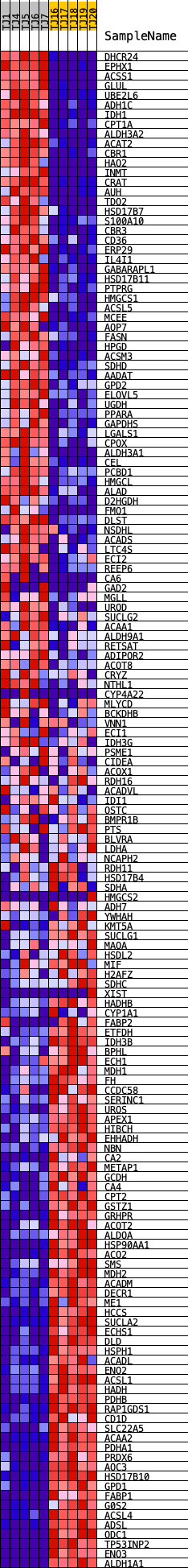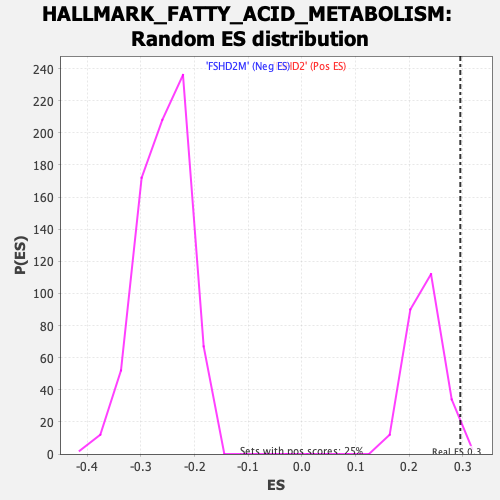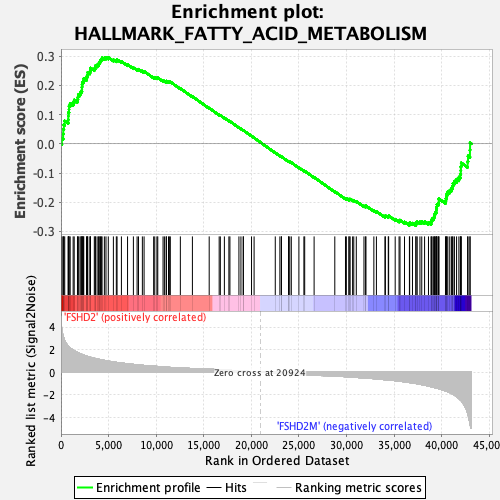
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_FATTY_ACID_METABOLISM |
| Enrichment Score (ES) | 0.29541174 |
| Normalized Enrichment Score (NES) | 1.2947463 |
| Nominal p-value | 0.01992032 |
| FDR q-value | 0.0646152 |
| FWER p-Value | 0.52 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DHCR24 | ENSG00000116133.13 | 91 | 3.768 | 0.0185 | Yes | ||
| 2 | EPHX1 | ENSG00000143819.12 | 223 | 3.237 | 0.0332 | Yes | ||
| 3 | ACSS1 | ENSG00000154930.15 | 241 | 3.164 | 0.0501 | Yes | ||
| 4 | GLUL | ENSG00000135821.18 | 300 | 2.991 | 0.0651 | Yes | ||
| 5 | UBE2L6 | ENSG00000156587.16 | 391 | 2.767 | 0.0781 | Yes | ||
| 6 | ADH1C | ENSG00000248144.6 | 760 | 2.300 | 0.0821 | Yes | ||
| 7 | IDH1 | ENSG00000138413.13 | 771 | 2.290 | 0.0944 | Yes | ||
| 8 | CPT1A | ENSG00000110090.13 | 776 | 2.287 | 0.1069 | Yes | ||
| 9 | ALDH3A2 | ENSG00000072210.19 | 840 | 2.223 | 0.1175 | Yes | ||
| 10 | ACAT2 | ENSG00000120437.9 | 847 | 2.219 | 0.1295 | Yes | ||
| 11 | CBR1 | ENSG00000159228.13 | 964 | 2.124 | 0.1385 | Yes | ||
| 12 | HAO2 | ENSG00000116882.14 | 1276 | 1.954 | 0.1419 | Yes | ||
| 13 | INMT | ENSG00000241644.2 | 1385 | 1.889 | 0.1497 | Yes | ||
| 14 | CRAT | ENSG00000095321.17 | 1730 | 1.743 | 0.1513 | Yes | ||
| 15 | AUH | ENSG00000148090.11 | 1776 | 1.726 | 0.1597 | Yes | ||
| 16 | TDO2 | ENSG00000151790.9 | 1817 | 1.707 | 0.1681 | Yes | ||
| 17 | HSD17B7 | ENSG00000132196.14 | 1994 | 1.633 | 0.1729 | Yes | ||
| 18 | S100A10 | ENSG00000197747.9 | 2097 | 1.600 | 0.1793 | Yes | ||
| 19 | CBR3 | ENSG00000159231.6 | 2198 | 1.567 | 0.1855 | Yes | ||
| 20 | CD36 | ENSG00000135218.19 | 2200 | 1.566 | 0.1941 | Yes | ||
| 21 | ERP29 | ENSG00000089248.7 | 2215 | 1.559 | 0.2023 | Yes | ||
| 22 | IL4I1 | ENSG00000104951.16 | 2235 | 1.554 | 0.2103 | Yes | ||
| 23 | GABARAPL1 | ENSG00000139112.11 | 2353 | 1.520 | 0.2159 | Yes | ||
| 24 | HSD17B11 | ENSG00000198189.11 | 2389 | 1.508 | 0.2234 | Yes | ||
| 25 | PTPRG | ENSG00000144724.20 | 2683 | 1.414 | 0.2243 | Yes | ||
| 26 | HMGCS1 | ENSG00000112972.15 | 2715 | 1.404 | 0.2312 | Yes | ||
| 27 | ACSL5 | ENSG00000197142.10 | 2795 | 1.380 | 0.2370 | Yes | ||
