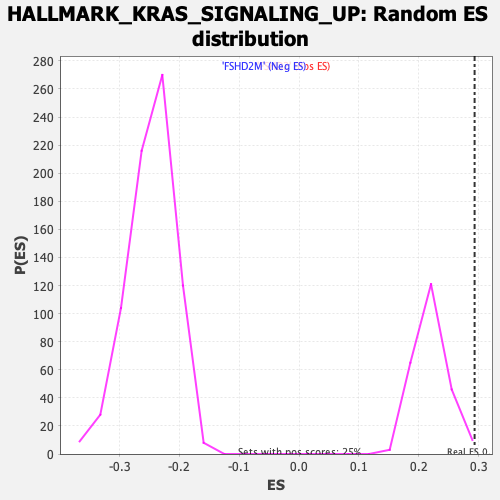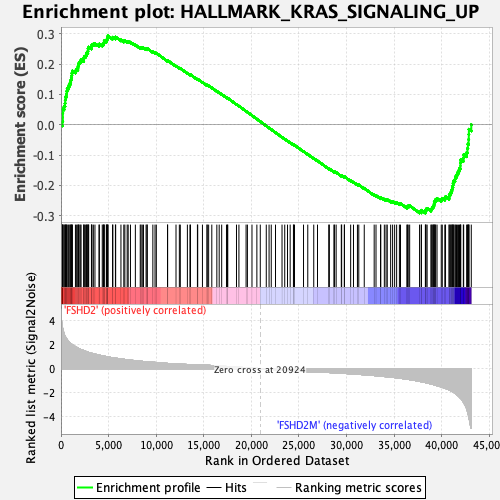
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_KRAS_SIGNALING_UP |
| Enrichment Score (ES) | 0.2934013 |
| Normalized Enrichment Score (NES) | 1.3333234 |
| Nominal p-value | 0.012244898 |
| FDR q-value | 0.057013694 |
| FWER p-Value | 0.403 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GPNMB | ENSG00000136235.16 | 163 | 3.424 | 0.0107 | Yes | ||
| 2 | GYPC | ENSG00000136732.16 | 189 | 3.328 | 0.0242 | Yes | ||
| 3 | ADGRA2 | ENSG00000020181.18 | 190 | 3.323 | 0.0382 | Yes | ||
| 4 | CFB | ENSG00000243649.8 | 196 | 3.306 | 0.0521 | Yes | ||
| 5 | ITGA2 | ENSG00000164171.11 | 327 | 2.899 | 0.0613 | Yes | ||
| 6 | KLF4 | ENSG00000136826.15 | 421 | 2.729 | 0.0707 | Yes | ||
| 7 | CTSS | ENSG00000163131.11 | 442 | 2.687 | 0.0816 | Yes | ||
| 8 | CAB39L | ENSG00000102547.19 | 467 | 2.647 | 0.0922 | Yes | ||
| 9 | AKAP12 | ENSG00000131016.17 | 561 | 2.526 | 0.1007 | Yes | ||
| 10 | LY96 | ENSG00000154589.7 | 593 | 2.480 | 0.1105 | Yes | ||
| 11 | EMP1 | ENSG00000134531.10 | 648 | 2.414 | 0.1194 | Yes | ||
| 12 | DNMBP | ENSG00000107554.17 | 772 | 2.289 | 0.1262 | Yes | ||
| 13 | ETV1 | ENSG00000006468.14 | 859 | 2.210 | 0.1336 | Yes | ||
| 14 | SATB1 | ENSG00000182568.17 | 963 | 2.125 | 0.1402 | Yes | ||
| 15 | PSMB8 | ENSG00000204264.11 | 1023 | 2.095 | 0.1476 | Yes | ||
| 16 | GNG11 | ENSG00000127920.6 | 1076 | 2.064 | 0.1551 | Yes | ||
| 17 | NRP1 | ENSG00000099250.18 | 1128 | 2.034 | 0.1626 | Yes | ||
| 18 | PRRX1 | ENSG00000116132.12 | 1144 | 2.020 | 0.1707 | Yes | ||
| 19 | ACE | ENSG00000159640.16 | 1197 | 1.991 | 0.1780 | Yes | ||
| 20 | EPB41L3 | ENSG00000082397.17 | 1516 | 1.836 | 0.1783 | Yes | ||
| 21 | DUSP6 | ENSG00000139318.8 | 1611 | 1.795 | 0.1837 | Yes | ||
| 22 | MAP7 | ENSG00000135525.18 | 1773 | 1.727 | 0.1873 | Yes | ||
| 23 | TNFRSF1B | ENSG00000028137.19 | 1790 | 1.722 | 0.1942 | Yes | ||
| 24 | IGFBP3 | ENSG00000146674.15 | 1828 | 1.702 | 0.2005 | Yes | ||
| 25 | APOD | ENSG00000189058.9 | 1901 | 1.668 | 0.2059 | Yes | ||
| 26 | TFPI | ENSG00000003436.16 | 2020 | 1.622 | 0.2100 | Yes | ||
| 27 | SPON1 | ENSG00000262655.4 | 2107 | 1.596 | 0.2147 | Yes | ||
| 28 | TMEM100 | ENSG00000166292.12 | 2391 | 1.508 | 0.2145 | Yes | ||
| 29 | IL1RL2 | ENSG00000115598.10 | 2396 | 1.506 | 0.2208 | Yes | ||
