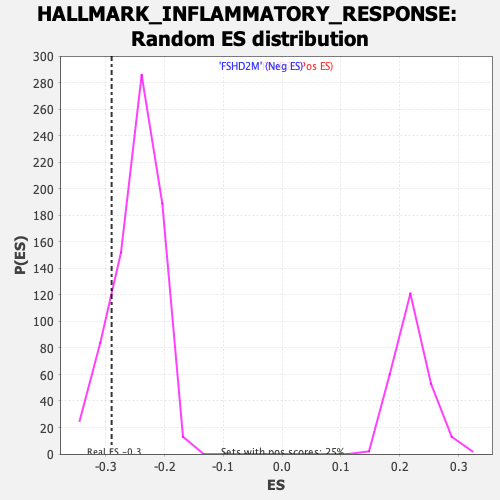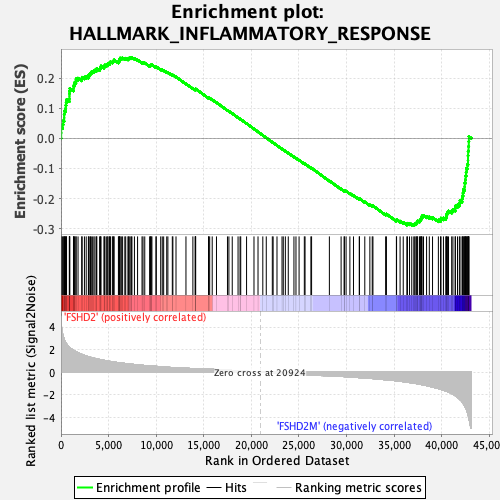
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_INFLAMMATORY_RESPONSE |
| Enrichment Score (ES) | -0.29006374 |
| Normalized Enrichment Score (NES) | -1.1721628 |
| Nominal p-value | 0.15086782 |
| FDR q-value | 0.22473446 |
| FWER p-Value | 0.997 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADM | ENSG00000148926.10 | 19 | 4.384 | 0.0178 | No | ||
| 2 | P2RX7 | ENSG00000089041.17 | 22 | 4.375 | 0.0359 | No | ||
| 3 | LPAR1 | ENSG00000198121.13 | 173 | 3.367 | 0.0464 | No | ||
| 4 | SLC1A2 | ENSG00000110436.13 | 232 | 3.201 | 0.0584 | No | ||
| 5 | TPBG | ENSG00000146242.8 | 331 | 2.893 | 0.0681 | No | ||
| 6 | PTGIR | ENSG00000160013.9 | 333 | 2.886 | 0.0801 | No | ||
| 7 | SCARF1 | ENSG00000074660.16 | 360 | 2.840 | 0.0913 | No | ||
| 8 | P2RX4 | ENSG00000135124.14 | 481 | 2.631 | 0.0995 | No | ||
| 9 | AHR | ENSG00000106546.14 | 498 | 2.602 | 0.1099 | No | ||
| 10 | IL1R1 | ENSG00000115594.12 | 542 | 2.559 | 0.1195 | No | ||
| 11 | TIMP1 | ENSG00000102265.12 | 582 | 2.491 | 0.1290 | No | ||
| 12 | IFNGR2 | ENSG00000159128.14 | 882 | 2.184 | 0.1311 | No | ||
| 13 | IFITM1 | ENSG00000185885.16 | 887 | 2.182 | 0.1401 | No | ||
| 14 | RTP4 | ENSG00000136514.3 | 888 | 2.182 | 0.1491 | No | ||
| 15 | LDLR | ENSG00000130164.13 | 900 | 2.174 | 0.1579 | No | ||
| 16 | LY6E | ENSG00000160932.11 | 924 | 2.150 | 0.1663 | No | ||
| 17 | CHST2 | ENSG00000175040.6 | 1311 | 1.932 | 0.1654 | No | ||
| 18 | GPR132 | ENSG00000183484.12 | 1315 | 1.930 | 0.1733 | No | ||
| 19 | IFNAR1 | ENSG00000142166.13 | 1384 | 1.890 | 0.1796 | No | ||
| 20 | IL15RA | ENSG00000134470.21 | 1434 | 1.873 | 0.1862 | No | ||
| 21 | TACR1 | ENSG00000115353.11 | 1567 | 1.814 | 0.1907 | No | ||
| 22 | CD14 | ENSG00000170458.14 | 1584 | 1.809 | 0.1979 | No | ||
| 23 | TNFRSF1B | ENSG00000028137.19 | 1790 | 1.722 | 0.2002 | No | ||
| 24 | TLR3 | ENSG00000164342.12 | 2177 | 1.573 | 0.1978 | No | ||
| 25 | CSF1 | ENSG00000184371.14 | 2219 | 1.559 | 0.2033 | No | ||
| 26 | RHOG | ENSG00000177105.10 | 2462 | 1.485 | 0.2039 | No | ||
| 27 | MXD1 | ENSG00000059728.11 | 2648 | 1.427 | 0.2055 | No | ||
| 28 | IRAK2 | ENSG00000134070.5 | 2876 | 1.361 | 0.2058 | No | ||
| 29 | EMP3 | ENSG00000142227.11 | 2904 | 1.355 | 0.2109 | No | ||
| 30 | HRH1 | ENSG00000196639.6 | 3030 | 1.327 | 0.2135 | No | ||
