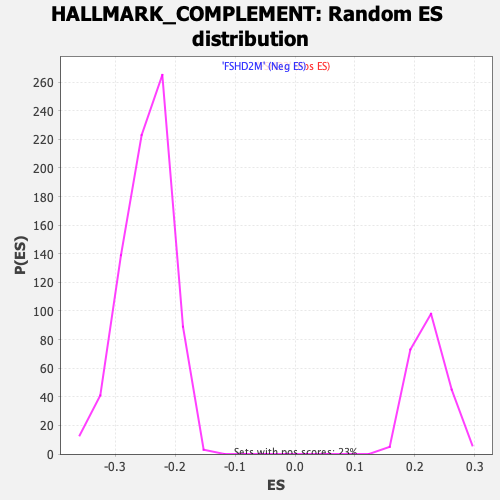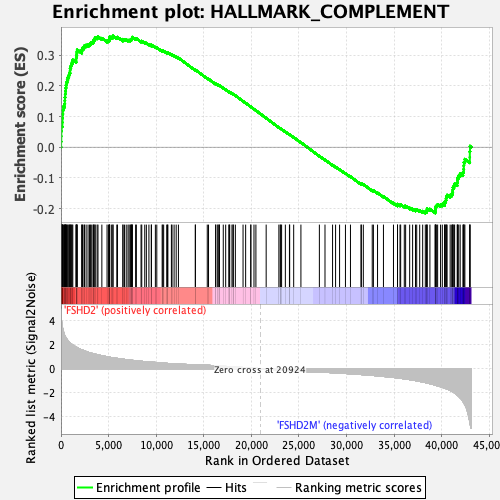
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_COMPLEMENT |
| Enrichment Score (ES) | 0.3629188 |
| Normalized Enrichment Score (NES) | 1.6337557 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.006208492 |
| FWER p-Value | 0.033 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CASP10 | ENSG00000003400.14 | 10 | 4.582 | 0.0184 | Yes | ||
| 2 | DPP4 | ENSG00000197635.10 | 30 | 4.288 | 0.0355 | Yes | ||
| 3 | C1S | ENSG00000182326.15 | 34 | 4.267 | 0.0528 | Yes | ||
| 4 | KCNIP3 | ENSG00000115041.13 | 68 | 3.944 | 0.0681 | Yes | ||
| 5 | CTSB | ENSG00000164733.21 | 145 | 3.504 | 0.0806 | Yes | ||
| 6 | LGALS3 | ENSG00000131981.16 | 150 | 3.486 | 0.0948 | Yes | ||
| 7 | ANG | ENSG00000214274.9 | 174 | 3.361 | 0.1079 | Yes | ||
| 8 | CFB | ENSG00000243649.8 | 196 | 3.306 | 0.1209 | Yes | ||
| 9 | C1R | ENSG00000159403.18 | 251 | 3.121 | 0.1324 | Yes | ||
| 10 | PRCP | ENSG00000137509.11 | 377 | 2.794 | 0.1409 | Yes | ||
| 11 | TIMP2 | ENSG00000035862.12 | 402 | 2.753 | 0.1515 | Yes | ||
| 12 | APOBEC3F | ENSG00000128394.17 | 410 | 2.739 | 0.1625 | Yes | ||
| 13 | CTSS | ENSG00000163131.11 | 442 | 2.687 | 0.1728 | Yes | ||
| 14 | CTSL | ENSG00000135047.15 | 458 | 2.665 | 0.1833 | Yes | ||
| 15 | TMPRSS6 | ENSG00000187045.18 | 499 | 2.600 | 0.1929 | Yes | ||
| 16 | LIPA | ENSG00000107798.18 | 528 | 2.571 | 0.2028 | Yes | ||
| 17 | TIMP1 | ENSG00000102265.12 | 582 | 2.491 | 0.2117 | Yes | ||
| 18 | PSMB9 | ENSG00000240065.8 | 653 | 2.409 | 0.2199 | Yes | ||
| 19 | CPQ | ENSG00000104324.16 | 721 | 2.349 | 0.2279 | Yes | ||
| 20 | PRSS36 | ENSG00000178226.11 | 819 | 2.244 | 0.2348 | Yes | ||
| 21 | APOBEC3G | ENSG00000239713.9 | 876 | 2.186 | 0.2424 | Yes | ||
| 22 | CTSO | ENSG00000256043.3 | 953 | 2.131 | 0.2493 | Yes | ||
| 23 | CALM3 | ENSG00000160014.17 | 958 | 2.130 | 0.2579 | Yes | ||
| 24 | PLSCR1 | ENSG00000188313.13 | 1031 | 2.090 | 0.2647 | Yes | ||
| 25 | MAFF | ENSG00000185022.12 | 1086 | 2.057 | 0.2719 | Yes | ||
| 26 | SRC | ENSG00000197122.11 | 1168 | 2.008 | 0.2782 | Yes | ||
| 27 | CEBPB | ENSG00000172216.6 | 1238 | 1.972 | 0.2846 | Yes | ||
| 28 | CD46 | ENSG00000117335.19 | 1590 | 1.805 | 0.2838 | Yes | ||
| 29 | CP | ENSG00000047457.14 | 1593 | 1.802 | 0.2911 | Yes | ||