| 30 | ALDH1A3 | ENSG00000184254.17 | 2445 | 1.491 | 0.2260 | Yes | ||
| 31 | FUCA1 | ENSG00000179163.11 | 2594 | 1.443 | 0.2286 | Yes | ||
| 32 | MALL | ENSG00000144063.3 | 2663 | 1.422 | 0.2330 | Yes | ||
| 33 | ERO1A | ENSG00000197930.13 | 2702 | 1.409 | 0.2381 | Yes | ||
| 34 | HOXD11 | ENSG00000128713.14 | 2820 | 1.373 | 0.2412 | Yes | ||
| 35 | PTGS2 | ENSG00000073756.12 | 2837 | 1.370 | 0.2466 | Yes | ||
| 36 | PRKG2 | ENSG00000138669.9 | 2843 | 1.368 | 0.2523 | Yes | ||
| 37 | CPE | ENSG00000109472.14 | 2910 | 1.353 | 0.2565 | Yes | ||
| 38 | ALDH1A2 | ENSG00000128918.15 | 3198 | 1.290 | 0.2552 | Yes | ||
| 39 | BMP2 | ENSG00000125845.7 | 3203 | 1.288 | 0.2606 | Yes | ||
| 40 | TRIB2 | ENSG00000071575.11 | 3254 | 1.277 | 0.2648 | Yes | ||
| 41 | GUCY1A1 | ENSG00000164116.16 | 3397 | 1.246 | 0.2668 | Yes | ||
| 42 | MAFB | ENSG00000204103.4 | 3567 | 1.214 | 0.2680 | Yes | ||
| 43 | IL33 | ENSG00000137033.11 | 3995 | 1.130 | 0.2628 | Yes | ||
| 44 | SLPI | ENSG00000124107.5 | 4026 | 1.122 | 0.2668 | Yes | ||
| 45 | TMEM176B | ENSG00000106565.18 | 4334 | 1.071 | 0.2642 | Yes | ||
| 46 | SNAP25 | ENSG00000132639.12 | 4440 | 1.054 | 0.2662 | Yes | ||
| 47 | ABCB1 | ENSG00000085563.14 | 4477 | 1.048 | 0.2698 | Yes | ||
| 48 | MMP10 | ENSG00000166670.10 | 4529 | 1.040 | 0.2730 | Yes | ||
| 49 | TMEM176A | ENSG00000002933.9 | 4536 | 1.038 | 0.2773 | Yes | ||
| 50 | SEMA3B | ENSG00000012171.20 | 4765 | 1.004 | 0.2762 | Yes | ||
| 51 | AKT2 | ENSG00000105221.18 | 4775 | 1.001 | 0.2802 | Yes | ||
| 52 | ITGBL1 | ENSG00000198542.14 | 4835 | 0.992 | 0.2831 | Yes | ||
| 53 | CDADC1 | ENSG00000102543.14 | 4890 | 0.985 | 0.2860 | Yes | ||
| 54 | FBXO4 | ENSG00000151876.13 | 4926 | 0.980 | 0.2893 | Yes | ||
| 55 | EREG | ENSG00000124882.4 | 4928 | 0.979 | 0.2934 | Yes | ||
| 56 | PRDM1 | ENSG00000057657.17 | 5413 | 0.912 | 0.2860 | No | ||
| 57 | TRIB1 | ENSG00000173334.4 | 5448 | 0.906 | 0.2890 | No | ||
| 58 | SDCCAG8 | ENSG00000054282.16 | 5716 | 0.874 | 0.2865 | No | ||
| 59 | HSD11B1 | ENSG00000117594.10 | 5743 | 0.870 | 0.2896 | No | ||
| 60 | ZNF277 | ENSG00000198839.10 | 6297 | 0.807 | 0.2801 | No | ||
| 61 | NGF | ENSG00000134259.3 | 6597 | 0.778 | 0.2764 | No | ||
| 62 | TRAF1 | ENSG00000056558.11 | 6707 | 0.767 | 0.2771 | No | ||
| 63 | ADAMDEC1 | ENSG00000134028.14 | 6951 | 0.743 | 0.2746 | No | ||
| 64 | DOCK2 | ENSG00000134516.17 | 7056 | 0.733 | 0.2753 | No | ||
