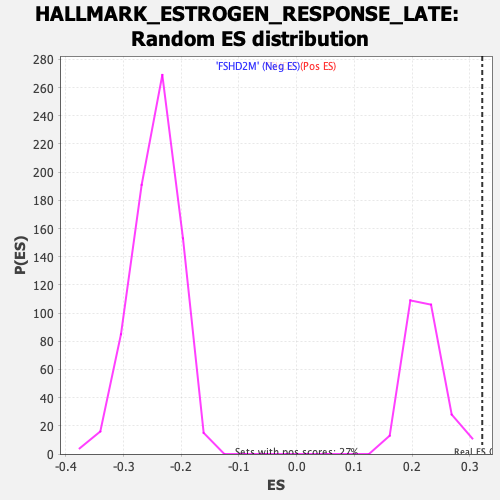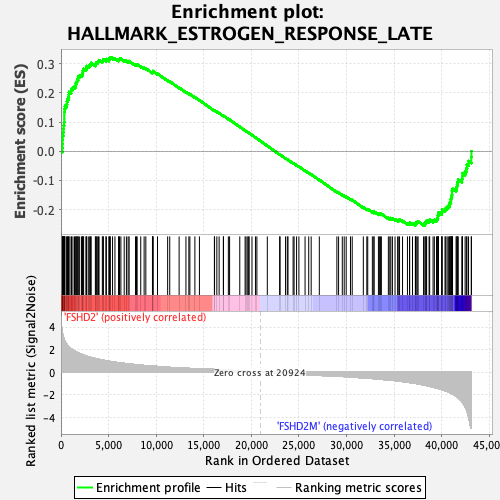
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_ESTROGEN_RESPONSE_LATE |
| Enrichment Score (ES) | 0.3217897 |
| Normalized Enrichment Score (NES) | 1.4546726 |
| Nominal p-value | 0.0037453184 |
| FDR q-value | 0.025801698 |
| FWER p-Value | 0.163 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NBL1 | ENSG00000158747.14 | 142 | 3.516 | 0.0111 | Yes | ||
| 2 | IGFBP4 | ENSG00000141753.7 | 151 | 3.478 | 0.0252 | Yes | ||
| 3 | CD44 | ENSG00000026508.18 | 157 | 3.446 | 0.0392 | Yes | ||
| 4 | SCNN1A | ENSG00000111319.13 | 172 | 3.375 | 0.0526 | Yes | ||
| 5 | FAM102A | ENSG00000167106.12 | 222 | 3.245 | 0.0648 | Yes | ||
| 6 | PGR | ENSG00000082175.15 | 256 | 3.111 | 0.0768 | Yes | ||
| 7 | IL6ST | ENSG00000134352.20 | 282 | 3.044 | 0.0887 | Yes | ||
| 8 | TPBG | ENSG00000146242.8 | 331 | 2.893 | 0.0994 | Yes | ||
| 9 | ZFP36 | ENSG00000128016.7 | 348 | 2.862 | 0.1107 | Yes | ||
| 10 | BCL2 | ENSG00000171791.13 | 349 | 2.862 | 0.1225 | Yes | ||
| 11 | PERP | ENSG00000112378.12 | 350 | 2.857 | 0.1342 | Yes | ||
| 12 | ACOX2 | ENSG00000168306.13 | 370 | 2.812 | 0.1452 | Yes | ||
| 13 | KLF4 | ENSG00000136826.15 | 421 | 2.729 | 0.1552 | Yes | ||
| 14 | ADD3 | ENSG00000148700.15 | 598 | 2.472 | 0.1613 | Yes | ||
| 15 | ATP2B4 | ENSG00000058668.14 | 627 | 2.432 | 0.1706 | Yes | ||
| 16 | CYP26B1 | ENSG00000003137.8 | 683 | 2.377 | 0.1790 | Yes | ||
| 17 | MICB | ENSG00000204516.10 | 794 | 2.266 | 0.1857 | Yes | ||
| 18 | OLFM1 | ENSG00000130558.19 | 825 | 2.239 | 0.1942 | Yes | ||
| 19 | ALDH3A2 | ENSG00000072210.19 | 840 | 2.223 | 0.2030 | Yes | ||
| 20 | CISH | ENSG00000114737.15 | 1052 | 2.083 | 0.2066 | Yes | ||
| 21 | RPS6KA2 | ENSG00000071242.12 | 1099 | 2.049 | 0.2139 | Yes | ||
| 22 | MAPK13 | ENSG00000156711.17 | 1253 | 1.961 | 0.2184 | Yes | ||
| 23 | PDCD4 | ENSG00000150593.18 | 1417 | 1.879 | 0.2223 | Yes | ||
| 24 | EGR3 | ENSG00000179388.9 | 1551 | 1.818 | 0.2266 | Yes | ||
| 25 | CD9 | ENSG00000010278.14 | 1562 | 1.814 | 0.2338 | Yes | ||
| 26 | BLVRB | ENSG00000090013.11 | 1648 | 1.782 | 0.2392 | Yes | ||
| 27 | PAPSS2 | ENSG00000198682.13 | 1705 | 1.753 | 0.2450 | Yes | ||
| 28 | CXCL12 | ENSG00000107562.16 | 1796 | 1.720 | 0.2500 | Yes | ||
| 29 | KRT19 | ENSG00000171345.13 | 1802 | 1.716 | 0.2569 | Yes | ||