| 30 | DUSP6 | ENSG00000139318.8 | 1611 | 1.795 | 0.2980 | Yes | ||
| 31 | CD59 | ENSG00000085063.16 | 1662 | 1.770 | 0.3041 | Yes | ||
| 32 | DUSP5 | ENSG00000138166.6 | 1674 | 1.764 | 0.3110 | Yes | ||
| 33 | IRF2 | ENSG00000168310.11 | 1700 | 1.754 | 0.3176 | Yes | ||
| 34 | CASP5 | ENSG00000137757.11 | 2183 | 1.572 | 0.3127 | Yes | ||
| 35 | CD36 | ENSG00000135218.19 | 2200 | 1.566 | 0.3188 | Yes | ||
| 36 | CASP4 | ENSG00000196954.14 | 2254 | 1.548 | 0.3238 | Yes | ||
| 37 | CLU | ENSG00000120885.22 | 2415 | 1.500 | 0.3262 | Yes | ||
| 38 | RHOG | ENSG00000177105.10 | 2462 | 1.485 | 0.3312 | Yes | ||
| 39 | CPM | ENSG00000135678.12 | 2638 | 1.429 | 0.3330 | Yes | ||
| 40 | CASP1 | ENSG00000137752.24 | 2840 | 1.369 | 0.3339 | Yes | ||
| 41 | LRP1 | ENSG00000123384.14 | 3000 | 1.333 | 0.3356 | Yes | ||
| 42 | SERPING1 | ENSG00000149131.15 | 3087 | 1.313 | 0.3389 | Yes | ||
| 43 | RASGRP1 | ENSG00000172575.12 | 3180 | 1.292 | 0.3421 | Yes | ||
| 44 | DOCK4 | ENSG00000128512.22 | 3363 | 1.252 | 0.3429 | Yes | ||
| 45 | PRSS3 | ENSG00000010438.16 | 3403 | 1.245 | 0.3471 | Yes | ||
| 46 | IRF1 | ENSG00000125347.14 | 3458 | 1.235 | 0.3509 | Yes | ||
| 47 | PHEX | ENSG00000102174.9 | 3561 | 1.215 | 0.3534 | Yes | ||
| 48 | PCSK9 | ENSG00000169174.10 | 3586 | 1.208 | 0.3578 | Yes | ||
| 49 | GPD2 | ENSG00000115159.16 | 3791 | 1.167 | 0.3578 | Yes | ||
| 50 | PDP1 | ENSG00000164951.16 | 3884 | 1.148 | 0.3604 | Yes | ||
| 51 | C3 | ENSG00000125730.17 | 4292 | 1.078 | 0.3553 | Yes | ||
| 52 | F10 | ENSG00000126218.12 | 4837 | 0.991 | 0.3466 | Yes | ||
| 53 | SERPINB2 | ENSG00000197632.9 | 4983 | 0.970 | 0.3472 | Yes | ||
| 54 | CR1 | ENSG00000203710.11 | 5061 | 0.959 | 0.3493 | Yes | ||
| 55 | GNAI2 | ENSG00000114353.17 | 5091 | 0.956 | 0.3526 | Yes | ||
| 56 | GP1BA | ENSG00000185245.8 | 5108 | 0.953 | 0.3561 | Yes | ||
| 57 | CD55 | ENSG00000196352.15 | 5115 | 0.952 | 0.3598 | Yes | ||
| 58 | DOCK10 | ENSG00000135905.19 | 5321 | 0.924 | 0.3588 | Yes | ||
| 59 | ANXA5 | ENSG00000164111.15 | 5451 | 0.906 | 0.3595 | Yes | ||
| 60 | AC016586.1 | ENSG00000268670.1 | 5463 | 0.904 | 0.3629 | Yes | ||
| 61 | PRKCD | ENSG00000163932.15 | 5863 | 0.856 | 0.3571 | No | ||
| 62 | FDX1 | ENSG00000137714.3 | 5937 | 0.849 | 0.3589 | No | ||
| 63 | PLA2G4A | ENSG00000116711.10 | 6498 | 0.788 | 0.3490 | No | ||