| 31 | CCL7 | ENSG00000108688.11 | 3096 | 1.310 | 0.2174 | No | ||
| 32 | RASGRP1 | ENSG00000172575.12 | 3180 | 1.292 | 0.2208 | No | ||
| 33 | KLF6 | ENSG00000067082.15 | 3324 | 1.262 | 0.2227 | No | ||
| 34 | IRF1 | ENSG00000125347.14 | 3458 | 1.235 | 0.2248 | No | ||
| 35 | ABCA1 | ENSG00000165029.16 | 3576 | 1.212 | 0.2271 | No | ||
| 36 | ITGB8 | ENSG00000105855.10 | 3741 | 1.176 | 0.2282 | No | ||
| 37 | TNFSF10 | ENSG00000121858.11 | 3755 | 1.173 | 0.2327 | No | ||
| 38 | EIF2AK2 | ENSG00000055332.18 | 4061 | 1.118 | 0.2303 | No | ||
| 39 | NAMPT | ENSG00000105835.12 | 4118 | 1.109 | 0.2336 | No | ||
| 40 | KCNA3 | ENSG00000177272.9 | 4151 | 1.103 | 0.2374 | No | ||
| 41 | ICAM1 | ENSG00000090339.9 | 4204 | 1.093 | 0.2408 | No | ||
| 42 | OSMR | ENSG00000145623.13 | 4521 | 1.041 | 0.2377 | No | ||
| 43 | TNFSF9 | ENSG00000125657.5 | 4548 | 1.036 | 0.2414 | No | ||
| 44 | BEST1 | ENSG00000167995.16 | 4619 | 1.026 | 0.2441 | No | ||
| 45 | CX3CL1 | ENSG00000006210.7 | 4796 | 0.999 | 0.2441 | No | ||
| 46 | SELL | ENSG00000188404.10 | 4919 | 0.981 | 0.2453 | No | ||
| 47 | EREG | ENSG00000124882.4 | 4928 | 0.979 | 0.2492 | No | ||
| 48 | GP1BA | ENSG00000185245.8 | 5108 | 0.953 | 0.2490 | No | ||
| 49 | CD55 | ENSG00000196352.15 | 5115 | 0.952 | 0.2528 | No | ||
| 50 | PDE4B | ENSG00000184588.18 | 5193 | 0.941 | 0.2550 | No | ||
| 51 | SLC11A2 | ENSG00000110911.16 | 5394 | 0.914 | 0.2541 | No | ||
| 52 | CXCL6 | ENSG00000124875.10 | 5459 | 0.905 | 0.2564 | No | ||
| 53 | AQP9 | ENSG00000103569.10 | 5524 | 0.897 | 0.2586 | No | ||
| 54 | GNA15 | ENSG00000060558.4 | 5581 | 0.890 | 0.2610 | No | ||
| 55 | CCR7 | ENSG00000126353.3 | 6031 | 0.837 | 0.2540 | No | ||
| 56 | GABBR1 | ENSG00000204681.11 | 6088 | 0.830 | 0.2562 | No | ||
| 57 | NFKB1 | ENSG00000109320.13 | 6111 | 0.828 | 0.2591 | No | ||
| 58 | IL15 | ENSG00000164136.17 | 6136 | 0.824 | 0.2620 | No | ||
| 59 | TAPBP | ENSG00000231925.11 | 6179 | 0.819 | 0.2644 | No | ||
| 60 | NLRP3 | ENSG00000162711.17 | 6212 | 0.815 | 0.2670 | No | ||
| 61 | GPC3 | ENSG00000147257.14 | 6374 | 0.800 | 0.2666 | No | ||
| 62 | IL2RB | ENSG00000100385.14 | 6472 | 0.790 | 0.2676 | No | ||
| 63 | SRI | ENSG00000075142.13 | 6706 | 0.767 | 0.2654 | No | ||
| 64 | IL18 | ENSG00000150782.12 | 6797 | 0.757 | 0.2665 | No | ||
| 65 | OLR1 | ENSG00000173391.9 | 7038 | 0.734 | 0.2639 | No | ||