| 65 | ANGPTL4 | ENSG00000167772.12 | 7297 | 0.710 | 0.2727 | No | ||
| 66 | MAP3K1 | ENSG00000095015.6 | 7811 | 0.662 | 0.2635 | No | ||
| 67 | RELN | ENSG00000189056.14 | 8324 | 0.618 | 0.2542 | No | ||
| 68 | CFH | ENSG00000000971.15 | 8408 | 0.612 | 0.2549 | No | ||
| 69 | TSPAN1 | ENSG00000117472.10 | 8534 | 0.603 | 0.2545 | No | ||
| 70 | IKZF1 | ENSG00000185811.19 | 8664 | 0.594 | 0.2540 | No | ||
| 71 | NAP1L2 | ENSG00000186462.9 | 8937 | 0.572 | 0.2501 | No | ||
| 72 | TLR8 | ENSG00000101916.11 | 9048 | 0.567 | 0.2499 | No | ||
| 73 | CROT | ENSG00000005469.11 | 9050 | 0.567 | 0.2523 | No | ||
| 74 | CLEC4A | ENSG00000111729.14 | 9649 | 0.526 | 0.2406 | No | ||
| 75 | LCP1 | ENSG00000136167.14 | 9877 | 0.517 | 0.2375 | No | ||
| 76 | MTMR10 | ENSG00000166912.17 | 10014 | 0.511 | 0.2365 | No | ||
| 77 | ARG1 | ENSG00000118520.15 | 11189 | 0.433 | 0.2110 | No | ||
| 78 | PCP4 | ENSG00000183036.11 | 11202 | 0.433 | 0.2125 | No | ||
| 79 | SCN1B | ENSG00000105711.12 | 12081 | 0.383 | 0.1937 | No | ||
| 80 | BIRC3 | ENSG00000023445.14 | 12432 | 0.366 | 0.1871 | No | ||
| 81 | WNT7A | ENSG00000154764.6 | 12532 | 0.364 | 0.1863 | No | ||
| 82 | ADGRL4 | ENSG00000162618.15 | 13276 | 0.341 | 0.1704 | No | ||
| 83 | ZNF639 | ENSG00000121864.10 | 13545 | 0.324 | 0.1656 | No | ||
| 84 | CFHR2 | ENSG00000080910.13 | 13602 | 0.322 | 0.1656 | No | ||
| 85 | PIGR | ENSG00000162896.6 | 14330 | 0.308 | 0.1500 | No | ||
| 86 | PPBP | ENSG00000163736.4 | 14361 | 0.308 | 0.1506 | No | ||
| 87 | PPP1R15A | ENSG00000087074.8 | 14858 | 0.289 | 0.1402 | No | ||
| 88 | SPARCL1 | ENSG00000152583.12 | 15335 | 0.283 | 0.1304 | No | ||
| 89 | CBX8 | ENSG00000141570.11 | 15355 | 0.282 | 0.1311 | No | ||
| 90 | GALNT3 | ENSG00000115339.14 | 15510 | 0.274 | 0.1287 | No | ||
| 91 | PCSK1N | ENSG00000102109.9 | 15841 | 0.258 | 0.1221 | No | ||
| 92 | SERPINA3 | ENSG00000196136.17 | 16370 | 0.226 | 0.1107 | No | ||
| 93 | EVI5 | ENSG00000067208.14 | 16617 | 0.213 | 0.1059 | No | ||
| 94 | ANKH | ENSG00000154122.14 | 16879 | 0.203 | 0.1007 | No | ||
| 95 | MMD | ENSG00000108960.9 | 17383 | 0.179 | 0.0897 | No | ||
| 96 | IL1B | ENSG00000125538.12 | 17487 | 0.172 | 0.0880 | No | ||
| 97 | SCG5 | ENSG00000166922.8 | 17518 | 0.171 | 0.0881 | No | ||
| 98 | HDAC9 | ENSG00000048052.21 | 18434 | 0.129 | 0.0673 | No | ||