| 64 | NOTCH4 | ENSG00000204301.6 | 6513 | 0.787 | 0.3519 | No | ||
| 65 | MMP8 | ENSG00000118113.12 | 6665 | 0.771 | 0.3515 | No | ||
| 66 | CDA | ENSG00000158825.6 | 6850 | 0.752 | 0.3503 | No | ||
| 67 | OLR1 | ENSG00000173391.9 | 7038 | 0.734 | 0.3490 | No | ||
| 68 | TFPI2 | ENSG00000105825.14 | 7157 | 0.723 | 0.3492 | No | ||
| 69 | KYNU | ENSG00000115919.15 | 7285 | 0.711 | 0.3491 | No | ||
| 70 | F8 | ENSG00000185010.15 | 7356 | 0.705 | 0.3503 | No | ||
| 71 | S100A13 | ENSG00000189171.14 | 7420 | 0.698 | 0.3517 | No | ||
| 72 | GNAI3 | ENSG00000065135.11 | 7451 | 0.695 | 0.3539 | No | ||
| 73 | CTSC | ENSG00000109861.16 | 7476 | 0.693 | 0.3561 | No | ||
| 74 | C4BPB | ENSG00000123843.13 | 7483 | 0.692 | 0.3588 | No | ||
| 75 | ATOX1 | ENSG00000177556.12 | 7840 | 0.661 | 0.3532 | No | ||
| 76 | DGKH | ENSG00000102780.16 | 7915 | 0.654 | 0.3541 | No | ||
| 77 | CFH | ENSG00000000971.15 | 8408 | 0.612 | 0.3452 | No | ||
| 78 | FCN1 | ENSG00000085265.11 | 8484 | 0.608 | 0.3459 | No | ||
| 79 | CCL5 | ENSG00000271503.6 | 8798 | 0.583 | 0.3410 | No | ||
| 80 | HSPA1A | ENSG00000204389.10 | 8966 | 0.570 | 0.3394 | No | ||
| 81 | SH2B3 | ENSG00000111252.10 | 9235 | 0.553 | 0.3354 | No | ||
| 82 | MMP14 | ENSG00000157227.13 | 9489 | 0.539 | 0.3317 | No | ||
| 83 | KLKB1 | ENSG00000164344.16 | 9496 | 0.538 | 0.3338 | No | ||
| 84 | FN1 | ENSG00000115414.19 | 9913 | 0.516 | 0.3262 | No | ||
| 85 | F7 | ENSG00000057593.13 | 10063 | 0.507 | 0.3248 | No | ||
| 86 | ITIH1 | ENSG00000055957.11 | 10626 | 0.470 | 0.3136 | No | ||
| 87 | C2 | ENSG00000166278.15 | 10689 | 0.466 | 0.3141 | No | ||
| 88 | LYN | ENSG00000254087.8 | 10784 | 0.460 | 0.3138 | No | ||
| 89 | LCP2 | ENSG00000043462.12 | 11106 | 0.438 | 0.3081 | No | ||
| 90 | VCPIP1 | ENSG00000175073.8 | 11186 | 0.433 | 0.3080 | No | ||
| 91 | F3 | ENSG00000117525.14 | 11227 | 0.431 | 0.3088 | No | ||
| 92 | GCA | ENSG00000115271.11 | 11596 | 0.408 | 0.3019 | No | ||
| 93 | USP8 | ENSG00000138592.14 | 11662 | 0.404 | 0.3020 | No | ||
| 94 | PLG | ENSG00000122194.18 | 11880 | 0.394 | 0.2986 | No | ||
| 95 | AKAP10 | ENSG00000108599.15 | 12100 | 0.382 | 0.2950 | No | ||
| 96 | ERAP2 | ENSG00000164308.16 | 12335 | 0.373 | 0.2911 | No | ||
| 97 | IRF7 | ENSG00000185507.21 | 14110 | 0.311 | 0.2511 | No | ||
| 98 | RNF4 | ENSG00000063978.16 | 14118 | 0.310 | 0.2522 | No | ||
