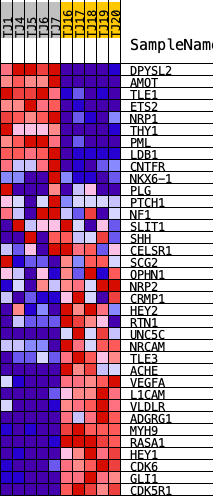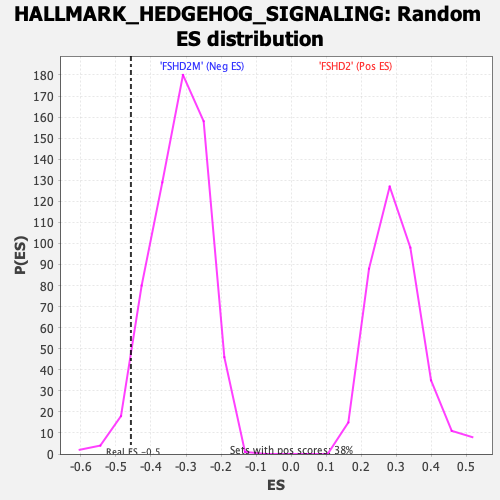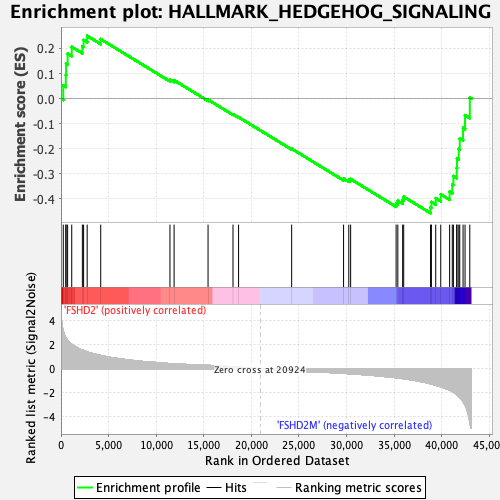
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_HEDGEHOG_SIGNALING |
| Enrichment Score (ES) | -0.45722064 |
| Normalized Enrichment Score (NES) | -1.4295732 |
| Nominal p-value | 0.03883495 |
| FDR q-value | 0.030804057 |
| FWER p-Value | 0.448 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DPYSL2 | ENSG00000092964.18 | 239 | 3.178 | 0.0525 | No | ||
| 2 | AMOT | ENSG00000126016.15 | 503 | 2.596 | 0.0938 | No | ||
| 3 | TLE1 | ENSG00000196781.15 | 537 | 2.564 | 0.1398 | No | ||
| 4 | ETS2 | ENSG00000157557.13 | 695 | 2.368 | 0.1794 | No | ||
| 5 | NRP1 | ENSG00000099250.18 | 1128 | 2.034 | 0.2065 | No | ||
| 6 | THY1 | ENSG00000154096.13 | 2244 | 1.552 | 0.2089 | No | ||
| 7 | PML | ENSG00000140464.20 | 2373 | 1.513 | 0.2336 | No | ||
| 8 | LDB1 | ENSG00000198728.10 | 2749 | 1.394 | 0.2503 | No | ||
| 9 | CNTFR | ENSG00000122756.15 | 4172 | 1.100 | 0.2374 | No | ||
| 10 | NKX6-1 | ENSG00000163623.9 | 11445 | 0.417 | 0.0762 | No | ||
| 11 | PLG | ENSG00000122194.18 | 11880 | 0.394 | 0.0734 | No | ||
| 12 | PTCH1 | ENSG00000185920.15 | 15447 | 0.278 | -0.0043 | No | ||
| 13 | NF1 | ENSG00000196712.17 | 18068 | 0.143 | -0.0625 | No | ||
| 14 | SLIT1 | ENSG00000187122.17 | 18654 | 0.116 | -0.0740 | No | ||
| 15 | SHH | ENSG00000164690.8 | 24230 | -0.184 | -0.2000 | No | ||
| 16 | CELSR1 | ENSG00000075275.16 | 29683 | -0.386 | -0.3195 | No | ||
| 17 | SCG2 | ENSG00000171951.5 | 30230 | -0.423 | -0.3245 | No | ||
| 18 | OPHN1 | ENSG00000079482.13 | 30426 | -0.433 | -0.3211 | No | ||
| 19 | NRP2 | ENSG00000118257.16 | 35202 | -0.760 | -0.4181 | No | ||
| 20 | CRMP1 | ENSG00000072832.15 | 35382 | -0.776 | -0.4081 | No | ||
| 21 | HEY2 | ENSG00000135547.9 | 35889 | -0.830 | -0.4047 | No | ||
| 22 | RTN1 | ENSG00000139970.17 | 36000 | -0.845 | -0.3918 | No | ||
| 23 | UNC5C | ENSG00000182168.15 | 38820 | -1.272 | -0.4340 | Yes | ||
| 24 | NRCAM | ENSG00000091129.20 | 38909 | -1.291 | -0.4125 | Yes | ||
| 25 | TLE3 | ENSG00000140332.16 | 39363 | -1.390 | -0.3976 | Yes | ||
| 26 | ACHE | ENSG00000087085.15 | 39892 | -1.520 | -0.3821 | Yes | ||
| 27 | VEGFA | ENSG00000112715.22 | 40824 | -1.792 | -0.3710 | Yes | ||
| 28 | L1CAM | ENSG00000198910.14 | 41102 | -1.908 | -0.3426 | Yes | ||
| 29 | VLDLR | ENSG00000147852.16 | 41219 | -1.967 | -0.3094 | Yes | ||
| 30 | ADGRG1 | ENSG00000205336.12 | 41567 | -2.173 | -0.2778 | Yes | ||
| 31 | MYH9 | ENSG00000100345.21 | 41599 | -2.193 | -0.2385 | Yes | ||
| 32 | RASA1 | ENSG00000145715.15 | 41787 | -2.315 | -0.2005 | Yes | ||
| 33 | HEY1 | ENSG00000164683.17 | 41887 | -2.391 | -0.1592 | Yes | ||
| 34 | CDK6 | ENSG00000105810.9 | 42235 | -2.748 | -0.1171 | Yes | ||
| 35 | GLI1 | ENSG00000111087.10 | 42443 | -3.048 | -0.0662 | Yes | ||
| 36 | CDK5R1 | ENSG00000176749.9 | 42942 | -4.486 | 0.0041 | Yes |