| 99 | CIDEA | ENSG00000176194.18 | 18690 | 0.113 | 0.0618 | No | ||
| 100 | ID2 | ENSG00000115738.10 | 19442 | 0.074 | 0.0446 | No | ||
| 101 | PEG3 | ENSG00000198300.14 | 19582 | 0.067 | 0.0417 | No | ||
| 102 | TSPAN13 | ENSG00000106537.8 | 20061 | 0.045 | 0.0307 | No | ||
| 103 | SPP1 | ENSG00000118785.14 | 20575 | 0.018 | 0.0189 | No | ||
| 104 | MMP11 | ENSG00000099953.10 | 20933 | -0.001 | 0.0106 | No | ||
| 105 | DCBLD2 | ENSG00000057019.16 | 21563 | -0.035 | -0.0040 | No | ||
| 106 | TOR1AIP2 | ENSG00000169905.12 | 21857 | -0.049 | -0.0106 | No | ||
| 107 | PTBP2 | ENSG00000117569.18 | 22086 | -0.062 | -0.0156 | No | ||
| 108 | ENG | ENSG00000106991.13 | 22533 | -0.086 | -0.0257 | No | ||
| 109 | IL7R | ENSG00000168685.15 | 23218 | -0.125 | -0.0411 | No | ||
| 110 | PECAM1 | ENSG00000261371.6 | 23502 | -0.141 | -0.0471 | No | ||
| 111 | ETV4 | ENSG00000175832.13 | 23795 | -0.157 | -0.0532 | No | ||
| 112 | TMEM158 | ENSG00000249992.2 | 24073 | -0.175 | -0.0589 | No | ||
| 113 | NR1H4 | ENSG00000012504.15 | 24416 | -0.194 | -0.0661 | No | ||
| 114 | SCG3 | ENSG00000104112.9 | 24442 | -0.195 | -0.0658 | No | ||
| 115 | NR0B2 | ENSG00000131910.5 | 24529 | -0.200 | -0.0670 | No | ||
| 116 | GFPT2 | ENSG00000131459.13 | 25476 | -0.226 | -0.0881 | No | ||
| 117 | SOX9 | ENSG00000125398.8 | 25922 | -0.255 | -0.0973 | No | ||
| 118 | HIST1H2BB | ENSG00000276410.3 | 26550 | -0.270 | -0.1108 | No | ||
| 119 | MPZL2 | ENSG00000149573.9 | 26944 | -0.275 | -0.1188 | No | ||
| 120 | F2RL1 | ENSG00000164251.5 | 28125 | -0.319 | -0.1450 | No | ||
| 121 | CBL | ENSG00000110395.6 | 28190 | -0.323 | -0.1451 | No | ||
| 122 | USH1C | ENSG00000006611.16 | 28668 | -0.342 | -0.1547 | No | ||
| 123 | ATG10 | ENSG00000152348.16 | 28738 | -0.347 | -0.1549 | No | ||
| 124 | IRF8 | ENSG00000140968.11 | 28903 | -0.351 | -0.1572 | No | ||
| 125 | RABGAP1L | ENSG00000152061.23 | 29444 | -0.371 | -0.1682 | No | ||
| 126 | BTBD3 | ENSG00000132640.15 | 29483 | -0.373 | -0.1675 | No | ||
| 127 | CXCL10 | ENSG00000169245.6 | 29765 | -0.392 | -0.1724 | No | ||
| 128 | PLEK2 | ENSG00000100558.9 | 29774 | -0.392 | -0.1710 | No | ||
| 129 | ADAM8 | ENSG00000151651.16 | 30427 | -0.433 | -0.1843 | No | ||
| 130 | CMKLR1 | ENSG00000174600.14 | 30724 | -0.448 | -0.1893 | No | ||
| 131 | MMP9 | ENSG00000100985.7 | 31139 | -0.471 | -0.1970 | No | ||
| 132 | EPHB2 | ENSG00000133216.16 | 31281 | -0.477 | -0.1982 | No | ||