| 99 | ADAM9 | ENSG00000168615.12 | 15364 | 0.282 | 0.2243 | No | ||
| 100 | LCK | ENSG00000182866.17 | 15496 | 0.275 | 0.2224 | No | ||
| 101 | LAMP2 | ENSG00000005893.15 | 16244 | 0.233 | 0.2059 | No | ||
| 102 | KLK1 | ENSG00000167748.11 | 16260 | 0.232 | 0.2065 | No | ||
| 103 | GNB4 | ENSG00000114450.10 | 16478 | 0.221 | 0.2024 | No | ||
| 104 | RABIF | ENSG00000183155.5 | 16494 | 0.220 | 0.2029 | No | ||
| 105 | WAS | ENSG00000015285.10 | 16612 | 0.214 | 0.2010 | No | ||
| 106 | PIM1 | ENSG00000137193.14 | 16645 | 0.213 | 0.2012 | No | ||
| 107 | CXCL1 | ENSG00000163739.5 | 17045 | 0.196 | 0.1927 | No | ||
| 108 | PLEK | ENSG00000115956.10 | 17299 | 0.183 | 0.1875 | No | ||
| 109 | CD40LG | ENSG00000102245.7 | 17639 | 0.167 | 0.1803 | No | ||
| 110 | LTA4H | ENSG00000111144.10 | 17667 | 0.165 | 0.1804 | No | ||
| 111 | PLA2G7 | ENSG00000146070.17 | 17722 | 0.162 | 0.1798 | No | ||
| 112 | LGMN | ENSG00000100600.15 | 17960 | 0.148 | 0.1748 | No | ||
| 113 | RBSN | ENSG00000131381.12 | 18086 | 0.142 | 0.1725 | No | ||
| 114 | CTSD | ENSG00000117984.14 | 18089 | 0.141 | 0.1730 | No | ||
| 115 | MMP12 | ENSG00000262406.3 | 18315 | 0.129 | 0.1683 | No | ||
| 116 | PRDM4 | ENSG00000110851.12 | 19123 | 0.093 | 0.1499 | No | ||
| 117 | GNB2 | ENSG00000172354.10 | 19396 | 0.077 | 0.1439 | No | ||
| 118 | RCE1 | ENSG00000173653.8 | 19938 | 0.051 | 0.1315 | No | ||
| 119 | PIK3CA | ENSG00000121879.5 | 19940 | 0.051 | 0.1317 | No | ||
| 120 | PSEN1 | ENSG00000080815.19 | 20272 | 0.034 | 0.1241 | No | ||
| 121 | GZMK | ENSG00000113088.6 | 20474 | 0.024 | 0.1195 | No | ||
| 122 | SERPINC1 | ENSG00000117601.13 | 21552 | -0.034 | 0.0945 | No | ||
| 123 | FYN | ENSG00000010810.17 | 22887 | -0.107 | 0.0639 | No | ||
| 124 | CASP7 | ENSG00000165806.20 | 23029 | -0.113 | 0.0611 | No | ||
| 125 | STX4 | ENSG00000103496.15 | 23081 | -0.117 | 0.0604 | No | ||
| 126 | GNG2 | ENSG00000186469.8 | 23173 | -0.123 | 0.0588 | No | ||
| 127 | PIK3R5 | ENSG00000141506.13 | 23567 | -0.145 | 0.0502 | No | ||
| 128 | PPP4C | ENSG00000149923.14 | 23997 | -0.171 | 0.0409 | No | ||
| 129 | F5 | ENSG00000198734.11 | 24021 | -0.173 | 0.0411 | No | ||
| 130 | SCG3 | ENSG00000104112.9 | 24442 | -0.195 | 0.0321 | No | ||
| 131 | GZMA | ENSG00000145649.8 | 25202 | -0.223 | 0.0153 | No | ||
| 132 | BRPF3 | ENSG00000096070.19 | 27141 | -0.283 | -0.0287 | No | ||