| 30 | TJP3 | ENSG00000105289.15 | 1957 | 1.648 | 0.2601 | Yes | ||
| 31 | MAPT | ENSG00000186868.15 | 2157 | 1.580 | 0.2619 | Yes | ||
| 32 | CCN5 | ENSG00000064205.10 | 2260 | 1.545 | 0.2659 | Yes | ||
| 33 | PRKAR2B | ENSG00000005249.13 | 2294 | 1.537 | 0.2714 | Yes | ||
| 34 | NAB2 | ENSG00000166886.13 | 2304 | 1.534 | 0.2774 | Yes | ||
| 35 | PPIF | ENSG00000108179.14 | 2376 | 1.511 | 0.2820 | Yes | ||
| 36 | IL17RB | ENSG00000056736.10 | 2614 | 1.438 | 0.2824 | Yes | ||
| 37 | CAV1 | ENSG00000105974.12 | 2696 | 1.410 | 0.2862 | Yes | ||
| 38 | AFF1 | ENSG00000172493.20 | 2700 | 1.409 | 0.2919 | Yes | ||
| 39 | CPE | ENSG00000109472.14 | 2910 | 1.353 | 0.2926 | Yes | ||
| 40 | PTGER3 | ENSG00000050628.20 | 3014 | 1.330 | 0.2957 | Yes | ||
| 41 | CA12 | ENSG00000074410.14 | 3104 | 1.310 | 0.2990 | Yes | ||
| 42 | ISG20 | ENSG00000172183.15 | 3169 | 1.296 | 0.3028 | Yes | ||
| 43 | SCARB1 | ENSG00000073060.16 | 3609 | 1.204 | 0.2975 | Yes | ||
| 44 | DHCR7 | ENSG00000172893.15 | 3663 | 1.193 | 0.3011 | Yes | ||
| 45 | FDFT1 | ENSG00000079459.13 | 3715 | 1.180 | 0.3048 | Yes | ||
| 46 | CACNA2D2 | ENSG00000007402.11 | 3855 | 1.154 | 0.3063 | Yes | ||
| 47 | ELOVL5 | ENSG00000012660.14 | 3930 | 1.140 | 0.3092 | Yes | ||
| 48 | UGDH | ENSG00000109814.12 | 3997 | 1.129 | 0.3123 | Yes | ||
| 49 | DNAJC1 | ENSG00000136770.11 | 4342 | 1.070 | 0.3087 | Yes | ||
| 50 | PTGES | ENSG00000148344.11 | 4412 | 1.060 | 0.3114 | Yes | ||
| 51 | SGK1 | ENSG00000118515.11 | 4450 | 1.052 | 0.3148 | Yes | ||
| 52 | CCND1 | ENSG00000110092.3 | 4717 | 1.011 | 0.3128 | Yes | ||
| 53 | SEMA3B | ENSG00000012171.20 | 4765 | 1.004 | 0.3158 | Yes | ||
| 54 | MDK | ENSG00000110492.15 | 5041 | 0.962 | 0.3133 | Yes | ||
| 55 | FOXC1 | ENSG00000054598.9 | 5058 | 0.960 | 0.3169 | Yes | ||
| 56 | CXCL14 | ENSG00000145824.12 | 5106 | 0.953 | 0.3197 | Yes | ||
| 57 | PRLR | ENSG00000113494.17 | 5183 | 0.942 | 0.3218 | Yes | ||
| 58 | GLA | ENSG00000102393.10 | 5414 | 0.912 | 0.3202 | No | ||
| 59 | GAL | ENSG00000069482.7 | 5666 | 0.880 | 0.3179 | No | ||
| 60 | GPER1 | ENSG00000164850.15 | 6050 | 0.836 | 0.3124 | No | ||
| 61 | RAB31 | ENSG00000168461.13 | 6077 | 0.831 | 0.3152 | No | ||
| 62 | TFAP2C | ENSG00000087510.7 | 6149 | 0.822 | 0.3169 | No | ||
| 63 | ITPK1 | ENSG00000100605.17 | 6290 | 0.807 | 0.3170 | No | ||
| 64 | ST14 | ENSG00000149418.11 | 6638 | 0.774 | 0.3121 | No | ||