| 28 | MCEE | ENSG00000124370.11 | 2803 | 1.377 | 0.2443 | Yes | ||
| 29 | AQP7 | ENSG00000165269.13 | 3039 | 1.324 | 0.2461 | Yes | ||
| 30 | FASN | ENSG00000169710.9 | 3072 | 1.316 | 0.2526 | Yes | ||
| 31 | HPGD | ENSG00000164120.14 | 3099 | 1.310 | 0.2591 | Yes | ||
| 32 | ACSM3 | ENSG00000005187.12 | 3499 | 1.225 | 0.2565 | Yes | ||
| 33 | SDHD | ENSG00000204370.12 | 3584 | 1.209 | 0.2612 | Yes | ||
| 34 | AADAT | ENSG00000109576.14 | 3613 | 1.204 | 0.2671 | Yes | ||
| 35 | GPD2 | ENSG00000115159.16 | 3791 | 1.167 | 0.2694 | Yes | ||
| 36 | ELOVL5 | ENSG00000012660.14 | 3930 | 1.140 | 0.2724 | Yes | ||
| 37 | UGDH | ENSG00000109814.12 | 3997 | 1.129 | 0.2771 | Yes | ||
| 38 | PPARA | ENSG00000186951.16 | 4083 | 1.114 | 0.2812 | Yes | ||
| 39 | GAPDHS | ENSG00000105679.9 | 4145 | 1.104 | 0.2858 | Yes | ||
| 40 | LGALS1 | ENSG00000100097.12 | 4241 | 1.085 | 0.2895 | Yes | ||
| 41 | CPOX | ENSG00000080819.9 | 4313 | 1.075 | 0.2937 | Yes | ||
| 42 | ALDH3A1 | ENSG00000108602.18 | 4556 | 1.035 | 0.2938 | Yes | ||
| 43 | CEL | ENSG00000170835.14 | 4724 | 1.010 | 0.2954 | Yes | ||
| 44 | PCBD1 | ENSG00000166228.9 | 4968 | 0.973 | 0.2951 | No | ||
| 45 | HMGCL | ENSG00000117305.15 | 5503 | 0.900 | 0.2876 | No | ||
| 46 | ALAD | ENSG00000148218.16 | 5811 | 0.863 | 0.2852 | No | ||
| 47 | D2HGDH | ENSG00000180902.18 | 5888 | 0.854 | 0.2881 | No | ||
| 48 | FMO1 | ENSG00000010932.17 | 6346 | 0.803 | 0.2818 | No | ||
| 49 | DLST | ENSG00000119689.15 | 6991 | 0.739 | 0.2709 | No | ||
| 50 | NSDHL | ENSG00000147383.11 | 7604 | 0.679 | 0.2603 | No | ||
| 51 | ACADS | ENSG00000122971.9 | 7993 | 0.648 | 0.2548 | No | ||
| 52 | LTC4S | ENSG00000213316.10 | 8139 | 0.635 | 0.2549 | No | ||
| 53 | ECI2 | ENSG00000198721.12 | 8547 | 0.602 | 0.2488 | No | ||
| 54 | REEP6 | ENSG00000115255.11 | 8727 | 0.589 | 0.2478 | No | ||
| 55 | CA6 | ENSG00000131686.15 | 9732 | 0.522 | 0.2273 | No | ||
| 56 | GAD2 | ENSG00000136750.13 | 9849 | 0.520 | 0.2275 | No | ||
| 57 | MGLL | ENSG00000074416.14 | 10076 | 0.505 | 0.2250 | No | ||
| 58 | UROD | ENSG00000126088.14 | 10165 | 0.499 | 0.2256 | No | ||
| 59 | SUCLG2 | ENSG00000172340.14 | 10720 | 0.464 | 0.2153 | No | ||
| 60 | ACAA1 | ENSG00000060971.18 | 10861 | 0.454 | 0.2145 | No | ||
| 61 | ALDH9A1 | ENSG00000143149.12 | 11047 | 0.441 | 0.2126 | No | ||