| 133 | JUP | ENSG00000173801.17 | 31855 | -0.505 | -0.2095 | No | ||
| 134 | GABRA3 | ENSG00000011677.13 | 32898 | -0.576 | -0.2313 | No | ||
| 135 | ETS1 | ENSG00000134954.14 | 33090 | -0.589 | -0.2333 | No | ||
| 136 | LAPTM5 | ENSG00000162511.8 | 33566 | -0.626 | -0.2417 | No | ||
| 137 | WDR33 | ENSG00000136709.12 | 33590 | -0.628 | -0.2396 | No | ||
| 138 | KIF5C | ENSG00000168280.17 | 33966 | -0.659 | -0.2455 | No | ||
| 139 | RBM4 | ENSG00000173933.20 | 34144 | -0.671 | -0.2468 | No | ||
| 140 | CXCR4 | ENSG00000121966.6 | 34272 | -0.681 | -0.2469 | No | ||
| 141 | PTCD2 | ENSG00000049883.15 | 34628 | -0.713 | -0.2521 | No | ||
| 142 | ST6GAL1 | ENSG00000073849.15 | 34834 | -0.728 | -0.2538 | No | ||
| 143 | RBP4 | ENSG00000138207.14 | 35047 | -0.745 | -0.2556 | No | ||
| 144 | TPH1 | ENSG00000129167.9 | 35258 | -0.764 | -0.2573 | No | ||
| 145 | PRELID3B | ENSG00000101166.16 | 35547 | -0.792 | -0.2606 | No | ||
| 146 | ANO1 | ENSG00000131620.17 | 35662 | -0.805 | -0.2599 | No | ||
| 147 | RGS16 | ENSG00000143333.7 | 36334 | -0.882 | -0.2718 | No | ||
| 148 | IL10RA | ENSG00000110324.10 | 36391 | -0.887 | -0.2694 | No | ||
| 149 | SNAP91 | ENSG00000065609.14 | 36474 | -0.897 | -0.2675 | No | ||
| 150 | CA2 | ENSG00000104267.10 | 36626 | -0.919 | -0.2671 | No | ||
| 151 | YRDC | ENSG00000196449.4 | 37665 | -1.061 | -0.2868 | No | ||
| 152 | ETV5 | ENSG00000244405.8 | 37843 | -1.090 | -0.2863 | No | ||
| 153 | CCL20 | ENSG00000115009.13 | 37860 | -1.093 | -0.2821 | No | ||
| 154 | AVL9 | ENSG00000105778.19 | 38275 | -1.167 | -0.2868 | No | ||
| 155 | MYCN | ENSG00000134323.12 | 38301 | -1.172 | -0.2824 | No | ||
| 156 | LAT2 | ENSG00000086730.17 | 38342 | -1.178 | -0.2784 | No | ||
| 157 | SPRY2 | ENSG00000136158.12 | 38459 | -1.199 | -0.2760 | No | ||
| 158 | PLVAP | ENSG00000130300.9 | 38855 | -1.279 | -0.2798 | No | ||
| 159 | CD37 | ENSG00000104894.12 | 38942 | -1.297 | -0.2763 | No | ||
| 160 | NIN | ENSG00000100503.23 | 38999 | -1.310 | -0.2721 | No | ||
| 161 | HKDC1 | ENSG00000156510.13 | 39104 | -1.332 | -0.2689 | No | ||
| 162 | CCSER2 | ENSG00000107771.17 | 39170 | -1.346 | -0.2647 | No | ||
| 163 | STRN | ENSG00000115808.12 | 39197 | -1.352 | -0.2596 | No | ||
| 164 | MAP4K1 | ENSG00000104814.13 | 39205 | -1.355 | -0.2540 | No | ||
| 165 | PDCD1LG2 | ENSG00000197646.8 | 39306 | -1.379 | -0.2505 | No | ||
