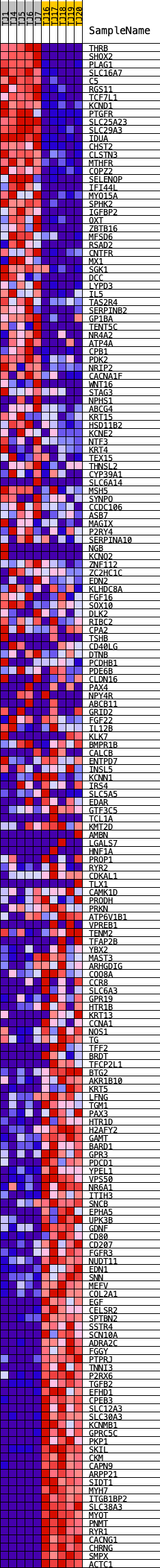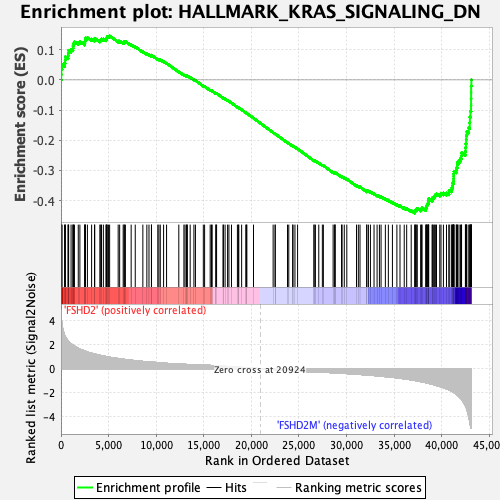
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
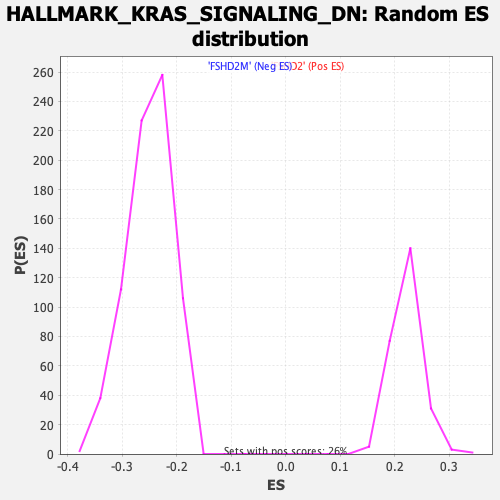
| GeneSet | HALLMARK_KRAS_SIGNALING_DN |
| Enrichment Score (ES) | -0.44192022 |
| Normalized Enrichment Score (NES) | -1.767425 |
| Nominal p-value | 0.0 |
| FDR q-value | 7.87772E-4 |
| FWER p-Value | 0.008 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | THRB | ENSG00000151090.19 | 25 | 4.314 | 0.0179 | No | ||
| 2 | SHOX2 | ENSG00000168779.19 | 40 | 4.232 | 0.0358 | No | ||
| 3 | PLAG1 | ENSG00000181690.8 | 114 | 3.626 | 0.0496 | No | ||
| 4 | SLC16A7 | ENSG00000118596.12 | 373 | 2.805 | 0.0556 | No | ||
| 5 | C5 | ENSG00000106804.7 | 443 | 2.687 | 0.0655 | No | ||
| 6 | RGS11 | ENSG00000076344.16 | 456 | 2.667 | 0.0767 | No | ||
| 7 | TCF7L1 | ENSG00000152284.5 | 744 | 2.321 | 0.0800 | No | ||
| 8 | KCND1 | ENSG00000102057.10 | 763 | 2.298 | 0.0894 | No | ||
| 9 | PTGFR | ENSG00000122420.10 | 801 | 2.260 | 0.0982 | No | ||
| 10 | SLC25A23 | ENSG00000125648.15 | 1058 | 2.079 | 0.1012 | No | ||
| 11 | SLC29A3 | ENSG00000198246.9 | 1251 | 1.962 | 0.1051 | No | ||
| 12 | IDUA | ENSG00000127415.13 | 1270 | 1.956 | 0.1131 | No | ||
| 13 | CHST2 | ENSG00000175040.6 | 1311 | 1.932 | 0.1205 | No | ||
| 14 | CLSTN3 | ENSG00000139182.15 | 1412 | 1.881 | 0.1262 | No | ||
| 15 | MTHFR | ENSG00000177000.12 | 1814 | 1.708 | 0.1242 | No | ||
| 16 | COPZ2 | ENSG00000005243.10 | 1984 | 1.638 | 0.1273 | No | ||
| 17 | SELENOP | ENSG00000250722.6 | 2483 | 1.478 | 0.1220 | No | ||
| 18 | IFI44L | ENSG00000137959.16 | 2506 | 1.473 | 0.1278 | No | ||
| 19 | MYO15A | ENSG00000091536.19 | 2545 | 1.458 | 0.1332 | No | ||
| 20 | SPHK2 | ENSG00000063176.16 | 2570 | 1.450 | 0.1389 | No | ||
| 21 | IGFBP2 | ENSG00000115457.10 | 2774 | 1.385 | 0.1401 | No | ||
| 22 | OXT | ENSG00000101405.3 | 3215 | 1.284 | 0.1353 | No | ||
| 23 | ZBTB16 | ENSG00000109906.14 | 3519 | 1.222 | 0.1335 | No | ||
| 24 | MFSD6 | ENSG00000151690.15 | 3545 | 1.217 | 0.1382 | No | ||
| 25 | RSAD2 | ENSG00000134321.12 | 4081 | 1.115 | 0.1305 | No | ||
| 26 | CNTFR | ENSG00000122756.15 | 4172 | 1.100 | 0.1331 | No | ||
| 27 | MX1 | ENSG00000157601.14 | 4253 | 1.082 | 0.1359 | No | ||