| 62 | RETSAT | ENSG00000042445.14 | 11072 | 0.439 | 0.2145 | No | ||
| 63 | ADIPOR2 | ENSG00000006831.10 | 11292 | 0.427 | 0.2117 | No | ||
| 64 | ACOT8 | ENSG00000101473.17 | 11383 | 0.422 | 0.2119 | No | ||
| 65 | CRYZ | ENSG00000116791.14 | 11483 | 0.415 | 0.2119 | No | ||
| 66 | NTHL1 | ENSG00000065057.9 | 12537 | 0.363 | 0.1894 | No | ||
| 67 | CYP4A22 | ENSG00000162365.12 | 13803 | 0.322 | 0.1617 | No | ||
| 68 | MLYCD | ENSG00000103150.7 | 15567 | 0.271 | 0.1221 | No | ||
| 69 | BCKDHB | ENSG00000083123.15 | 16615 | 0.213 | 0.0989 | No | ||
| 70 | VNN1 | ENSG00000112299.8 | 16738 | 0.208 | 0.0972 | No | ||
| 71 | ECI1 | ENSG00000167969.13 | 17162 | 0.189 | 0.0884 | No | ||
| 72 | IDH3G | ENSG00000067829.19 | 17636 | 0.167 | 0.0783 | No | ||
| 73 | PSME1 | ENSG00000092010.15 | 17744 | 0.161 | 0.0767 | No | ||
| 74 | CIDEA | ENSG00000176194.18 | 18690 | 0.113 | 0.0553 | No | ||
| 75 | ACOX1 | ENSG00000161533.12 | 18902 | 0.103 | 0.0510 | No | ||
| 76 | RDH16 | ENSG00000139547.7 | 19139 | 0.093 | 0.0460 | No | ||
| 77 | ACADVL | ENSG00000072778.20 | 19159 | 0.091 | 0.0460 | No | ||
| 78 | IDI1 | ENSG00000067064.11 | 20014 | 0.048 | 0.0264 | No | ||
| 79 | OSTC | ENSG00000198856.13 | 20284 | 0.033 | 0.0203 | No | ||
| 80 | BMPR1B | ENSG00000138696.11 | 22508 | -0.085 | -0.0309 | No | ||
| 81 | PTS | ENSG00000150787.8 | 22993 | -0.112 | -0.0416 | No | ||
| 82 | BLVRA | ENSG00000106605.11 | 23148 | -0.121 | -0.0445 | No | ||
| 83 | LDHA | ENSG00000134333.14 | 23158 | -0.122 | -0.0441 | No | ||
| 84 | NCAPH2 | ENSG00000025770.19 | 23906 | -0.165 | -0.0605 | No | ||
| 85 | RDH11 | ENSG00000072042.13 | 23910 | -0.166 | -0.0597 | No | ||
| 86 | HSD17B4 | ENSG00000133835.16 | 24029 | -0.173 | -0.0615 | No | ||
| 87 | SDHA | ENSG00000073578.17 | 24192 | -0.182 | -0.0643 | No | ||
| 88 | HMGCS2 | ENSG00000134240.11 | 24990 | -0.223 | -0.0816 | No | ||
| 89 | ADH7 | ENSG00000196344.11 | 25515 | -0.228 | -0.0926 | No | ||
| 90 | YWHAH | ENSG00000128245.15 | 25599 | -0.233 | -0.0932 | No | ||
| 91 | KMT5A | ENSG00000183955.13 | 26588 | -0.273 | -0.1147 | No | ||
| 92 | SUCLG1 | ENSG00000163541.12 | 28761 | -0.349 | -0.1634 | No | ||
| 93 | MAOA | ENSG00000189221.9 | 29884 | -0.397 | -0.1873 | No | ||