| 66 | IL18RAP | ENSG00000115607.9 | 7077 | 0.730 | 0.2661 | No | ||
| 67 | GCH1 | ENSG00000131979.19 | 7161 | 0.723 | 0.2671 | No | ||
| 68 | NMI | ENSG00000123609.11 | 7263 | 0.713 | 0.2677 | No | ||
| 69 | CCL2 | ENSG00000108691.9 | 7391 | 0.701 | 0.2677 | No | ||
| 70 | GNAI3 | ENSG00000065135.11 | 7451 | 0.695 | 0.2692 | No | ||
| 71 | PTGER4 | ENSG00000171522.6 | 7713 | 0.671 | 0.2659 | No | ||
| 72 | CD40 | ENSG00000101017.14 | 8044 | 0.644 | 0.2609 | No | ||
| 73 | SPHK1 | ENSG00000176170.13 | 8512 | 0.605 | 0.2526 | No | ||
| 74 | MSR1 | ENSG00000038945.15 | 8615 | 0.598 | 0.2527 | No | ||
| 75 | CCL5 | ENSG00000271503.6 | 8798 | 0.583 | 0.2508 | No | ||
| 76 | TLR2 | ENSG00000137462.8 | 9313 | 0.549 | 0.2412 | No | ||
| 77 | ATP2C1 | ENSG00000017260.19 | 9381 | 0.545 | 0.2419 | No | ||
| 78 | CSF3R | ENSG00000119535.17 | 9393 | 0.544 | 0.2439 | No | ||
| 79 | MMP14 | ENSG00000157227.13 | 9489 | 0.539 | 0.2439 | No | ||
| 80 | CXCL9 | ENSG00000138755.6 | 9551 | 0.534 | 0.2447 | No | ||
| 81 | ADGRE1 | ENSG00000174837.15 | 9975 | 0.514 | 0.2370 | No | ||
| 82 | IL1A | ENSG00000115008.5 | 10008 | 0.511 | 0.2384 | No | ||
| 83 | NFKBIA | ENSG00000100906.10 | 10473 | 0.480 | 0.2295 | No | ||
| 84 | ACVR1B | ENSG00000135503.13 | 10636 | 0.469 | 0.2277 | No | ||
| 85 | LYN | ENSG00000254087.8 | 10784 | 0.460 | 0.2262 | No | ||
| 86 | LCP2 | ENSG00000043462.12 | 11106 | 0.438 | 0.2205 | No | ||
| 87 | F3 | ENSG00000117525.14 | 11227 | 0.431 | 0.2195 | No | ||
| 88 | IL18R1 | ENSG00000115604.10 | 11692 | 0.402 | 0.2104 | No | ||
| 89 | SELENOS | ENSG00000131871.15 | 11752 | 0.400 | 0.2107 | No | ||
| 90 | SCN1B | ENSG00000105711.12 | 12081 | 0.383 | 0.2046 | No | ||
| 91 | OPRK1 | ENSG00000082556.12 | 13126 | 0.350 | 0.1818 | No | ||
| 92 | PROK2 | ENSG00000163421.9 | 13853 | 0.322 | 0.1662 | No | ||
| 93 | PDPN | ENSG00000162493.16 | 14099 | 0.311 | 0.1618 | No | ||
| 94 | IRF7 | ENSG00000185507.21 | 14110 | 0.311 | 0.1628 | No | ||
| 95 | PTGER2 | ENSG00000125384.7 | 14137 | 0.308 | 0.1635 | No | ||
| 96 | C5AR1 | ENSG00000197405.8 | 14141 | 0.308 | 0.1647 | No | ||
| 97 | LCK | ENSG00000182866.17 | 15496 | 0.275 | 0.1343 | No | ||
| 98 | KCNMB2 | ENSG00000197584.12 | 15516 | 0.274 | 0.1350 | No | ||
| 99 | HPN | ENSG00000105707.14 | 15619 | 0.269 | 0.1338 | No | ||
