040 | 0.1454 | No | ||
| 86 | FGG | ENSG00000171557.17 | 22680 | -0.095 | 0.1224 | No | ||
| 87 | FYN | ENSG00000010810.17 | 22887 | -0.107 | 0.1183 | No | ||
| 88 | PECAM1 | ENSG00000261371.6 | 23502 | -0.141 | 0.1049 | No | ||
| 89 | HTRA1 | ENSG00000166033.13 | 24116 | -0.177 | 0.0917 | No | ||
| 90 | DCT | ENSG00000080166.16 | 24780 | -0.217 | 0.0777 | No | ||
| 91 | HMGCS2 | ENSG00000134240.11 | 24990 | -0.223 | 0.0742 | No | ||
| 92 | MSRB2 | ENSG00000148450.13 | 25953 | -0.258 | 0.0534 | No | ||
| 93 | F13B | ENSG00000143278.4 | 26031 | -0.262 | 0.0532 | No | ||
| 94 | MEP1A | ENSG00000112818.10 | 26323 | -0.263 | 0.0481 | No | ||
| 95 | PROZ | ENSG00000126231.14 | 27498 | -0.297 | 0.0226 | No | ||
| 96 | F11 | ENSG00000088926.13 | 28806 | -0.351 | -0.0056 | No | ||
| 97 | C1QA | ENSG00000173372.17 | 28844 | -0.351 | -0.0043 | No | ||
| 98 | APOC3 | ENSG00000110245.12 | 29586 | -0.380 | -0.0192 | No | ||
| 99 | F12 | ENSG00000131187.9 | 29857 | -0.394 | -0.0230 | No | ||
| 100 | HNF4A | ENSG00000101076.16 | 30407 | -0.432 | -0.0331 | No | ||
| 101 | MMP9 | ENSG00000100985.7 | 31139 | -0.471 | -0.0472 | No | ||
| 102 | ITGB3 | ENSG00000259207.7 | 31330 | -0.481 | -0.0486 | No | ||
| 103 | CPB2 | ENSG00000080618.16 | 32522 | -0.544 | -0.0729 | No | ||
| 104 | GSN | ENSG00000148180.19 | 32592 | -0.551 | -0.0711 | No | ||
| 105 | DUSP14 | ENSG00000276023.5 | 32940 | -0.578 | -0.0756 | No | ||
| 106 | APOC1 | ENSG00000130208.9 | 33251 | -0.601 | -0.0791 | No | ||
| 107 | LEFTY2 | ENSG00000143768.13 | 34745 | -0.720 | -0.1093 | No | ||
| 108 | PF4 | ENSG00000163737.4 | 34978 | -0.739 | -0.1101 | No | ||
| 109 | C9 | ENSG00000113600.11 | 35681 | -0.807 | -0.1215 | No | ||
| 110 | GP9 | ENSG00000169704.4 | 36202 | -0.868 | -0.1282 | No | ||
| 111 | SPARC | ENSG00000113140.11 | 37052 | -0.974 | -0.1419 | No | ||
| 112 | PREP | ENSG00000085377.14 | 37237 | -0.997 | -0.1400 | No | ||
| 113 | FURIN | ENSG00000140564.12 | 37563 | -1.044 | -0.1410 | No | ||
| 114 | SERPINE1 | ENSG00000106366.8 | 38403 | -1.189 | -0.1532 | No | ||
| 115 | BMP1 | ENSG00000168487.19 | 39505 | -1.423 | -0.1700 | No | ||
| 116 | KLF7 | ENSG00000118263.15 | 39515 | -1.424 | -0.1613 | No | ||
| 117 | CASP9 | ENSG00000132906.18 | 39565 | -1.434 | -0.1536 | No | ||
| 118 | WDR1 | ENSG00000071127.17 | 39815 | -1.497 | -0.1501 | No | ||
| 119 | THBD | ENSG00000178726.6 | 39927 | -1.529 | -0.1431 | No | ||
| 120 | CFI | ENSG00000205403.13 | 40164 | -1.591 | -0.1388 | No | ||
| 121 | CTSH | ENSG00000103811.16 | 40296 | -1.626 | -0.1317 | No | ||
| 122 | CSRP1 | ENSG00000159176.14 | 40304 | -1.628 | -0.1218 | No | ||
| 123 | USP11 | ENSG00000102226.9 | 40376 | -1.650 | -0.1132 | No | ||
| 124 | SERPINA1 | ENSG00000197249.13 | 40558 | -1.704 | -0.1068 | No | ||
| 125 | CTSV | ENSG00000136943.11 | 41043 | -1.887 | -0.1064 | No | ||
| 126 | APOA1 | ENSG00000118137.9 | 41195 | -1.956 | -0.0977 | No | ||
| 127 | MMP15 | ENSG00000102996.5 | 41275 | -1.992 | -0.0872 | No | ||
| 128 | PLAT | ENSG00000104368.18 | 41640 | -2.216 | -0.0819 | No | ||
| 129 | F2 | ENSG00000180210.14 | 41777 | -2.308 | -0.0707 | No | ||
| 130 | PDGFB | ENSG00000100311.17 | 41932 | -2.434 | -0.0592 | No | ||
| 131 | SIRT2 | ENSG00000068903.20 | 42338 | -2.901 | -0.0506 | No | ||
| 132 | PLAU | ENSG00000122861.16 | 42698 | -3.591 | -0.0367 | No | ||
| 133 | SH2B2 | ENSG00000160999.10 | 42724 | -3.659 | -0.0146 | No | ||
| 134 | CRIP2 | ENSG00000182809.10 | 42778 | -3.821 | 0.0079 | No |