| 65 | FABP5 | ENSG00000164687.11 | 6871 | 0.750 | 0.3097 | No | ||
| 66 | ASS1 | ENSG00000130707.18 | 7046 | 0.733 | 0.3087 | No | ||
| 67 | TFPI2 | ENSG00000105825.14 | 7157 | 0.723 | 0.3091 | No | ||
| 68 | FARP1 | ENSG00000152767.17 | 7828 | 0.661 | 0.2962 | No | ||
| 69 | SCUBE2 | ENSG00000175356.13 | 7929 | 0.652 | 0.2965 | No | ||
| 70 | ST6GALNAC2 | ENSG00000070731.11 | 7997 | 0.647 | 0.2976 | No | ||
| 71 | TRIM29 | ENSG00000137699.17 | 8376 | 0.614 | 0.2913 | No | ||
| 72 | PRSS23 | ENSG00000150687.12 | 8732 | 0.588 | 0.2855 | No | ||
| 73 | SULT2B1 | ENSG00000088002.11 | 8911 | 0.575 | 0.2837 | No | ||
| 74 | SNX10 | ENSG00000086300.15 | 9608 | 0.529 | 0.2696 | No | ||
| 75 | TST | ENSG00000128311.14 | 9625 | 0.528 | 0.2714 | No | ||
| 76 | HSPA4L | ENSG00000164070.12 | 9659 | 0.525 | 0.2728 | No | ||
| 77 | FOS | ENSG00000170345.10 | 9664 | 0.525 | 0.2749 | No | ||
| 78 | DLG5 | ENSG00000151208.17 | 10138 | 0.501 | 0.2659 | No | ||
| 79 | PCP4 | ENSG00000183036.11 | 11202 | 0.433 | 0.2429 | No | ||
| 80 | ASCL1 | ENSG00000139352.4 | 11422 | 0.419 | 0.2395 | No | ||
| 81 | RET | ENSG00000165731.20 | 12408 | 0.369 | 0.2181 | No | ||
| 82 | CALCR | ENSG00000004948.15 | 13124 | 0.350 | 0.2028 | No | ||
| 83 | NMU | ENSG00000109255.11 | 13431 | 0.331 | 0.1971 | No | ||
| 84 | IMPA2 | ENSG00000141401.12 | 13538 | 0.324 | 0.1959 | No | ||
| 85 | ALDH3B1 | ENSG00000006534.16 | 14064 | 0.314 | 0.1850 | No | ||
| 86 | BATF | ENSG00000156127.7 | 14541 | 0.308 | 0.1752 | No | ||
| 87 | GJB3 | ENSG00000188910.8 | 16113 | 0.242 | 0.1395 | No | ||
| 88 | SLC2A8 | ENSG00000136856.18 | 16130 | 0.241 | 0.1402 | No | ||
| 89 | SERPINA3 | ENSG00000196136.17 | 16370 | 0.226 | 0.1355 | No | ||
| 90 | NPY1R | ENSG00000164128.7 | 16610 | 0.214 | 0.1308 | No | ||
| 91 | LSR | ENSG00000105699.16 | 17054 | 0.196 | 0.1213 | No | ||
| 92 | HR | ENSG00000168453.15 | 17070 | 0.194 | 0.1218 | No | ||
| 93 | KLK10 | ENSG00000129451.12 | 17579 | 0.170 | 0.1106 | No | ||
| 94 | CHPT1 | ENSG00000111666.11 | 17710 | 0.162 | 0.1083 | No | ||
| 95 | TSTA3 | ENSG00000104522.16 | 18779 | 0.108 | 0.0838 | No | ||
| 96 | DCXR | ENSG00000169738.8 | 19340 | 0.081 | 0.0711 | No | ||
| 97 | ID2 | ENSG00000115738.10 | 19442 | 0.074 | 0.0690 | No | ||
| 98 | JAK1 | ENSG00000162434.13 | 19624 | 0.065 | 0.0651 | No | ||