| 28 | SGK1 | ENSG00000118515.11 | 4450 | 1.052 | 0.1358 | No | ||
| 29 | DCC | ENSG00000187323.12 | 4719 | 1.011 | 0.1339 | No | ||
| 30 | LYPD3 | ENSG00000124466.9 | 4788 | 1.000 | 0.1366 | No | ||
| 31 | IL5 | ENSG00000113525.10 | 4807 | 0.996 | 0.1405 | No | ||
| 32 | TAS2R4 | ENSG00000127364.3 | 4857 | 0.988 | 0.1436 | No | ||
| 33 | SERPINB2 | ENSG00000197632.9 | 4983 | 0.970 | 0.1448 | No | ||
| 34 | GP1BA | ENSG00000185245.8 | 5108 | 0.953 | 0.1460 | No | ||
| 35 | TENT5C | ENSG00000183508.5 | 6006 | 0.841 | 0.1288 | No | ||
| 36 | NR4A2 | ENSG00000153234.14 | 6161 | 0.821 | 0.1287 | No | ||
| 37 | ATP4A | ENSG00000105675.9 | 6532 | 0.785 | 0.1234 | No | ||
| 38 | CPB1 | ENSG00000153002.12 | 6644 | 0.773 | 0.1242 | No | ||
| 39 | PDK2 | ENSG00000005882.12 | 6646 | 0.773 | 0.1275 | No | ||
| 40 | NRIP2 | ENSG00000053702.15 | 6765 | 0.761 | 0.1280 | No | ||
| 41 | CACNA1F | ENSG00000102001.12 | 7370 | 0.703 | 0.1169 | No | ||
| 42 | WNT16 | ENSG00000002745.13 | 7799 | 0.664 | 0.1098 | No | ||
| 43 | STAG3 | ENSG00000066923.17 | 8598 | 0.599 | 0.0938 | No | ||
| 44 | NPHS1 | ENSG00000161270.19 | 9018 | 0.568 | 0.0865 | No | ||
| 45 | ABCG4 | ENSG00000172350.10 | 9242 | 0.552 | 0.0836 | No | ||
| 46 | KRT15 | ENSG00000171346.16 | 9499 | 0.538 | 0.0800 | No | ||
| 47 | HSD11B2 | ENSG00000176387.7 | 9508 | 0.537 | 0.0821 | No | ||
| 48 | KCNE2 | ENSG00000159197.3 | 10168 | 0.499 | 0.0689 | No | ||
| 49 | NTF3 | ENSG00000185652.12 | 10285 | 0.492 | 0.0683 | No | ||
| 50 | KRT4 | ENSG00000170477.13 | 10429 | 0.483 | 0.0670 | No | ||
| 51 | TEX15 | ENSG00000133863.8 | 10777 | 0.460 | 0.0609 | No | ||
| 52 | THNSL2 | ENSG00000144115.16 | 11090 | 0.438 | 0.0555 | No | ||
| 53 | CYP39A1 | ENSG00000146233.8 | 12375 | 0.371 | 0.0272 | No | ||
| 54 | SLC6A14 | ENSG00000268104.3 | 12912 | 0.355 | 0.0163 | No | ||
| 55 | MSH5 | ENSG00000204410.15 | 13032 | 0.352 | 0.0150 | No | ||
| 56 | SYNPO | ENSG00000171992.13 | 13181 | 0.347 | 0.0130 | No | ||
| 57 | CCDC106 | ENSG00000173581.7 | 13230 | 0.344 | 0.0134 | No | ||
| 58 | ASB7 | ENSG00000183475.12 | 13270 | 0.341 | 0.0140 | No | ||
| 59 | MAGIX | ENSG00000269313.5 | 13569 | 0.323 | 0.0084 | No | ||
| 60 | P2RY4 | ENSG00000186912.6 | 13962 | 0.319 | 0.0006 | No | ||
