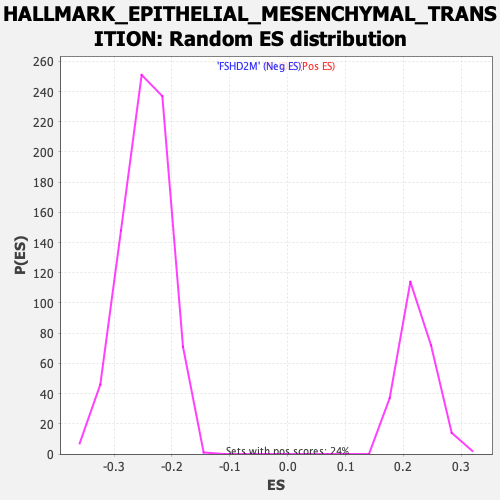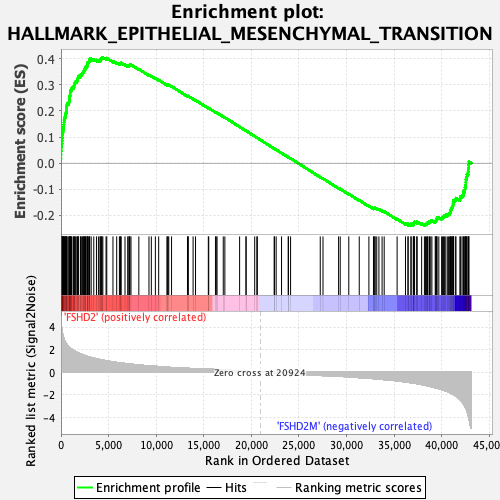
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION |
| Enrichment Score (ES) | 0.4048666 |
| Normalized Enrichment Score (NES) | 1.8267726 |
| Nominal p-value | 0.0 |
| FDR q-value | 5.192308E-4 |
| FWER p-Value | 0.002 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLIT3 | ENSG00000184347.14 | 0 | 4.923 | 0.0165 | Yes | ||
| 2 | DCN | ENSG00000011465.17 | 6 | 4.734 | 0.0322 | Yes | ||
| 3 | FBLN5 | ENSG00000140092.14 | 11 | 4.579 | 0.0475 | Yes | ||
| 4 | PCOLCE | ENSG00000106333.13 | 33 | 4.267 | 0.0613 | Yes | ||
| 5 | FBLN1 | ENSG00000077942.19 | 63 | 3.993 | 0.0740 | Yes | ||
| 6 | PDGFRB | ENSG00000113721.14 | 106 | 3.655 | 0.0853 | Yes | ||
| 7 | CTHRC1 | ENSG00000164932.13 | 123 | 3.593 | 0.0970 | Yes | ||
| 8 | IGFBP4 | ENSG00000141753.7 | 151 | 3.478 | 0.1080 | Yes | ||
| 9 | CD44 | ENSG00000026508.18 | 157 | 3.446 | 0.1194 | Yes | ||
| 10 | FMOD | ENSG00000122176.12 | 215 | 3.256 | 0.1290 | Yes | ||
| 11 | ANPEP | ENSG00000166825.14 | 265 | 3.089 | 0.1382 | Yes | ||
| 12 | PCOLCE2 | ENSG00000163710.9 | 294 | 3.004 | 0.1476 | Yes | ||
| 13 | PMP22 | ENSG00000109099.15 | 310 | 2.946 | 0.1571 | Yes | ||
| 14 | ITGA2 | ENSG00000164171.11 | 327 | 2.899 | 0.1665 | Yes | ||
| 15 | COL6A3 | ENSG00000163359.15 | 371 | 2.812 | 0.1749 | Yes | ||
| 16 | SLIT2 | ENSG00000145147.20 | 459 | 2.664 | 0.1818 | Yes | ||
| 17 | TGFBR3 | ENSG00000069702.11 | 474 | 2.641 | 0.1903 | Yes | ||
| 18 | SERPINE2 | ENSG00000135919.13 | 544 | 2.548 | 0.1973 | Yes | ||
| 19 | MATN2 | ENSG00000132561.14 | 579 | 2.497 | 0.2048 | Yes | ||
| 20 | TIMP1 | ENSG00000102265.12 | 582 | 2.491 | 0.2131 | Yes | ||
| 21 | COL16A1 | ENSG00000084636.18 | 589 | 2.483 | 0.2213 | Yes | ||
| 22 | WIPF1 | ENSG00000115935.17 | 654 | 2.409 | 0.2279 | Yes | ||
| 23 | COL6A2 | ENSG00000142173.15 | 816 | 2.246 | 0.2317 | Yes | ||
| 24 | VIM | ENSG00000026025.16 | 852 | 2.214 | 0.2383 | Yes | ||
| 25 | SDC4 | ENSG00000124145.6 | 886 | 2.182 | 0.2448 | Yes | ||
| 26 | TIMP3 | ENSG00000100234.11 | 892 | 2.178 | 0.2520 | Yes | ||
| 27 | FAS | ENSG00000026103.22 | 911 | 2.164 | 0.2588 | Yes | ||
| 28 | MSX1 | ENSG00000163132.8 | 976 | 2.117 | 0.2644 | Yes | ||
| 29 | FBLN2 | ENSG00000163520.14 | 977 | 2.116 | 0.2715 | Yes | ||
| 30 | CAPG | ENSG00000042493.16 | 994 | 2.108 | 0.2782 | Yes | ||