| 133 | APOA4 | ENSG00000110244.7 | 27731 | -0.302 | -0.0412 | No | ||
| 134 | GMFB | ENSG00000197045.13 | 28527 | -0.338 | -0.0583 | No | ||
| 135 | C1QA | ENSG00000173372.17 | 28844 | -0.351 | -0.0643 | No | ||
| 136 | L3MBTL4 | ENSG00000154655.15 | 29264 | -0.360 | -0.0726 | No | ||
| 137 | LAP3 | ENSG00000002549.12 | 29880 | -0.396 | -0.0853 | No | ||
| 138 | HNF4A | ENSG00000101076.16 | 30407 | -0.432 | -0.0958 | No | ||
| 139 | CR2 | ENSG00000117322.17 | 31514 | -0.489 | -0.1195 | No | ||
| 140 | GATA3 | ENSG00000107485.18 | 31550 | -0.493 | -0.1183 | No | ||
| 141 | JAK2 | ENSG00000096968.13 | 31758 | -0.501 | -0.1211 | No | ||
| 142 | RAF1 | ENSG00000132155.11 | 32694 | -0.559 | -0.1406 | No | ||
| 143 | SPOCK2 | ENSG00000107742.13 | 32816 | -0.570 | -0.1411 | No | ||
| 144 | APOC1 | ENSG00000130208.9 | 33251 | -0.601 | -0.1488 | No | ||
| 145 | USP16 | ENSG00000156256.15 | 33872 | -0.651 | -0.1606 | No | ||
| 146 | PPP2CB | ENSG00000104695.13 | 34925 | -0.735 | -0.1821 | No | ||
| 147 | GZMB | ENSG00000100453.13 | 35349 | -0.772 | -0.1888 | No | ||
| 148 | CBLB | ENSG00000114423.22 | 35351 | -0.773 | -0.1857 | No | ||
| 149 | USP15 | ENSG00000135655.16 | 35611 | -0.799 | -0.1884 | No | ||
| 150 | C9 | ENSG00000113600.11 | 35681 | -0.807 | -0.1868 | No | ||
| 151 | KCNIP2 | ENSG00000120049.19 | 36062 | -0.851 | -0.1921 | No | ||
| 152 | GP9 | ENSG00000169704.4 | 36202 | -0.868 | -0.1918 | No | ||
| 153 | CA2 | ENSG00000104267.10 | 36626 | -0.919 | -0.1979 | No | ||
| 154 | MT3 | ENSG00000087250.8 | 36936 | -0.957 | -0.2012 | No | ||
| 155 | PREP | ENSG00000085377.14 | 37237 | -0.997 | -0.2042 | No | ||
| 156 | IL6 | ENSG00000136244.12 | 37366 | -1.015 | -0.2030 | No | ||
| 157 | PIK3CG | ENSG00000105851.11 | 37680 | -1.063 | -0.2060 | No | ||
| 158 | EHD1 | ENSG00000110047.18 | 37992 | -1.116 | -0.2087 | No | ||
| 159 | ZEB1 | ENSG00000148516.21 | 38269 | -1.166 | -0.2103 | No | ||
| 160 | SERPINE1 | ENSG00000106366.8 | 38403 | -1.189 | -0.2086 | No | ||
| 161 | MSRB1 | ENSG00000198736.11 | 38437 | -1.196 | -0.2045 | No | ||
| 162 | HPCAL4 | ENSG00000116983.13 | 38464 | -1.199 | -0.2002 | No | ||
| 163 | PCLO | ENSG00000186472.20 | 38753 | -1.255 | -0.2018 | No | ||
| 164 | GRB2 | ENSG00000177885.15 | 39308 | -1.380 | -0.2091 | No | ||