| 61 | SERPINA10 | ENSG00000140093.10 | 14122 | 0.310 | -0.0017 | No | ||
| 62 | NGB | ENSG00000165553.4 | 14950 | 0.287 | -0.0198 | No | ||
| 63 | KCNQ2 | ENSG00000075043.18 | 15085 | 0.287 | -0.0217 | No | ||
| 64 | ZNF112 | ENSG00000062370.16 | 15672 | 0.265 | -0.0342 | No | ||
| 65 | ZC2HC1C | ENSG00000119703.14 | 15808 | 0.259 | -0.0362 | No | ||
| 66 | EDN2 | ENSG00000127129.10 | 15879 | 0.255 | -0.0367 | No | ||
| 67 | KLHDC8A | ENSG00000162873.14 | 16272 | 0.231 | -0.0449 | No | ||
| 68 | FGF16 | ENSG00000196468.8 | 16277 | 0.231 | -0.0440 | No | ||
| 69 | SOX10 | ENSG00000100146.17 | 16344 | 0.228 | -0.0445 | No | ||
| 70 | DLK2 | ENSG00000171462.15 | 17021 | 0.197 | -0.0594 | No | ||
| 71 | RIBC2 | ENSG00000128408.9 | 17060 | 0.195 | -0.0595 | No | ||
| 72 | CPA2 | ENSG00000158516.12 | 17237 | 0.186 | -0.0628 | No | ||
| 73 | TSHB | ENSG00000134200.3 | 17499 | 0.172 | -0.0681 | No | ||
| 74 | CD40LG | ENSG00000102245.7 | 17639 | 0.167 | -0.0706 | No | ||
| 75 | DTNB | ENSG00000138101.18 | 17908 | 0.151 | -0.0762 | No | ||
| 76 | PCDHB1 | ENSG00000171815.5 | 18530 | 0.122 | -0.0902 | No | ||
| 77 | PDE6B | ENSG00000133256.12 | 18632 | 0.117 | -0.0920 | No | ||
| 78 | CLDN16 | ENSG00000113946.3 | 18678 | 0.114 | -0.0926 | No | ||
| 79 | PAX4 | ENSG00000106331.17 | 18975 | 0.100 | -0.0991 | No | ||
| 80 | NPY4R | ENSG00000204174.8 | 19410 | 0.076 | -0.1088 | No | ||
| 81 | ABCB11 | ENSG00000073734.10 | 19500 | 0.072 | -0.1106 | No | ||
| 82 | GRID2 | ENSG00000152208.13 | 19514 | 0.071 | -0.1106 | No | ||
| 83 | FGF22 | ENSG00000070388.11 | 20221 | 0.037 | -0.1269 | No | ||
| 84 | IL12B | ENSG00000113302.4 | 22283 | -0.073 | -0.1746 | No | ||
| 85 | KLK7 | ENSG00000169035.12 | 22487 | -0.084 | -0.1790 | No | ||
| 86 | BMPR1B | ENSG00000138696.11 | 22508 | -0.085 | -0.1791 | No | ||
| 87 | CALCB | ENSG00000175868.14 | 23794 | -0.157 | -0.2083 | No | ||
| 88 | ENTPD7 | ENSG00000198018.7 | 23927 | -0.167 | -0.2107 | No | ||
| 89 | INSL5 | ENSG00000172410.4 | 24334 | -0.190 | -0.2193 | No | ||
| 90 | KCNN1 | ENSG00000105642.15 | 24372 | -0.192 | -0.2193 | No | ||
| 91 | IRS4 | ENSG00000133124.11 | 24575 | -0.203 | -0.2232 | No | ||
| 92 | SLC5A5 | ENSG00000105641.4 | 24852 | -0.221 | -0.2287 | No | ||