| 31 | ECM1 | ENSG00000143369.15 | 1104 | 2.046 | 0.2825 | Yes | ||
| 32 | PRRX1 | ENSG00000116132.12 | 1144 | 2.020 | 0.2884 | Yes | ||
| 33 | DKK1 | ENSG00000107984.10 | 1299 | 1.941 | 0.2913 | Yes | ||
| 34 | TNFRSF11B | ENSG00000164761.9 | 1360 | 1.901 | 0.2963 | Yes | ||
| 35 | LUM | ENSG00000139329.5 | 1391 | 1.886 | 0.3019 | Yes | ||
| 36 | THBS2 | ENSG00000186340.15 | 1447 | 1.867 | 0.3069 | Yes | ||
| 37 | LOX | ENSG00000113083.14 | 1511 | 1.838 | 0.3116 | Yes | ||
| 38 | CD59 | ENSG00000085063.16 | 1662 | 1.770 | 0.3140 | Yes | ||
| 39 | PPIB | ENSG00000166794.4 | 1719 | 1.748 | 0.3186 | Yes | ||
| 40 | GEM | ENSG00000164949.8 | 1767 | 1.729 | 0.3233 | Yes | ||
| 41 | CXCL12 | ENSG00000107562.16 | 1796 | 1.720 | 0.3284 | Yes | ||
| 42 | IGFBP3 | ENSG00000146674.15 | 1828 | 1.702 | 0.3334 | Yes | ||
| 43 | NT5E | ENSG00000135318.12 | 2035 | 1.617 | 0.3340 | Yes | ||
| 44 | LAMA1 | ENSG00000101680.15 | 2061 | 1.609 | 0.3388 | Yes | ||
| 45 | SFRP1 | ENSG00000104332.12 | 2175 | 1.573 | 0.3414 | Yes | ||
| 46 | THY1 | ENSG00000154096.13 | 2244 | 1.552 | 0.3450 | Yes | ||
| 47 | DPYSL3 | ENSG00000113657.13 | 2327 | 1.526 | 0.3483 | Yes | ||
| 48 | GJA1 | ENSG00000152661.9 | 2383 | 1.509 | 0.3520 | Yes | ||
| 49 | MXRA5 | ENSG00000101825.8 | 2436 | 1.494 | 0.3558 | Yes | ||
| 50 | MMP1 | ENSG00000196611.5 | 2473 | 1.481 | 0.3600 | Yes | ||
| 51 | MMP3 | ENSG00000149968.12 | 2562 | 1.453 | 0.3628 | Yes | ||
| 52 | FUCA1 | ENSG00000179163.11 | 2594 | 1.443 | 0.3669 | Yes | ||
| 53 | MMP2 | ENSG00000087245.13 | 2616 | 1.437 | 0.3712 | Yes | ||
| 54 | PTHLH | ENSG00000087494.16 | 2751 | 1.393 | 0.3728 | Yes | ||
| 55 | CDH11 | ENSG00000140937.14 | 2757 | 1.391 | 0.3773 | Yes | ||
| 56 | IGFBP2 | ENSG00000115457.10 | 2774 | 1.385 | 0.3816 | Yes | ||
| 57 | PTX3 | ENSG00000163661.4 | 2789 | 1.381 | 0.3859 | Yes | ||
| 58 | EMP3 | ENSG00000142227.11 | 2904 | 1.355 | 0.3878 | Yes | ||
| 59 | EFEMP2 | ENSG00000172638.13 | 2973 | 1.339 | 0.3907 | Yes | ||
| 60 | LRP1 | ENSG00000123384.14 | 3000 | 1.333 | 0.3945 | Yes | ||
| 61 | GAS1 | ENSG00000180447.7 | 3007 | 1.331 | 0.3988 | Yes | ||
| 62 | ECM2 | ENSG00000106823.12 | 3170 | 1.296 | 0.3994 | Yes | ||
| 63 | FAP | ENSG00000078098.14 | 3443 | 1.238 | 0.3972 | Yes | ||
| 64 | MGP | ENSG00000111341.10 | 3725 | 1.178 | 0.3946 | Yes | ||
| 65 | VEGFC | ENSG00000150630.4 | 3931 | 1.140 | 0.3937 | Yes | ||