| 99 | ANXA9 | ENSG00000143412.10 | 19674 | 0.062 | 0.0642 | No | ||
| 100 | XRCC3 | ENSG00000126215.14 | 19768 | 0.058 | 0.0623 | No | ||
| 101 | TSPAN13 | ENSG00000106537.8 | 20061 | 0.045 | 0.0557 | No | ||
| 102 | KLK11 | ENSG00000167757.14 | 20427 | 0.026 | 0.0473 | No | ||
| 103 | MYOF | ENSG00000138119.17 | 20576 | 0.018 | 0.0439 | No | ||
| 104 | BAG1 | ENSG00000107262.23 | 21672 | -0.041 | 0.0185 | No | ||
| 105 | NXT1 | ENSG00000132661.4 | 22966 | -0.111 | -0.0111 | No | ||
| 106 | SIAH2 | ENSG00000181788.4 | 23013 | -0.113 | -0.0117 | No | ||
| 107 | CHST8 | ENSG00000124302.13 | 23588 | -0.146 | -0.0245 | No | ||
| 108 | WFS1 | ENSG00000109501.13 | 23770 | -0.155 | -0.0281 | No | ||
| 109 | SFN | ENSG00000175793.12 | 23845 | -0.161 | -0.0292 | No | ||
| 110 | SOX3 | ENSG00000134595.8 | 24382 | -0.192 | -0.0409 | No | ||
| 111 | AMFR | ENSG00000159461.15 | 24485 | -0.198 | -0.0424 | No | ||
| 112 | ETFB | ENSG00000105379.9 | 24745 | -0.215 | -0.0476 | No | ||
| 113 | HMGCS2 | ENSG00000134240.11 | 24990 | -0.223 | -0.0524 | No | ||
| 114 | ARL3 | ENSG00000138175.9 | 25638 | -0.236 | -0.0665 | No | ||
| 115 | RBBP8 | ENSG00000101773.19 | 26021 | -0.261 | -0.0743 | No | ||
| 116 | OVOL2 | ENSG00000125850.11 | 26280 | -0.263 | -0.0792 | No | ||
| 117 | PLXNB1 | ENSG00000164050.13 | 27135 | -0.283 | -0.0980 | No | ||
| 118 | TPSAB1 | ENSG00000172236.17 | 28979 | -0.351 | -0.1395 | No | ||
| 119 | AREG | ENSG00000109321.11 | 29159 | -0.357 | -0.1422 | No | ||
| 120 | SLC26A2 | ENSG00000155850.7 | 29548 | -0.377 | -0.1497 | No | ||
| 121 | KIF20A | ENSG00000112984.12 | 29750 | -0.391 | -0.1527 | No | ||
| 122 | HPRT1 | ENSG00000165704.15 | 29958 | -0.404 | -0.1559 | No | ||
| 123 | RABEP1 | ENSG00000029725.17 | 30408 | -0.432 | -0.1646 | No | ||
| 124 | SLC27A2 | ENSG00000140284.11 | 30591 | -0.441 | -0.1670 | No | ||
| 125 | JAK2 | ENSG00000096968.13 | 31758 | -0.501 | -0.1922 | No | ||
| 126 | GINS2 | ENSG00000131153.9 | 32126 | -0.521 | -0.1986 | No | ||
| 127 | XBP1 | ENSG00000100219.16 | 32221 | -0.529 | -0.1986 | No | ||
| 128 | STIL | ENSG00000123473.15 | 32707 | -0.560 | -0.2076 | No | ||
| 129 | FLNB | ENSG00000136068.14 | 32870 | -0.574 | -0.2090 | No | ||
| 130 | MEST | ENSG00000106484.16 | 32879 | -0.574 | -0.2069 | No | ||
| 131 | TH | ENSG00000180176.14 | 33344 | -0.609 | -0.2152 | No | ||
