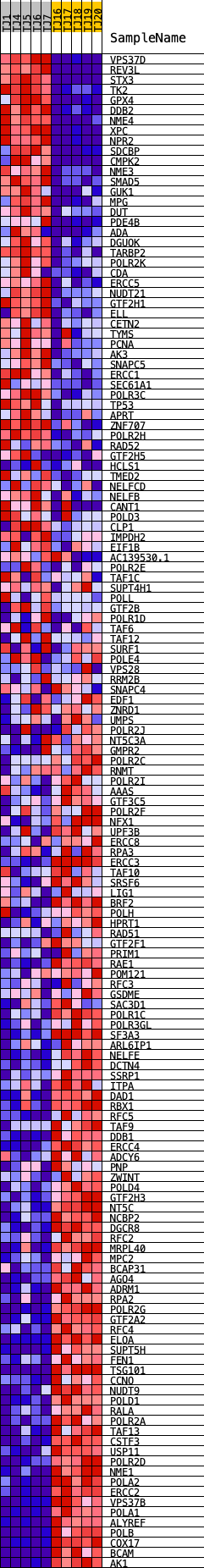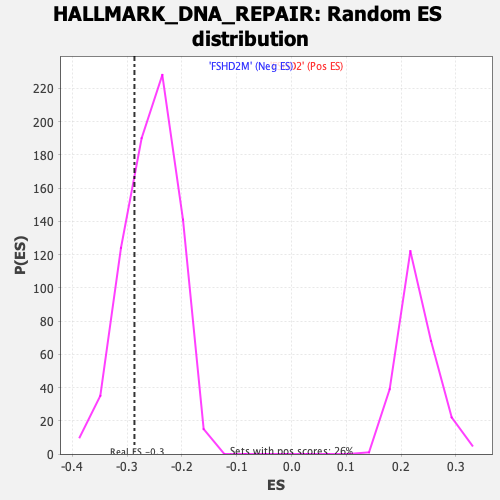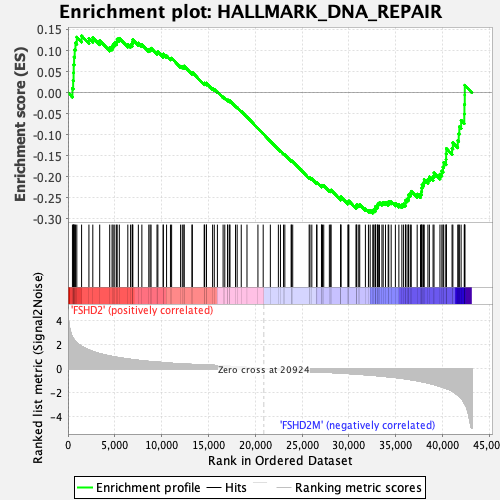
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_DNA_REPAIR |
| Enrichment Score (ES) | -0.2867701 |
| Normalized Enrichment Score (NES) | -1.1164588 |
| Nominal p-value | 0.25975773 |
| FDR q-value | 0.2660153 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VPS37D | ENSG00000176428.6 | 473 | 2.643 | 0.0100 | No | ||
| 2 | REV3L | ENSG00000009413.15 | 550 | 2.543 | 0.0285 | No | ||
| 3 | STX3 | ENSG00000166900.17 | 596 | 2.475 | 0.0471 | No | ||
| 4 | TK2 | ENSG00000166548.16 | 609 | 2.459 | 0.0664 | No | ||
| 5 | GPX4 | ENSG00000167468.17 | 652 | 2.411 | 0.0846 | No | ||
| 6 | DDB2 | ENSG00000134574.11 | 714 | 2.356 | 0.1020 | No | ||
| 7 | NME4 | ENSG00000103202.13 | 795 | 2.266 | 0.1181 | No | ||
| 8 | XPC | ENSG00000154767.14 | 941 | 2.136 | 0.1318 | No | ||
| 9 | NPR2 | ENSG00000159899.14 | 1451 | 1.865 | 0.1348 | No | ||
| 10 | SDCBP | ENSG00000137575.12 | 2234 | 1.554 | 0.1289 | No | ||
| 11 | CMPK2 | ENSG00000134326.11 | 2649 | 1.426 | 0.1306 | No | ||
| 12 | NME3 | ENSG00000103024.7 | 3385 | 1.247 | 0.1235 | No | ||
| 13 | SMAD5 | ENSG00000113658.18 | 4453 | 1.052 | 0.1070 | No | ||
| 14 | GUK1 | ENSG00000143774.16 | 4705 | 1.013 | 0.1092 | No | ||
| 15 | MPG | ENSG00000103152.11 | 4822 | 0.994 | 0.1144 | No | ||
| 16 | DUT | ENSG00000128951.14 | 4996 | 0.969 | 0.1181 | No | ||
| 17 | PDE4B | ENSG00000184588.18 | 5193 | 0.941 | 0.1210 | No | ||
| 18 | ADA | ENSG00000196839.13 | 5246 | 0.934 | 0.1272 | No | ||
| 19 | DGUOK | ENSG00000114956.20 | 5480 | 0.902 | 0.1290 | No | ||
| 20 | TARBP2 | ENSG00000139546.11 | 6387 | 0.799 | 0.1143 | No | ||
| 21 | POLR2K | ENSG00000147669.11 | 6681 | 0.769 | 0.1136 | No | ||
| 22 | CDA | ENSG00000158825.6 | 6850 | 0.752 | 0.1157 | No | ||
| 23 | ERCC5 | ENSG00000134899.21 | 6886 | 0.749 | 0.1208 | No | ||
| 24 | NUDT21 | ENSG00000167005.14 | 6924 | 0.744 | 0.1259 | No | ||
| 25 | GTF2H1 | ENSG00000110768.12 | 7513 | 0.689 | 0.1177 | No | ||
| 26 | ELL | ENSG00000105656.13 | 7887 | 0.656 | 0.1142 | No | ||
| 27 | CETN2 | ENSG00000147400.9 | 8628 | 0.597 | 0.1017 | No | ||