| 100 | BST2 | ENSG00000130303.13 | 15884 | 0.255 | 0.1287 | No | ||
| 101 | AXL | ENSG00000167601.12 | 16332 | 0.229 | 0.1192 | No | ||
| 102 | IL1B | ENSG00000125538.12 | 17487 | 0.172 | 0.0930 | No | ||
| 103 | ADORA2B | ENSG00000170425.3 | 17637 | 0.167 | 0.0903 | No | ||
| 104 | CCL24 | ENSG00000106178.6 | 17988 | 0.147 | 0.0827 | No | ||
| 105 | PTAFR | ENSG00000169403.12 | 18599 | 0.119 | 0.0690 | No | ||
| 106 | CXCL11 | ENSG00000169248.13 | 18808 | 0.107 | 0.0646 | No | ||
| 107 | RELA | ENSG00000173039.19 | 18880 | 0.104 | 0.0634 | No | ||
| 108 | NPFFR2 | ENSG00000056291.17 | 19493 | 0.072 | 0.0494 | No | ||
| 109 | PSEN1 | ENSG00000080815.19 | 20272 | 0.034 | 0.0314 | No | ||
| 110 | SLC31A1 | ENSG00000136868.11 | 20696 | 0.013 | 0.0216 | No | ||
| 111 | APLNR | ENSG00000134817.10 | 21199 | -0.016 | 0.0100 | No | ||
| 112 | DCBLD2 | ENSG00000057019.16 | 21563 | -0.035 | 0.0017 | No | ||
| 113 | CD70 | ENSG00000125726.11 | 22197 | -0.068 | -0.0128 | No | ||
| 114 | IL12B | ENSG00000113302.4 | 22283 | -0.073 | -0.0144 | No | ||
| 115 | ACVR2A | ENSG00000121989.15 | 22689 | -0.096 | -0.0235 | No | ||
| 116 | IL7R | ENSG00000168685.15 | 23218 | -0.125 | -0.0353 | No | ||
| 117 | EBI3 | ENSG00000105246.6 | 23348 | -0.132 | -0.0377 | No | ||
| 118 | PIK3R5 | ENSG00000141506.13 | 23567 | -0.145 | -0.0422 | No | ||
| 119 | MARCO | ENSG00000019169.10 | 23874 | -0.163 | -0.0487 | No | ||
| 120 | RIPK2 | ENSG00000104312.8 | 24469 | -0.197 | -0.0617 | No | ||
| 121 | CXCR6 | ENSG00000172215.6 | 24683 | -0.210 | -0.0658 | No | ||
| 122 | FFAR2 | ENSG00000126262.4 | 25009 | -0.223 | -0.0724 | No | ||
| 123 | SEMA4D | ENSG00000187764.11 | 25560 | -0.231 | -0.0843 | No | ||
| 124 | NMUR1 | ENSG00000171596.7 | 25637 | -0.236 | -0.0850 | No | ||
| 125 | CD69 | ENSG00000110848.8 | 26251 | -0.263 | -0.0982 | No | ||
| 126 | MEP1A | ENSG00000112818.10 | 26323 | -0.263 | -0.0988 | No | ||
| 127 | SLC28A2 | ENSG00000137860.12 | 28192 | -0.323 | -0.1410 | No | ||
| 128 | CLEC5A | ENSG00000258227.7 | 29424 | -0.370 | -0.1681 | No | ||
| 129 | STAB1 | ENSG00000010327.10 | 29747 | -0.390 | -0.1740 | No | ||
| 130 | CXCL10 | ENSG00000169245.6 | 29765 | -0.392 | -0.1728 | No | ||
| 131 | SGMS2 | ENSG00000164023.14 | 29947 | -0.403 | -0.1753 | No | ||
| 132 | CCL22 | ENSG00000102962.5 | 30346 | -0.429 | -0.1828 | No | ||
| 133 | CMKLR1 | ENSG00000174600.14 | 30724 | -0.448 | -0.1897 | No | ||