| 66 | DAB2 | ENSG00000153071.15 | 4095 | 1.112 | 0.3936 | Yes | ||
| 67 | THBS1 | ENSG00000137801.11 | 4127 | 1.107 | 0.3966 | Yes | ||
| 68 | GREM1 | ENSG00000166923.12 | 4206 | 1.092 | 0.3984 | Yes | ||
| 69 | LGALS1 | ENSG00000100097.12 | 4241 | 1.085 | 0.4013 | Yes | ||
| 70 | ITGB5 | ENSG00000082781.12 | 4315 | 1.074 | 0.4032 | Yes | ||
| 71 | TNC | ENSG00000041982.16 | 4396 | 1.062 | 0.4049 | Yes | ||
| 72 | FGF2 | ENSG00000138685.15 | 4734 | 1.009 | 0.4004 | No | ||
| 73 | SNAI2 | ENSG00000019549.13 | 4808 | 0.996 | 0.4020 | No | ||
| 74 | CXCL6 | ENSG00000124875.10 | 5459 | 0.905 | 0.3899 | No | ||
| 75 | FBN1 | ENSG00000166147.13 | 5850 | 0.858 | 0.3837 | No | ||
| 76 | IL15 | ENSG00000164136.17 | 6136 | 0.824 | 0.3798 | No | ||
| 77 | COMP | ENSG00000105664.11 | 6225 | 0.814 | 0.3805 | No | ||
| 78 | CADM1 | ENSG00000182985.17 | 6243 | 0.812 | 0.3828 | No | ||
| 79 | COL1A2 | ENSG00000164692.17 | 6324 | 0.804 | 0.3837 | No | ||
| 80 | COL12A1 | ENSG00000111799.21 | 6723 | 0.765 | 0.3770 | No | ||
| 81 | P3H1 | ENSG00000117385.15 | 7005 | 0.738 | 0.3729 | No | ||
| 82 | BGN | ENSG00000182492.16 | 7136 | 0.724 | 0.3723 | No | ||
| 83 | TFPI2 | ENSG00000105825.14 | 7157 | 0.723 | 0.3742 | No | ||
| 84 | SPOCK1 | ENSG00000152377.14 | 7208 | 0.718 | 0.3755 | No | ||
| 85 | COL3A1 | ENSG00000168542.14 | 7229 | 0.715 | 0.3774 | No | ||
| 86 | LRRC15 | ENSG00000172061.8 | 7351 | 0.706 | 0.3770 | No | ||
| 87 | QSOX1 | ENSG00000116260.17 | 8176 | 0.631 | 0.3599 | No | ||
| 88 | LOXL1 | ENSG00000129038.16 | 9240 | 0.552 | 0.3370 | No | ||
| 89 | MMP14 | ENSG00000157227.13 | 9489 | 0.539 | 0.3330 | No | ||
| 90 | FN1 | ENSG00000115414.19 | 9913 | 0.516 | 0.3249 | No | ||
| 91 | PLOD3 | ENSG00000106397.12 | 10261 | 0.493 | 0.3184 | No | ||
| 92 | TPM4 | ENSG00000167460.17 | 11125 | 0.437 | 0.2998 | No | ||
| 93 | FBN2 | ENSG00000138829.12 | 11219 | 0.432 | 0.2991 | No | ||
| 94 | COPA | ENSG00000122218.16 | 11223 | 0.431 | 0.3004 | No | ||
| 95 | GLIPR1 | ENSG00000139278.10 | 11320 | 0.426 | 0.2996 | No | ||
| 96 | FSTL1 | ENSG00000163430.12 | 11613 | 0.407 | 0.2942 | No | ||
| 97 | SDC1 | ENSG00000115884.11 | 13286 | 0.340 | 0.2564 | No | ||
| 98 | BDNF | ENSG00000176697.19 | 13340 | 0.337 | 0.2563 | No | ||
| 99 | BASP1 | ENSG00000176788.9 | 13365 | 0.335 | 0.2568 | No | ||
| 100 | PRSS2 | ENSG00000275896.5 | 13875 | 0.322 | 0.2461 | No | ||