| 28 | TYMS | ENSG00000176890.16 | 8763 | 0.586 | 0.1033 | No | ||
| 29 | PCNA | ENSG00000132646.11 | 8869 | 0.578 | 0.1054 | No | ||
| 30 | AK3 | ENSG00000147853.17 | 9505 | 0.537 | 0.0949 | No | ||
| 31 | SNAPC5 | ENSG00000174446.13 | 9597 | 0.530 | 0.0970 | No | ||
| 32 | ERCC1 | ENSG00000012061.15 | 10169 | 0.499 | 0.0877 | No | ||
| 33 | SEC61A1 | ENSG00000058262.10 | 10170 | 0.499 | 0.0917 | No | ||
| 34 | POLR3C | ENSG00000186141.9 | 10525 | 0.476 | 0.0872 | No | ||
| 35 | TP53 | ENSG00000141510.17 | 10940 | 0.449 | 0.0812 | No | ||
| 36 | APRT | ENSG00000198931.11 | 11064 | 0.440 | 0.0818 | No | ||
| 37 | ZNF707 | ENSG00000181135.16 | 12022 | 0.387 | 0.0626 | No | ||
| 38 | POLR2H | ENSG00000163882.9 | 12205 | 0.377 | 0.0614 | No | ||
| 39 | RAD52 | ENSG00000002016.18 | 12389 | 0.370 | 0.0601 | No | ||
| 40 | GTF2H5 | ENSG00000272047.2 | 12391 | 0.370 | 0.0630 | No | ||
| 41 | HCLS1 | ENSG00000180353.11 | 13234 | 0.344 | 0.0461 | No | ||
| 42 | TMED2 | ENSG00000086598.11 | 13277 | 0.341 | 0.0479 | No | ||
| 43 | NELFCD | ENSG00000101158.15 | 14542 | 0.308 | 0.0209 | No | ||
| 44 | NELFB | ENSG00000188986.7 | 14578 | 0.305 | 0.0225 | No | ||
| 45 | CANT1 | ENSG00000171302.17 | 14775 | 0.293 | 0.0203 | No | ||
| 46 | POLD3 | ENSG00000077514.9 | 14781 | 0.293 | 0.0225 | No | ||
| 47 | CLP1 | ENSG00000172409.6 | 15441 | 0.279 | 0.0094 | No | ||
| 48 | IMPDH2 | ENSG00000178035.12 | 15620 | 0.269 | 0.0074 | No | ||
| 49 | EIF1B | ENSG00000114784.4 | 15953 | 0.252 | 0.0016 | No | ||
| 50 | AC139530.1 | ENSG00000262049.1 | 16561 | 0.216 | -0.0108 | No | ||
| 51 | POLR2E | ENSG00000099817.12 | 16742 | 0.208 | -0.0133 | No | ||
| 52 | TAF1C | ENSG00000103168.16 | 17033 | 0.196 | -0.0185 | No | ||
| 53 | SUPT4H1 | ENSG00000213246.7 | 17067 | 0.194 | -0.0177 | No | ||
| 54 | POLL | ENSG00000166169.17 | 17266 | 0.185 | -0.0208 | No | ||
| 55 | GTF2B | ENSG00000137947.12 | 17284 | 0.184 | -0.0198 | No | ||
| 56 | POLR1D | ENSG00000186184.18 | 17889 | 0.152 | -0.0326 | No | ||
| 57 | TAF6 | ENSG00000106290.15 | 18064 | 0.143 | -0.0355 | No | ||
| 58 | TAF12 | ENSG00000120656.11 | 18500 | 0.125 | -0.0447 | No | ||