| 94 | HSDL2 | ENSG00000119471.15 | 29975 | -0.405 | -0.1872 | No | ||
| 95 | MIF | ENSG00000240972.2 | 30243 | -0.424 | -0.1911 | No | ||
| 96 | H2AFZ | ENSG00000164032.12 | 30267 | -0.427 | -0.1893 | No | ||
| 97 | SDHC | ENSG00000143252.14 | 30372 | -0.429 | -0.1894 | No | ||
| 98 | XIST | ENSG00000229807.12 | 30632 | -0.445 | -0.1929 | No | ||
| 99 | HADHB | ENSG00000138029.14 | 30755 | -0.451 | -0.1933 | No | ||
| 100 | CYP1A1 | ENSG00000140465.14 | 31019 | -0.461 | -0.1969 | No | ||
| 101 | FABP2 | ENSG00000145384.4 | 31826 | -0.503 | -0.2129 | No | ||
| 102 | ETFDH | ENSG00000171503.12 | 32005 | -0.514 | -0.2143 | No | ||
| 103 | IDH3B | ENSG00000101365.21 | 32024 | -0.515 | -0.2119 | No | ||
| 104 | BPHL | ENSG00000137274.13 | 32864 | -0.573 | -0.2283 | No | ||
| 105 | ECH1 | ENSG00000104823.9 | 33133 | -0.593 | -0.2312 | No | ||
| 106 | MDH1 | ENSG00000014641.19 | 34029 | -0.662 | -0.2485 | No | ||
| 107 | FH | ENSG00000091483.6 | 34076 | -0.665 | -0.2459 | No | ||
| 108 | CCDC58 | ENSG00000160124.9 | 34379 | -0.691 | -0.2491 | No | ||
| 109 | SERINC1 | ENSG00000111897.7 | 34417 | -0.694 | -0.2462 | No | ||
| 110 | UROS | ENSG00000188690.15 | 35105 | -0.751 | -0.2581 | No | ||
| 111 | APEX1 | ENSG00000100823.12 | 35512 | -0.788 | -0.2632 | No | ||
| 112 | HIBCH | ENSG00000198130.16 | 35612 | -0.799 | -0.2612 | No | ||
| 113 | EHHADH | ENSG00000113790.11 | 36083 | -0.853 | -0.2674 | No | ||
| 114 | NBN | ENSG00000104320.13 | 36600 | -0.915 | -0.2744 | No | ||
| 115 | CA2 | ENSG00000104267.10 | 36626 | -0.919 | -0.2700 | No | ||
| 116 | METAP1 | ENSG00000164024.12 | 36921 | -0.956 | -0.2716 | No | ||
| 117 | GCDH | ENSG00000105607.13 | 37278 | -1.003 | -0.2744 | No | ||
| 118 | CA4 | ENSG00000167434.10 | 37287 | -1.005 | -0.2691 | No | ||
| 119 | CPT2 | ENSG00000157184.7 | 37417 | -1.022 | -0.2665 | No | ||
| 120 | GSTZ1 | ENSG00000100577.19 | 37706 | -1.068 | -0.2674 | No | ||
| 121 | GRHPR | ENSG00000137106.18 | 37896 | -1.100 | -0.2657 | No | ||
| 122 | ACOT2 | ENSG00000119673.14 | 38194 | -1.152 | -0.2663 | No | ||
| 123 | ALDOA | ENSG00000149925.20 | 38602 | -1.223 | -0.2691 | No | ||
| 124 | HSP90AA1 | ENSG00000080824.18 | 38879 | -1.285 | -0.2685 | No | ||
| 125 | ACO2 | ENSG00000100412.16 | 38900 | -1.290 | -0.2619 | No | ||