| 93 | EDAR | ENSG00000135960.10 | 26574 | -0.272 | -0.2676 | No | ||
| 94 | GTF3C5 | ENSG00000148308.17 | 26585 | -0.273 | -0.2666 | No | ||
| 95 | TCL1A | ENSG00000100721.11 | 26725 | -0.275 | -0.2687 | No | ||
| 96 | KMT2D | ENSG00000167548.15 | 27078 | -0.279 | -0.2757 | No | ||
| 97 | AMBN | ENSG00000178522.14 | 27454 | -0.297 | -0.2832 | No | ||
| 98 | LGALS7 | ENSG00000205076.4 | 27560 | -0.302 | -0.2843 | No | ||
| 99 | HNF1A | ENSG00000135100.17 | 28586 | -0.341 | -0.3067 | No | ||
| 100 | PROP1 | ENSG00000175325.2 | 28747 | -0.348 | -0.3090 | No | ||
| 101 | RYR2 | ENSG00000198626.17 | 28784 | -0.351 | -0.3083 | No | ||
| 102 | CDKAL1 | ENSG00000145996.11 | 28787 | -0.351 | -0.3068 | No | ||
| 103 | TLX1 | ENSG00000107807.13 | 29469 | -0.372 | -0.3211 | No | ||
| 104 | CAMK1D | ENSG00000183049.12 | 29527 | -0.376 | -0.3208 | No | ||
| 105 | PRODH | ENSG00000100033.16 | 29757 | -0.391 | -0.3245 | No | ||
| 106 | PRKN | ENSG00000185345.21 | 30023 | -0.409 | -0.3289 | No | ||
| 107 | ATP6V1B1 | ENSG00000116039.13 | 31042 | -0.463 | -0.3506 | No | ||
| 108 | VPREB1 | ENSG00000169575.5 | 31234 | -0.473 | -0.3530 | No | ||
| 109 | TENM2 | ENSG00000145934.16 | 31409 | -0.486 | -0.3550 | No | ||
| 110 | TFAP2B | ENSG00000008196.13 | 32116 | -0.521 | -0.3692 | No | ||
| 111 | YBX2 | ENSG00000006047.13 | 32122 | -0.521 | -0.3671 | No | ||
| 112 | MAST3 | ENSG00000099308.10 | 32294 | -0.534 | -0.3688 | No | ||
| 113 | ARHGDIG | ENSG00000242173.10 | 32505 | -0.543 | -0.3713 | No | ||
| 114 | COQ8A | ENSG00000163050.17 | 32883 | -0.574 | -0.3777 | No | ||
| 115 | CCR8 | ENSG00000179934.7 | 33204 | -0.599 | -0.3825 | No | ||
| 116 | SLC6A3 | ENSG00000142319.18 | 33448 | -0.617 | -0.3856 | No | ||
| 117 | GPR19 | ENSG00000183150.8 | 33621 | -0.631 | -0.3869 | No | ||
| 118 | HTR1B | ENSG00000135312.6 | 34073 | -0.665 | -0.3945 | No | ||
| 119 | KRT13 | ENSG00000171401.15 | 34387 | -0.691 | -0.3988 | No | ||
| 120 | CCNA1 | ENSG00000133101.10 | 34815 | -0.726 | -0.4057 | No | ||
| 121 | NOS1 | ENSG00000089250.19 | 35284 | -0.766 | -0.4133 | No | ||
| 122 | TG | ENSG00000042832.12 | 35610 | -0.799 | -0.4174 | No | ||
| 123 | TFF2 | ENSG00000160181.9 | 36055 | -0.850 | -0.4241 | No | ||
| 124 | BRDT | ENSG00000137948.18 | 36312 | -0.879 | -0.4263 | No | ||