| 101 | TGM2 | ENSG00000198959.12 | 14132 | 0.309 | 0.2411 | No | ||
| 102 | PLOD1 | ENSG00000083444.17 | 15475 | 0.276 | 0.2108 | No | ||
| 103 | SAT1 | ENSG00000130066.16 | 15520 | 0.274 | 0.2107 | No | ||
| 104 | MAGEE1 | ENSG00000198934.5 | 16225 | 0.235 | 0.1951 | No | ||
| 105 | APLP1 | ENSG00000105290.12 | 16304 | 0.230 | 0.1940 | No | ||
| 106 | CDH6 | ENSG00000113361.13 | 16393 | 0.226 | 0.1927 | No | ||
| 107 | CXCL1 | ENSG00000163739.5 | 17045 | 0.196 | 0.1782 | No | ||
| 108 | WNT5A | ENSG00000114251.14 | 17212 | 0.187 | 0.1750 | No | ||
| 109 | RGS4 | ENSG00000117152.13 | 18763 | 0.109 | 0.1392 | No | ||
| 110 | MYLK | ENSG00000065534.18 | 19434 | 0.074 | 0.1239 | No | ||
| 111 | ID2 | ENSG00000115738.10 | 19442 | 0.074 | 0.1239 | No | ||
| 112 | COL8A2 | ENSG00000171812.13 | 20347 | 0.029 | 0.1030 | No | ||
| 113 | SPP1 | ENSG00000118785.14 | 20575 | 0.018 | 0.0977 | No | ||
| 114 | MCM7 | ENSG00000166508.17 | 20623 | 0.016 | 0.0967 | No | ||
| 115 | CCN1 | ENSG00000142871.17 | 22390 | -0.079 | 0.0558 | No | ||
| 116 | LAMC1 | ENSG00000135862.6 | 22432 | -0.081 | 0.0551 | No | ||
| 117 | LAMA2 | ENSG00000196569.12 | 22603 | -0.091 | 0.0515 | No | ||
| 118 | GPX7 | ENSG00000116157.6 | 23167 | -0.122 | 0.0388 | No | ||
| 119 | COLGALT1 | ENSG00000130309.11 | 23871 | -0.162 | 0.0229 | No | ||
| 120 | NOTCH2 | ENSG00000134250.20 | 23887 | -0.164 | 0.0231 | No | ||
| 121 | HTRA1 | ENSG00000166033.13 | 24116 | -0.177 | 0.0184 | No | ||
| 122 | FZD8 | ENSG00000177283.7 | 27233 | -0.290 | -0.0532 | No | ||
| 123 | COL5A3 | ENSG00000080573.7 | 27521 | -0.299 | -0.0589 | No | ||
| 124 | AREG | ENSG00000109321.11 | 29159 | -0.357 | -0.0958 | No | ||
| 125 | SERPINH1 | ENSG00000149257.14 | 29331 | -0.364 | -0.0986 | No | ||
| 126 | SCG2 | ENSG00000171951.5 | 30230 | -0.423 | -0.1181 | No | ||
| 127 | ITGB3 | ENSG00000259207.7 | 31330 | -0.481 | -0.1421 | No | ||
| 128 | COL1A1 | ENSG00000108821.13 | 32343 | -0.536 | -0.1639 | No | ||
| 129 | CCN2 | ENSG00000118523.6 | 32823 | -0.570 | -0.1731 | No | ||
| 130 | MEST | ENSG00000106484.16 | 32879 | -0.574 | -0.1725 | No | ||
| 131 | ADAM12 | ENSG00000148848.14 | 32976 | -0.582 | -0.1728 | No | ||
| 132 | LAMA3 | ENSG00000053747.16 | 32988 | -0.582 | -0.1711 | No | ||
| 133 | OXTR | ENSG00000180914.10 | 33167 | -0.595 | -0.1732 | No | ||
| 134 | POSTN | ENSG00000133110.15 | 33395 | -0.612 | -0.1765 | No | ||