| 59 | SURF1 | ENSG00000148290.10 | 19112 | 0.094 | -0.0581 | No | ||
| 60 | POLE4 | ENSG00000115350.12 | 20281 | 0.033 | -0.0850 | No | ||
| 61 | VPS28 | ENSG00000160948.14 | 20843 | 0.005 | -0.0981 | No | ||
| 62 | RRM2B | ENSG00000048392.11 | 21607 | -0.038 | -0.1155 | No | ||
| 63 | SNAPC4 | ENSG00000165684.4 | 22464 | -0.083 | -0.1348 | No | ||
| 64 | EDF1 | ENSG00000107223.13 | 22686 | -0.096 | -0.1392 | No | ||
| 65 | ZNRD1 | ENSG00000066379.15 | 23017 | -0.113 | -0.1459 | No | ||
| 66 | UMPS | ENSG00000114491.14 | 23154 | -0.122 | -0.1481 | No | ||
| 67 | POLR2J | ENSG00000005075.15 | 23836 | -0.160 | -0.1627 | No | ||
| 68 | NT5C3A | ENSG00000122643.21 | 23852 | -0.162 | -0.1618 | No | ||
| 69 | GMPR2 | ENSG00000100938.17 | 23995 | -0.171 | -0.1637 | No | ||
| 70 | POLR2C | ENSG00000102978.13 | 25751 | -0.243 | -0.2026 | No | ||
| 71 | RNMT | ENSG00000101654.17 | 25839 | -0.250 | -0.2027 | No | ||
| 72 | POLR2I | ENSG00000105258.9 | 26035 | -0.262 | -0.2051 | No | ||
| 73 | AAAS | ENSG00000094914.14 | 26543 | -0.269 | -0.2148 | No | ||
| 74 | GTF3C5 | ENSG00000148308.17 | 26585 | -0.273 | -0.2135 | No | ||
| 75 | POLR2F | ENSG00000100142.15 | 27064 | -0.278 | -0.2225 | No | ||
| 76 | NFX1 | ENSG00000086102.19 | 27077 | -0.279 | -0.2205 | No | ||
| 77 | UPF3B | ENSG00000125351.11 | 27177 | -0.286 | -0.2206 | No | ||
| 78 | ERCC8 | ENSG00000049167.14 | 27308 | -0.296 | -0.2212 | No | ||
| 79 | RPA3 | ENSG00000106399.11 | 27907 | -0.302 | -0.2327 | No | ||
| 80 | ERCC3 | ENSG00000163161.13 | 28052 | -0.314 | -0.2336 | No | ||
| 81 | TAF10 | ENSG00000166337.10 | 28088 | -0.316 | -0.2319 | No | ||
| 82 | SRSF6 | ENSG00000124193.16 | 29117 | -0.354 | -0.2530 | No | ||
| 83 | LIG1 | ENSG00000105486.14 | 29118 | -0.354 | -0.2502 | No | ||
| 84 | BRF2 | ENSG00000104221.13 | 29144 | -0.357 | -0.2479 | No | ||
| 85 | POLH | ENSG00000170734.11 | 29886 | -0.397 | -0.2620 | No | ||
| 86 | HPRT1 | ENSG00000165704.15 | 29958 | -0.404 | -0.2604 | No | ||
| 87 | RAD51 | ENSG00000051180.17 | 29987 | -0.406 | -0.2579 | No | ||
| 88 | GTF2F1 | ENSG00000125651.14 | 30762 | -0.451 | -0.2723 | No | ||
| 89 | PRIM1 | ENSG00000198056.15 | 30787 | -0.453 | -0.2692 | No | ||