| 165 | ME1 | ENSG00000065833.9 | 39319 | -1.382 | -0.2037 | No | ||
| 166 | XPNPEP1 | ENSG00000108039.18 | 39332 | -1.384 | -0.1983 | No | ||
| 167 | TNFAIP3 | ENSG00000118503.15 | 39359 | -1.390 | -0.1933 | No | ||
| 168 | COL4A2 | ENSG00000134871.19 | 39460 | -1.413 | -0.1898 | No | ||
| 169 | CASP9 | ENSG00000132906.18 | 39565 | -1.434 | -0.1864 | No | ||
| 170 | USP14 | ENSG00000101557.15 | 39871 | -1.514 | -0.1873 | No | ||
| 171 | PFN1 | ENSG00000108518.7 | 40055 | -1.560 | -0.1852 | No | ||
| 172 | CTSH | ENSG00000103811.16 | 40296 | -1.626 | -0.1842 | No | ||
| 173 | CSRP1 | ENSG00000159176.14 | 40304 | -1.628 | -0.1777 | No | ||
| 174 | LTF | ENSG00000012223.13 | 40415 | -1.661 | -0.1735 | No | ||
| 175 | KIF2A | ENSG00000068796.16 | 40453 | -1.672 | -0.1676 | No | ||
| 176 | ADRA2B | ENSG00000274286.2 | 40474 | -1.681 | -0.1612 | No | ||
| 177 | SERPINA1 | ENSG00000197249.13 | 40558 | -1.704 | -0.1562 | No | ||
| 178 | ITGAM | ENSG00000169896.18 | 40880 | -1.815 | -0.1562 | No | ||
| 179 | CTSV | ENSG00000136943.11 | 41043 | -1.887 | -0.1523 | No | ||
| 180 | ZFPM2 | ENSG00000169946.14 | 41138 | -1.926 | -0.1467 | No | ||
| 181 | CDH13 | ENSG00000140945.17 | 41157 | -1.936 | -0.1392 | No | ||
| 182 | PLAUR | ENSG00000011422.12 | 41170 | -1.945 | -0.1315 | No | ||
| 183 | MMP15 | ENSG00000102996.5 | 41275 | -1.992 | -0.1258 | No | ||
| 184 | MMP13 | ENSG00000137745.12 | 41348 | -2.033 | -0.1192 | No | ||
| 185 | DYRK2 | ENSG00000127334.10 | 41627 | -2.208 | -0.1167 | No | ||
| 186 | PLAT | ENSG00000104368.18 | 41640 | -2.216 | -0.1079 | No | ||
| 187 | HSPA5 | ENSG00000044574.8 | 41668 | -2.232 | -0.0995 | No | ||
| 188 | F2 | ENSG00000180210.14 | 41777 | -2.308 | -0.0926 | No | ||
| 189 | PDGFB | ENSG00000100311.17 | 41932 | -2.434 | -0.0862 | No | ||
| 190 | GNGT2 | ENSG00000167083.7 | 42221 | -2.737 | -0.0818 | No | ||
| 191 | FCER1G | ENSG00000158869.11 | 42299 | -2.847 | -0.0720 | No | ||
| 192 | CASP3 | ENSG00000164305.19 | 42313 | -2.869 | -0.0606 | No | ||
| 193 | CALM1 | ENSG00000198668.12 | 42323 | -2.877 | -0.0491 | No | ||
| 194 | DOCK9 | ENSG00000088387.19 | 42425 | -3.027 | -0.0391 | No | ||
| 195 | CDK5R1 | ENSG00000176749.9 | 42942 | -4.486 | -0.0328 | No | ||
| 196 | DGKG | ENSG00000058866.15 | 42945 | -4.495 | -0.0145 | No | ||
| 197 | ACTN2 | ENSG00000077522.13 | 42966 | -4.553 | 0.0036 | No |