| 132 | PLK4 | ENSG00000142731.11 | 33445 | -0.616 | -0.2150 | No | ||
| 133 | TOP2A | ENSG00000131747.15 | 33556 | -0.624 | -0.2150 | No | ||
| 134 | SERPINA5 | ENSG00000188488.14 | 33654 | -0.635 | -0.2146 | No | ||
| 135 | KRT13 | ENSG00000171401.15 | 34387 | -0.691 | -0.2289 | No | ||
| 136 | KCNK5 | ENSG00000164626.9 | 34538 | -0.705 | -0.2295 | No | ||
| 137 | UNC13B | ENSG00000198722.14 | 34650 | -0.714 | -0.2291 | No | ||
| 138 | CCNA1 | ENSG00000133101.10 | 34815 | -0.726 | -0.2300 | No | ||
| 139 | CYP4F11 | ENSG00000171903.16 | 35089 | -0.750 | -0.2333 | No | ||
| 140 | CLIC3 | ENSG00000169583.13 | 35360 | -0.774 | -0.2364 | No | ||
| 141 | DUSP2 | ENSG00000158050.5 | 35482 | -0.785 | -0.2360 | No | ||
| 142 | PKP3 | ENSG00000184363.10 | 35564 | -0.794 | -0.2346 | No | ||
| 143 | CDC20 | ENSG00000117399.14 | 35877 | -0.828 | -0.2385 | No | ||
| 144 | BTG3 | ENSG00000154640.14 | 36393 | -0.887 | -0.2469 | No | ||
| 145 | RAPGEFL1 | ENSG00000108352.12 | 36612 | -0.917 | -0.2482 | No | ||
| 146 | CA2 | ENSG00000104267.10 | 36626 | -0.919 | -0.2447 | No | ||
| 147 | AGR2 | ENSG00000106541.12 | 36923 | -0.956 | -0.2477 | No | ||
| 148 | DHRS2 | ENSG00000100867.14 | 37225 | -0.995 | -0.2506 | No | ||
| 149 | CDH1 | ENSG00000039068.19 | 37235 | -0.997 | -0.2468 | No | ||
| 150 | TOB1 | ENSG00000141232.5 | 37303 | -1.007 | -0.2442 | No | ||
| 151 | LAMC2 | ENSG00000058085.15 | 37414 | -1.022 | -0.2426 | No | ||
| 152 | LLGL2 | ENSG00000073350.13 | 37508 | -1.035 | -0.2405 | No | ||
| 153 | DYNLT3 | ENSG00000165169.11 | 38084 | -1.132 | -0.2493 | No | ||
| 154 | MOCS2 | ENSG00000164172.19 | 38242 | -1.161 | -0.2482 | No | ||
| 155 | IGSF1 | ENSG00000147255.19 | 38251 | -1.163 | -0.2436 | No | ||
| 156 | METTL3 | ENSG00000165819.12 | 38364 | -1.181 | -0.2414 | No | ||
| 157 | PTPN6 | ENSG00000111679.17 | 38407 | -1.189 | -0.2375 | No | ||
| 158 | PDZK1 | ENSG00000174827.13 | 38667 | -1.237 | -0.2384 | No | ||
| 159 | SORD | ENSG00000140263.14 | 38729 | -1.249 | -0.2347 | No | ||
| 160 | TFF3 | ENSG00000160180.15 | 39084 | -1.327 | -0.2376 | No | ||
| 161 | PLAC1 | ENSG00000170965.10 | 39219 | -1.357 | -0.2351 | No | ||
| 162 | FGFR3 | ENSG00000068078.18 | 39445 | -1.408 | -0.2346 | No | ||
| 163 | SLC9A3R1 | ENSG00000109062.12 | 39524 | -1.426 | -0.2306 | No | ||
| 164 | COX6C | ENSG00000164919.11 | 39588 | -1.439 | -0.2261 | No | ||