| 125 | TFCP2L1 | ENSG00000115112.8 | 36787 | -0.939 | -0.4333 | No | ||
| 126 | BTG2 | ENSG00000159388.6 | 37157 | -0.987 | -0.4377 | Yes | ||
| 127 | AKR1B10 | ENSG00000198074.10 | 37193 | -0.991 | -0.4343 | Yes | ||
| 128 | KRT5 | ENSG00000186081.12 | 37249 | -0.999 | -0.4312 | Yes | ||
| 129 | LFNG | ENSG00000106003.13 | 37364 | -1.015 | -0.4295 | Yes | ||
| 130 | TGM1 | ENSG00000092295.12 | 37419 | -1.022 | -0.4264 | Yes | ||
| 131 | PAX3 | ENSG00000135903.19 | 37778 | -1.080 | -0.4301 | Yes | ||
| 132 | HTR1D | ENSG00000179546.4 | 37875 | -1.096 | -0.4277 | Yes | ||
| 133 | H2AFY2 | ENSG00000099284.14 | 37927 | -1.105 | -0.4241 | Yes | ||
| 134 | GAMT | ENSG00000130005.13 | 38317 | -1.174 | -0.4281 | Yes | ||
| 135 | BARD1 | ENSG00000138376.11 | 38347 | -1.179 | -0.4238 | Yes | ||
| 136 | GPR3 | ENSG00000181773.7 | 38401 | -1.187 | -0.4199 | Yes | ||
| 137 | PDCD1 | ENSG00000188389.11 | 38434 | -1.195 | -0.4155 | Yes | ||
| 138 | YPEL1 | ENSG00000100027.16 | 38485 | -1.204 | -0.4115 | Yes | ||
| 139 | VPS50 | ENSG00000004766.17 | 38559 | -1.216 | -0.4080 | Yes | ||
| 140 | NR6A1 | ENSG00000148200.17 | 38598 | -1.222 | -0.4036 | Yes | ||
| 141 | ITIH3 | ENSG00000162267.12 | 38611 | -1.225 | -0.3987 | Yes | ||
| 142 | SNCB | ENSG00000074317.11 | 38623 | -1.228 | -0.3937 | Yes | ||
| 143 | EPHA5 | ENSG00000145242.13 | 38998 | -1.309 | -0.3968 | Yes | ||
| 144 | UPK3B | ENSG00000243566.6 | 39001 | -1.310 | -0.3912 | Yes | ||
| 145 | GDNF | ENSG00000168621.15 | 39146 | -1.341 | -0.3888 | Yes | ||
| 146 | CD80 | ENSG00000121594.12 | 39254 | -1.364 | -0.3854 | Yes | ||
| 147 | CD207 | ENSG00000116031.9 | 39320 | -1.382 | -0.3810 | Yes | ||
| 148 | FGFR3 | ENSG00000068078.18 | 39445 | -1.408 | -0.3779 | Yes | ||
| 149 | NUDT11 | ENSG00000196368.5 | 39788 | -1.490 | -0.3794 | Yes | ||
| 150 | EDN1 | ENSG00000078401.7 | 39917 | -1.527 | -0.3759 | Yes | ||
| 151 | SNN | ENSG00000184602.6 | 40170 | -1.592 | -0.3749 | Yes | ||
| 152 | MEFV | ENSG00000103313.12 | 40494 | -1.686 | -0.3752 | Yes | ||
| 153 | COL2A1 | ENSG00000139219.19 | 40754 | -1.768 | -0.3736 | Yes | ||
| 154 | EGF | ENSG00000138798.13 | 40769 | -1.774 | -0.3664 | Yes | ||
| 155 | CELSR2 | ENSG00000143126.8 | 41001 | -1.869 | -0.3637 | Yes | ||