| 135 | EDIL3 | ENSG00000164176.13 | 33729 | -0.640 | -0.1821 | No | ||
| 136 | CALD1 | ENSG00000122786.20 | 33950 | -0.658 | -0.1850 | No | ||
| 137 | ABI3BP | ENSG00000154175.17 | 35307 | -0.769 | -0.2140 | No | ||
| 138 | VCAN | ENSG00000038427.16 | 36187 | -0.866 | -0.2316 | No | ||
| 139 | ITGB1 | ENSG00000150093.19 | 36454 | -0.895 | -0.2348 | No | ||
| 140 | FLNA | ENSG00000196924.17 | 36458 | -0.895 | -0.2319 | No | ||
| 141 | COL5A2 | ENSG00000204262.14 | 36764 | -0.936 | -0.2358 | No | ||
| 142 | CALU | ENSG00000128595.17 | 36775 | -0.937 | -0.2329 | No | ||
| 143 | ELN | ENSG00000049540.17 | 37004 | -0.966 | -0.2350 | No | ||
| 144 | SPARC | ENSG00000113140.11 | 37052 | -0.974 | -0.2328 | No | ||
| 145 | NID2 | ENSG00000087303.18 | 37054 | -0.975 | -0.2296 | No | ||
| 146 | COL5A1 | ENSG00000130635.15 | 37069 | -0.977 | -0.2266 | No | ||
| 147 | JUN | ENSG00000177606.7 | 37141 | -0.985 | -0.2250 | No | ||
| 148 | IL6 | ENSG00000136244.12 | 37366 | -1.015 | -0.2268 | No | ||
| 149 | LAMC2 | ENSG00000058085.15 | 37414 | -1.022 | -0.2245 | No | ||
| 150 | CXCL8 | ENSG00000169429.11 | 37884 | -1.098 | -0.2317 | No | ||
| 151 | SGCB | ENSG00000163069.12 | 38176 | -1.149 | -0.2347 | No | ||
| 152 | FOXC2 | ENSG00000176692.8 | 38284 | -1.169 | -0.2332 | No | ||
| 153 | SERPINE1 | ENSG00000106366.8 | 38403 | -1.189 | -0.2320 | No | ||
| 154 | TGFB1 | ENSG00000105329.10 | 38478 | -1.201 | -0.2297 | No | ||
| 155 | RHOB | ENSG00000143878.10 | 38518 | -1.210 | -0.2265 | No | ||
| 156 | MFAP5 | ENSG00000197614.11 | 38668 | -1.237 | -0.2259 | No | ||
| 157 | SLC6A8 | ENSG00000130821.16 | 38709 | -1.245 | -0.2226 | No | ||
| 158 | PFN2 | ENSG00000070087.14 | 38838 | -1.276 | -0.2213 | No | ||
| 159 | LOXL2 | ENSG00000134013.16 | 38961 | -1.300 | -0.2198 | No | ||
| 160 | COL4A1 | ENSG00000187498.16 | 39310 | -1.380 | -0.2233 | No | ||
| 161 | TNFAIP3 | ENSG00000118503.15 | 39359 | -1.390 | -0.2198 | No | ||
| 162 | COL7A1 | ENSG00000114270.17 | 39421 | -1.404 | -0.2165 | No | ||
| 163 | COL4A2 | ENSG00000134871.19 | 39460 | -1.413 | -0.2126 | No | ||
| 164 | BMP1 | ENSG00000168487.19 | 39505 | -1.423 | -0.2089 | No | ||
| 165 | ENO2 | ENSG00000111674.9 | 39689 | -1.464 | -0.2083 | No | ||
| 166 | CRLF1 | ENSG00000006016.11 | 39981 | -1.543 | -0.2099 | No | ||
| 167 | MATN3 | ENSG00000132031.13 | 40083 | -1.566 | -0.2070 | No | ||