| 134 | ITGB3 | ENSG00000259207.7 | 31330 | -0.481 | -0.2018 | No | ||
| 135 | BDKRB1 | ENSG00000100739.11 | 31357 | -0.483 | -0.2004 | No | ||
| 136 | CCL17 | ENSG00000102970.11 | 31912 | -0.509 | -0.2112 | No | ||
| 137 | SELE | ENSG00000007908.16 | 32444 | -0.541 | -0.2213 | No | ||
| 138 | RAF1 | ENSG00000132155.11 | 32694 | -0.559 | -0.2248 | No | ||
| 139 | KCNJ2 | ENSG00000123700.4 | 32768 | -0.566 | -0.2241 | No | ||
| 140 | PCDH7 | ENSG00000169851.15 | 34101 | -0.667 | -0.2524 | No | ||
| 141 | VIP | ENSG00000146469.13 | 34186 | -0.675 | -0.2515 | No | ||
| 142 | LTA | ENSG00000226979.8 | 35231 | -0.762 | -0.2727 | No | ||
| 143 | MYC | ENSG00000136997.19 | 35251 | -0.763 | -0.2700 | No | ||
| 144 | IL4R | ENSG00000077238.14 | 35619 | -0.801 | -0.2752 | No | ||
| 145 | NDP | ENSG00000124479.11 | 35951 | -0.838 | -0.2794 | No | ||
| 146 | RGS16 | ENSG00000143333.7 | 36334 | -0.882 | -0.2847 | No | ||
| 147 | IL10RA | ENSG00000110324.10 | 36391 | -0.887 | -0.2823 | No | ||
| 148 | TACR3 | ENSG00000169836.5 | 36611 | -0.917 | -0.2836 | No | ||
| 149 | OSM | ENSG00000099985.4 | 36846 | -0.947 | -0.2851 | No | ||
| 150 | CD48 | ENSG00000117091.10 | 37061 | -0.976 | -0.2860 | Yes | ||
| 151 | BTG2 | ENSG00000159388.6 | 37157 | -0.987 | -0.2841 | Yes | ||
| 152 | ADRM1 | ENSG00000130706.13 | 37285 | -1.005 | -0.2829 | Yes | ||
| 153 | IL6 | ENSG00000136244.12 | 37366 | -1.015 | -0.2805 | Yes | ||
| 154 | TNFAIP6 | ENSG00000123610.5 | 37407 | -1.021 | -0.2772 | Yes | ||
| 155 | CDKN1A | ENSG00000124762.13 | 37465 | -1.029 | -0.2743 | Yes | ||
| 156 | TLR1 | ENSG00000174125.8 | 37664 | -1.061 | -0.2745 | Yes | ||
| 157 | ABI1 | ENSG00000136754.17 | 37724 | -1.071 | -0.2714 | Yes | ||
| 158 | HIF1A | ENSG00000100644.17 | 37783 | -1.081 | -0.2683 | Yes | ||
| 159 | CCL20 | ENSG00000115009.13 | 37860 | -1.093 | -0.2655 | Yes | ||
| 160 | CXCL8 | ENSG00000169429.11 | 37884 | -1.098 | -0.2615 | Yes | ||
| 161 | CYBB | ENSG00000165168.7 | 37925 | -1.105 | -0.2578 | Yes | ||
| 162 | ICOSLG | ENSG00000160223.17 | 38080 | -1.131 | -0.2567 | Yes | ||
| 163 | SERPINE1 | ENSG00000106366.8 | 38403 | -1.189 | -0.2592 | Yes | ||
| 164 | LAMP3 | ENSG00000078081.8 | 38690 | -1.241 | -0.2607 | Yes | ||
| 165 | MET | ENSG00000105976.15 | 39028 | -1.316 | -0.2631 | Yes | ||