| 165 | OPN3 | ENSG00000054277.14 | 39590 | -1.440 | -0.2203 | No | ||
| 166 | MYB | ENSG00000118513.19 | 39639 | -1.451 | -0.2155 | No | ||
| 167 | EMP2 | ENSG00000213853.10 | 39656 | -1.455 | -0.2099 | No | ||
| 168 | NRIP1 | ENSG00000180530.11 | 39986 | -1.544 | -0.2112 | No | ||
| 169 | FKBP5 | ENSG00000096060.14 | 40021 | -1.553 | -0.2056 | No | ||
| 170 | RNASEH2A | ENSG00000104889.6 | 40034 | -1.556 | -0.1995 | No | ||
| 171 | SLC1A4 | ENSG00000115902.11 | 40339 | -1.639 | -0.1999 | No | ||
| 172 | LTF | ENSG00000012223.13 | 40415 | -1.661 | -0.1949 | No | ||
| 173 | SERPINA1 | ENSG00000197249.13 | 40558 | -1.704 | -0.1912 | No | ||
| 174 | ABHD2 | ENSG00000140526.18 | 40689 | -1.746 | -0.1871 | No | ||
| 175 | FRK | ENSG00000111816.8 | 40827 | -1.793 | -0.1829 | No | ||
| 176 | SLC22A5 | ENSG00000197375.12 | 40836 | -1.797 | -0.1757 | No | ||
| 177 | NCOR2 | ENSG00000196498.13 | 40948 | -1.842 | -0.1708 | No | ||
| 178 | LARGE1 | ENSG00000133424.20 | 40958 | -1.846 | -0.1634 | No | ||
| 179 | CELSR2 | ENSG00000143126.8 | 41001 | -1.869 | -0.1567 | No | ||
| 180 | TPD52L1 | ENSG00000111907.21 | 41057 | -1.891 | -0.1503 | No | ||
| 181 | CKB | ENSG00000166165.13 | 41077 | -1.898 | -0.1430 | No | ||
| 182 | FKBP4 | ENSG00000004478.8 | 41079 | -1.899 | -0.1352 | No | ||
| 183 | SLC24A3 | ENSG00000185052.12 | 41129 | -1.924 | -0.1285 | No | ||
| 184 | GALE | ENSG00000117308.15 | 41501 | -2.130 | -0.1284 | No | ||
| 185 | HOMER2 | ENSG00000103942.13 | 41576 | -2.179 | -0.1212 | No | ||
| 186 | CDC6 | ENSG00000094804.12 | 41614 | -2.200 | -0.1130 | No | ||
| 187 | DNAJC12 | ENSG00000108176.15 | 41665 | -2.231 | -0.1051 | No | ||
| 188 | SLC16A1 | ENSG00000155380.11 | 41724 | -2.273 | -0.0971 | No | ||
| 189 | ABCA3 | ENSG00000167972.14 | 42113 | -2.590 | -0.0955 | No | ||
| 190 | TIAM1 | ENSG00000156299.13 | 42155 | -2.641 | -0.0857 | No | ||
| 191 | PLAAT3 | ENSG00000176485.11 | 42174 | -2.666 | -0.0752 | No | ||
| 192 | SLC29A1 | ENSG00000112759.18 | 42451 | -3.060 | -0.0691 | No | ||
| 193 | IDH2 | ENSG00000182054.9 | 42583 | -3.291 | -0.0586 | No | ||
| 194 | SLC7A5 | ENSG00000103257.9 | 42645 | -3.449 | -0.0459 | No | ||
| 195 | HSPB8 | ENSG00000152137.6 | 42810 | -3.949 | -0.0336 | No | ||
| 196 | TNNC1 | ENSG00000114854.8 | 43100 | -4.968 | -0.0200 | No | ||
| 197 | PDLIM3 | ENSG00000154553.15 | 43113 | -4.975 | 0.0001 | No |