| 126 | SMS | ENSG00000102172.16 | 38982 | -1.305 | -0.2567 | No | ||
| 127 | MDH2 | ENSG00000146701.12 | 39149 | -1.341 | -0.2532 | No | ||
| 128 | ACADM | ENSG00000117054.13 | 39201 | -1.353 | -0.2470 | No | ||
| 129 | DECR1 | ENSG00000104325.7 | 39242 | -1.363 | -0.2405 | No | ||
| 130 | ME1 | ENSG00000065833.9 | 39319 | -1.382 | -0.2347 | No | ||
| 131 | HCCS | ENSG00000004961.15 | 39422 | -1.404 | -0.2294 | No | ||
| 132 | SUCLA2 | ENSG00000136143.16 | 39428 | -1.405 | -0.2218 | No | ||
| 133 | ECHS1 | ENSG00000127884.5 | 39473 | -1.417 | -0.2151 | No | ||
| 134 | DLD | ENSG00000091140.14 | 39495 | -1.421 | -0.2078 | No | ||
| 135 | HSPH1 | ENSG00000120694.20 | 39642 | -1.452 | -0.2032 | No | ||
| 136 | ACADL | ENSG00000115361.8 | 39688 | -1.463 | -0.1963 | No | ||
| 137 | ENO2 | ENSG00000111674.9 | 39689 | -1.464 | -0.1883 | No | ||
| 138 | ACSL1 | ENSG00000151726.14 | 40389 | -1.654 | -0.1955 | No | ||
| 139 | HADH | ENSG00000138796.16 | 40435 | -1.667 | -0.1874 | No | ||
| 140 | PDHB | ENSG00000168291.13 | 40487 | -1.684 | -0.1794 | No | ||
| 141 | RAP1GDS1 | ENSG00000138698.14 | 40532 | -1.697 | -0.1711 | No | ||
| 142 | CD1D | ENSG00000158473.6 | 40635 | -1.730 | -0.1640 | No | ||
| 143 | SLC22A5 | ENSG00000197375.12 | 40836 | -1.797 | -0.1589 | No | ||
| 144 | ACAA2 | ENSG00000167315.18 | 41011 | -1.873 | -0.1527 | No | ||
| 145 | PDHA1 | ENSG00000131828.14 | 41112 | -1.913 | -0.1445 | No | ||
| 146 | PRDX6 | ENSG00000117592.9 | 41173 | -1.946 | -0.1353 | No | ||
| 147 | AOC3 | ENSG00000131471.7 | 41327 | -2.021 | -0.1278 | No | ||
| 148 | HSD17B10 | ENSG00000072506.12 | 41554 | -2.165 | -0.1212 | No | ||
| 149 | GPD1 | ENSG00000167588.13 | 41796 | -2.325 | -0.1141 | No | ||
| 150 | FABP1 | ENSG00000163586.10 | 41951 | -2.456 | -0.1042 | No | ||
| 151 | G0S2 | ENSG00000123689.6 | 41996 | -2.485 | -0.0917 | No | ||
| 152 | ACSL4 | ENSG00000068366.20 | 42004 | -2.493 | -0.0782 | No | ||
| 153 | ADSL | ENSG00000239900.12 | 42041 | -2.520 | -0.0652 | No | ||
| 154 | ODC1 | ENSG00000115758.13 | 42710 | -3.622 | -0.0610 | No | ||
| 155 | TP53INP2 | ENSG00000078804.13 | 42758 | -3.743 | -0.0416 | No | ||
| 156 | ENO3 | ENSG00000108515.17 | 42958 | -4.537 | -0.0214 | No | ||
| 157 | ALDH1A1 | ENSG00000165092.13 | 42972 | -4.592 | 0.0034 | No |