| 166 | PTPRE | ENSG00000132334.16 | 39641 | -1.452 | -0.2713 | Yes | ||
| 167 | CALCRL | ENSG00000064989.13 | 39870 | -1.513 | -0.2704 | Yes | ||
| 168 | EDN1 | ENSG00000078401.7 | 39917 | -1.527 | -0.2651 | Yes | ||
| 169 | ATP2B1 | ENSG00000070961.15 | 40174 | -1.593 | -0.2644 | Yes | ||
| 170 | ITGA5 | ENSG00000161638.11 | 40419 | -1.662 | -0.2632 | Yes | ||
| 171 | MEFV | ENSG00000103313.12 | 40494 | -1.686 | -0.2579 | Yes | ||
| 172 | P2RY2 | ENSG00000175591.11 | 40519 | -1.694 | -0.2514 | Yes | ||
| 173 | SLC31A2 | ENSG00000136867.11 | 40590 | -1.714 | -0.2459 | Yes | ||
| 174 | SLC7A1 | ENSG00000139514.13 | 40707 | -1.750 | -0.2414 | Yes | ||
| 175 | GPR183 | ENSG00000169508.7 | 41040 | -1.884 | -0.2413 | Yes | ||
| 176 | PLAUR | ENSG00000011422.12 | 41170 | -1.945 | -0.2362 | Yes | ||
| 177 | CCRL2 | ENSG00000121797.10 | 41412 | -2.072 | -0.2332 | Yes | ||
| 178 | INHBA | ENSG00000122641.11 | 41421 | -2.082 | -0.2247 | Yes | ||
| 179 | RGS1 | ENSG00000090104.12 | 41652 | -2.222 | -0.2208 | Yes | ||
| 180 | TNFRSF9 | ENSG00000049249.8 | 41845 | -2.364 | -0.2155 | Yes | ||
| 181 | SLC7A2 | ENSG00000003989.17 | 41914 | -2.420 | -0.2070 | Yes | ||
| 182 | CSF3 | ENSG00000108342.12 | 42130 | -2.616 | -0.2011 | Yes | ||
| 183 | ATP2A2 | ENSG00000174437.17 | 42170 | -2.663 | -0.1910 | Yes | ||
| 184 | FZD5 | ENSG00000163251.4 | 42233 | -2.747 | -0.1810 | Yes | ||
| 185 | C3AR1 | ENSG00000171860.5 | 42263 | -2.787 | -0.1701 | Yes | ||
| 186 | HAS2 | ENSG00000170961.7 | 42403 | -2.985 | -0.1609 | Yes | ||
| 187 | PVR | ENSG00000073008.15 | 42407 | -2.989 | -0.1485 | Yes | ||
| 188 | TNFSF15 | ENSG00000181634.8 | 42478 | -3.101 | -0.1373 | Yes | ||
| 189 | RNF144B | ENSG00000137393.9 | 42503 | -3.129 | -0.1248 | Yes | ||
| 190 | SLC4A4 | ENSG00000080493.17 | 42558 | -3.226 | -0.1127 | Yes | ||
| 191 | KIF1B | ENSG00000054523.17 | 42584 | -3.294 | -0.0996 | Yes | ||
| 192 | ROS1 | ENSG00000047936.10 | 42683 | -3.542 | -0.0871 | Yes | ||
| 193 | FPR1 | ENSG00000171051.8 | 42753 | -3.730 | -0.0732 | Yes | ||
| 194 | IL10 | ENSG00000136634.7 | 42757 | -3.743 | -0.0577 | Yes | ||
| 195 | CD82 | ENSG00000085117.12 | 42768 | -3.776 | -0.0423 | Yes | ||
| 196 | NOD2 | ENSG00000167207.13 | 42814 | -3.963 | -0.0268 | Yes | ||
| 197 | HBEGF | ENSG00000113070.8 | 42836 | -4.053 | -0.0105 | Yes | ||
| 198 | LIF | ENSG00000128342.5 | 42847 | -4.098 | 0.0063 | Yes |