| 90 | RAE1 | ENSG00000101146.13 | 30841 | -0.456 | -0.2668 | No | ||
| 91 | POM121 | ENSG00000196313.11 | 31048 | -0.463 | -0.2679 | No | ||
| 92 | RFC3 | ENSG00000133119.13 | 31151 | -0.472 | -0.2666 | No | ||
| 93 | GSDME | ENSG00000105928.16 | 31756 | -0.501 | -0.2766 | No | ||
| 94 | SAC3D1 | ENSG00000168061.15 | 32081 | -0.520 | -0.2800 | No | ||
| 95 | POLR1C | ENSG00000171453.20 | 32265 | -0.532 | -0.2801 | No | ||
| 96 | POLR3GL | ENSG00000121851.13 | 32554 | -0.547 | -0.2824 | Yes | ||
| 97 | SF3A3 | ENSG00000183431.12 | 32599 | -0.551 | -0.2791 | Yes | ||
| 98 | ARL6IP1 | ENSG00000170540.15 | 32793 | -0.568 | -0.2790 | Yes | ||
| 99 | NELFE | ENSG00000204356.14 | 32806 | -0.569 | -0.2748 | Yes | ||
| 100 | DCTN4 | ENSG00000132912.12 | 32834 | -0.571 | -0.2709 | Yes | ||
| 101 | SSRP1 | ENSG00000149136.9 | 33026 | -0.585 | -0.2706 | Yes | ||
| 102 | ITPA | ENSG00000125877.13 | 33045 | -0.587 | -0.2664 | Yes | ||
| 103 | DAD1 | ENSG00000129562.11 | 33139 | -0.593 | -0.2638 | Yes | ||
| 104 | RBX1 | ENSG00000100387.9 | 33259 | -0.602 | -0.2618 | Yes | ||
| 105 | RFC5 | ENSG00000111445.14 | 33522 | -0.623 | -0.2630 | Yes | ||
| 106 | TAF9 | ENSG00000273841.5 | 33655 | -0.635 | -0.2610 | Yes | ||
| 107 | DDB1 | ENSG00000167986.13 | 33905 | -0.654 | -0.2616 | Yes | ||
| 108 | ERCC4 | ENSG00000175595.14 | 34189 | -0.675 | -0.2628 | Yes | ||
| 109 | ADCY6 | ENSG00000174233.11 | 34252 | -0.679 | -0.2588 | Yes | ||
| 110 | PNP | ENSG00000198805.11 | 34508 | -0.703 | -0.2592 | Yes | ||
| 111 | ZWINT | ENSG00000122952.17 | 34958 | -0.738 | -0.2637 | Yes | ||
| 112 | POLD4 | ENSG00000175482.9 | 35331 | -0.771 | -0.2663 | Yes | ||
| 113 | GTF2H3 | ENSG00000111358.14 | 35650 | -0.804 | -0.2673 | Yes | ||
| 114 | NT5C | ENSG00000125458.7 | 35836 | -0.822 | -0.2650 | Yes | ||
| 115 | NCBP2 | ENSG00000114503.11 | 36040 | -0.848 | -0.2630 | Yes | ||
| 116 | DGCR8 | ENSG00000128191.16 | 36043 | -0.849 | -0.2563 | Yes | ||
| 117 | RFC2 | ENSG00000049541.11 | 36206 | -0.868 | -0.2532 | Yes | ||
| 118 | MRPL40 | ENSG00000185608.8 | 36363 | -0.884 | -0.2498 | Yes | ||
| 119 | MPC2 | ENSG00000143158.11 | 36372 | -0.885 | -0.2429 | Yes | ||