| 168 | MYL9 | ENSG00000101335.10 | 40187 | -1.596 | -0.2040 | No | ||
| 169 | COL11A1 | ENSG00000060718.22 | 40275 | -1.618 | -0.2006 | No | ||
| 170 | ITGA5 | ENSG00000161638.11 | 40419 | -1.662 | -0.1984 | No | ||
| 171 | DST | ENSG00000151914.20 | 40581 | -1.710 | -0.1964 | No | ||
| 172 | TGFBI | ENSG00000120708.17 | 40715 | -1.752 | -0.1936 | No | ||
| 173 | VEGFA | ENSG00000112715.22 | 40824 | -1.792 | -0.1901 | No | ||
| 174 | PMEPA1 | ENSG00000124225.16 | 40906 | -1.826 | -0.1859 | No | ||
| 175 | NNMT | ENSG00000166741.7 | 40927 | -1.833 | -0.1802 | No | ||
| 176 | GADD45B | ENSG00000099860.9 | 40977 | -1.852 | -0.1752 | No | ||
| 177 | SGCD | ENSG00000170624.13 | 41042 | -1.886 | -0.1703 | No | ||
| 178 | ITGAV | ENSG00000138448.12 | 41100 | -1.907 | -0.1653 | No | ||
| 179 | PLAUR | ENSG00000011422.12 | 41170 | -1.945 | -0.1604 | No | ||
| 180 | TNFRSF12A | ENSG00000006327.13 | 41188 | -1.954 | -0.1542 | No | ||
| 181 | GADD45A | ENSG00000116717.13 | 41218 | -1.966 | -0.1483 | No | ||
| 182 | VCAM1 | ENSG00000162692.12 | 41228 | -1.969 | -0.1419 | No | ||
| 183 | INHBA | ENSG00000122641.11 | 41421 | -2.082 | -0.1394 | No | ||
| 184 | FERMT2 | ENSG00000073712.15 | 41491 | -2.127 | -0.1339 | No | ||
| 185 | CDH2 | ENSG00000170558.9 | 41908 | -2.414 | -0.1355 | No | ||
| 186 | TAGLN | ENSG00000149591.17 | 41950 | -2.456 | -0.1282 | No | ||
| 187 | NTM | ENSG00000182667.14 | 42122 | -2.597 | -0.1235 | No | ||
| 188 | ACTA2 | ENSG00000107796.13 | 42259 | -2.777 | -0.1174 | No | ||
| 189 | SGCG | ENSG00000102683.7 | 42272 | -2.813 | -0.1082 | No | ||
| 190 | PVR | ENSG00000073008.15 | 42407 | -2.989 | -0.1013 | No | ||
| 191 | IL32 | ENSG00000008517.16 | 42438 | -3.039 | -0.0918 | No | ||
| 192 | PDLIM4 | ENSG00000131435.13 | 42504 | -3.130 | -0.0829 | No | ||
| 193 | PLOD2 | ENSG00000152952.12 | 42521 | -3.156 | -0.0727 | No | ||
| 194 | GPC1 | ENSG00000063660.9 | 42528 | -3.165 | -0.0622 | No | ||
| 195 | SNTB1 | ENSG00000172164.15 | 42613 | -3.372 | -0.0528 | No | ||
| 196 | FSTL3 | ENSG00000070404.10 | 42644 | -3.449 | -0.0420 | No | ||
| 197 | SFRP4 | ENSG00000106483.12 | 42775 | -3.803 | -0.0323 | No | ||
| 198 | TPM1 | ENSG00000140416.21 | 42794 | -3.877 | -0.0197 | No | ||
| 199 | CAP2 | ENSG00000112186.12 | 42820 | -3.986 | -0.0069 | No | ||
| 200 | TPM2 | ENSG00000198467.15 | 42854 | -4.136 | 0.0062 | No |