| 156 | SPTBN2 | ENSG00000173898.13 | 41074 | -1.896 | -0.3573 | Yes | ||
| 157 | SSTR4 | ENSG00000132671.5 | 41135 | -1.925 | -0.3504 | Yes | ||
| 158 | SCN10A | ENSG00000185313.8 | 41147 | -1.932 | -0.3424 | Yes | ||
| 159 | ADRA2C | ENSG00000184160.8 | 41237 | -1.974 | -0.3360 | Yes | ||
| 160 | FGGY | ENSG00000172456.17 | 41242 | -1.975 | -0.3276 | Yes | ||
| 161 | PTPRJ | ENSG00000149177.13 | 41249 | -1.977 | -0.3193 | Yes | ||
| 162 | TNNI3 | ENSG00000129991.13 | 41255 | -1.980 | -0.3109 | Yes | ||
| 163 | P2RX6 | ENSG00000099957.16 | 41282 | -1.994 | -0.3029 | Yes | ||
| 164 | TGFB2 | ENSG00000092969.12 | 41517 | -2.139 | -0.2992 | Yes | ||
| 165 | EFHD1 | ENSG00000115468.12 | 41559 | -2.169 | -0.2909 | Yes | ||
| 166 | CPEB3 | ENSG00000107864.15 | 41631 | -2.211 | -0.2830 | Yes | ||
| 167 | SLC12A3 | ENSG00000070915.9 | 41643 | -2.219 | -0.2738 | Yes | ||
| 168 | SLC30A3 | ENSG00000115194.11 | 41805 | -2.332 | -0.2675 | Yes | ||
| 169 | KCNMB1 | ENSG00000145936.9 | 41959 | -2.461 | -0.2605 | Yes | ||
| 170 | GPRC5C | ENSG00000170412.18 | 42060 | -2.537 | -0.2520 | Yes | ||
| 171 | PKP1 | ENSG00000081277.12 | 42078 | -2.554 | -0.2414 | Yes | ||
| 172 | SKIL | ENSG00000136603.14 | 42481 | -3.103 | -0.2375 | Yes | ||
| 173 | CKM | ENSG00000104879.5 | 42498 | -3.122 | -0.2244 | Yes | ||
| 174 | CAPN9 | ENSG00000135773.13 | 42532 | -3.169 | -0.2116 | Yes | ||
| 175 | ARPP21 | ENSG00000172995.16 | 42566 | -3.237 | -0.1985 | Yes | ||
| 176 | SIDT1 | ENSG00000072858.10 | 42578 | -3.265 | -0.1848 | Yes | ||
| 177 | MYH7 | ENSG00000092054.13 | 42632 | -3.422 | -0.1713 | Yes | ||
| 178 | ITGB1BP2 | ENSG00000147166.11 | 42834 | -4.040 | -0.1587 | Yes | ||
| 179 | SLC38A3 | ENSG00000188338.15 | 42936 | -4.446 | -0.1419 | Yes | ||
| 180 | MYOT | ENSG00000120729.9 | 42951 | -4.511 | -0.1229 | Yes | ||
| 181 | PNMT | ENSG00000141744.4 | 43014 | -4.774 | -0.1039 | Yes | ||
| 182 | RYR1 | ENSG00000196218.12 | 43074 | -4.942 | -0.0841 | Yes | ||
| 183 | CACNG1 | ENSG00000108878.5 | 43077 | -4.944 | -0.0629 | Yes | ||
| 184 | CHRNG | ENSG00000196811.13 | 43082 | -4.950 | -0.0418 | Yes | ||
| 185 | SMPX | ENSG00000091482.8 | 43084 | -4.952 | -0.0205 | Yes | ||
| 186 | ACTC1 | ENSG00000159251.8 | 43108 | -4.973 | 0.0003 | Yes |