| 120 | BCAP31 | ENSG00000185825.17 | 36556 | -0.907 | -0.2399 | Yes | ||
| 121 | AGO4 | ENSG00000134698.11 | 36650 | -0.922 | -0.2348 | Yes | ||
| 122 | ADRM1 | ENSG00000130706.13 | 37285 | -1.005 | -0.2415 | Yes | ||
| 123 | RPA2 | ENSG00000117748.10 | 37640 | -1.058 | -0.2414 | Yes | ||
| 124 | POLR2G | ENSG00000168002.12 | 37749 | -1.077 | -0.2353 | Yes | ||
| 125 | GTF2A2 | ENSG00000140307.11 | 37772 | -1.079 | -0.2272 | Yes | ||
| 126 | RFC4 | ENSG00000163918.10 | 37809 | -1.085 | -0.2194 | Yes | ||
| 127 | ELOA | ENSG00000011007.12 | 37965 | -1.112 | -0.2142 | Yes | ||
| 128 | SUPT5H | ENSG00000196235.14 | 38036 | -1.123 | -0.2069 | Yes | ||
| 129 | FEN1 | ENSG00000168496.4 | 38431 | -1.194 | -0.2066 | Yes | ||
| 130 | TSG101 | ENSG00000074319.13 | 38589 | -1.221 | -0.2005 | Yes | ||
| 131 | CCNO | ENSG00000152669.9 | 38981 | -1.305 | -0.1992 | Yes | ||
| 132 | NUDT9 | ENSG00000170502.13 | 39083 | -1.327 | -0.1910 | Yes | ||
| 133 | POLD1 | ENSG00000062822.14 | 39718 | -1.471 | -0.1941 | Yes | ||
| 134 | RALA | ENSG00000006451.8 | 39921 | -1.528 | -0.1866 | Yes | ||
| 135 | POLR2A | ENSG00000181222.16 | 40044 | -1.559 | -0.1770 | Yes | ||
| 136 | TAF13 | ENSG00000197780.10 | 40119 | -1.578 | -0.1662 | Yes | ||
| 137 | CSTF3 | ENSG00000176102.13 | 40367 | -1.645 | -0.1589 | Yes | ||
| 138 | USP11 | ENSG00000102226.9 | 40376 | -1.650 | -0.1459 | Yes | ||
| 139 | POLR2D | ENSG00000144231.11 | 40398 | -1.657 | -0.1332 | Yes | ||
| 140 | NME1 | ENSG00000239672.7 | 41010 | -1.873 | -0.1325 | Yes | ||
| 141 | POLA2 | ENSG00000014138.9 | 41096 | -1.906 | -0.1193 | Yes | ||
| 142 | ERCC2 | ENSG00000104884.15 | 41619 | -2.203 | -0.1140 | Yes | ||
| 143 | VPS37B | ENSG00000139722.7 | 41722 | -2.271 | -0.0983 | Yes | ||
| 144 | POLA1 | ENSG00000101868.11 | 41785 | -2.314 | -0.0813 | Yes | ||
| 145 | ALYREF | ENSG00000183684.8 | 41977 | -2.473 | -0.0661 | Yes | ||
| 146 | POLB | ENSG00000070501.12 | 42316 | -2.869 | -0.0511 | Yes | ||
| 147 | COX17 | ENSG00000138495.6 | 42332 | -2.886 | -0.0285 | Yes | ||
| 148 | BCAM | ENSG00000187244.11 | 42367 | -2.933 | -0.0059 | Yes | ||
| 149 | AK1 | ENSG00000106992.19 | 42370 | -2.941 | 0.0174 | Yes |