| 166 | TNFAIP3 | ENSG00000118503.15 | 39359 | -1.390 | -0.2459 | No | ||
| 167 | BPGM | ENSG00000172331.12 | 39501 | -1.423 | -0.2431 | No | ||
| 168 | ADAM17 | ENSG00000151694.14 | 39944 | -1.533 | -0.2469 | No | ||
| 169 | FGF9 | ENSG00000102678.7 | 40051 | -1.560 | -0.2428 | No | ||
| 170 | CSF2 | ENSG00000164400.6 | 40320 | -1.633 | -0.2422 | No | ||
| 171 | GPRC5B | ENSG00000167191.12 | 40394 | -1.657 | -0.2369 | No | ||
| 172 | IGF2 | ENSG00000167244.21 | 40765 | -1.773 | -0.2380 | No | ||
| 173 | AMMECR1 | ENSG00000101935.10 | 40781 | -1.779 | -0.2308 | No | ||
| 174 | BTC | ENSG00000174808.12 | 40898 | -1.823 | -0.2258 | No | ||
| 175 | GADD45G | ENSG00000130222.11 | 40980 | -1.854 | -0.2199 | No | ||
| 176 | CBR4 | ENSG00000145439.12 | 41059 | -1.892 | -0.2137 | No | ||
| 177 | GLRX | ENSG00000173221.14 | 41105 | -1.909 | -0.2067 | No | ||
| 178 | PLAUR | ENSG00000011422.12 | 41170 | -1.945 | -0.1999 | No | ||
| 179 | USP12 | ENSG00000152484.14 | 41186 | -1.952 | -0.1920 | No | ||
| 180 | ITGB2 | ENSG00000160255.18 | 41252 | -1.979 | -0.1852 | No | ||
| 181 | VWA5A | ENSG00000110002.16 | 41384 | -2.054 | -0.1795 | No | ||
| 182 | INHBA | ENSG00000122641.11 | 41421 | -2.082 | -0.1716 | No | ||
| 183 | TSPAN7 | ENSG00000156298.12 | 41547 | -2.161 | -0.1654 | No | ||
| 184 | PLAT | ENSG00000104368.18 | 41640 | -2.216 | -0.1581 | No | ||
| 185 | KCNN4 | ENSG00000104783.14 | 41760 | -2.297 | -0.1512 | No | ||
| 186 | IL2RG | ENSG00000147168.12 | 41862 | -2.374 | -0.1435 | No | ||
| 187 | FLT4 | ENSG00000037280.16 | 41958 | -2.460 | -0.1353 | No | ||
| 188 | CCND2 | ENSG00000118971.8 | 41960 | -2.462 | -0.1249 | No | ||
| 189 | G0S2 | ENSG00000123689.6 | 41996 | -2.485 | -0.1152 | No | ||
| 190 | C3AR1 | ENSG00000171860.5 | 42263 | -2.787 | -0.1097 | No | ||
| 191 | FCER1G | ENSG00000158869.11 | 42299 | -2.847 | -0.0984 | No | ||
| 192 | PTPRR | ENSG00000153233.13 | 42612 | -3.370 | -0.0915 | No | ||
| 193 | PLAU | ENSG00000122861.16 | 42698 | -3.591 | -0.0783 | No | ||
| 194 | ANXA10 | ENSG00000109511.12 | 42740 | -3.706 | -0.0635 | No | ||
| 195 | F13A1 | ENSG00000124491.15 | 42818 | -3.981 | -0.0485 | No | ||
| 196 | HBEGF | ENSG00000113070.8 | 42836 | -4.053 | -0.0318 | No | ||
| 197 | LIF | ENSG00000128342.5 | 42847 | -4.098 | -0.0147 | No | ||
| 198 | TNNT2 | ENSG00000118194.20 | 43098 | -4.967 | 0.0005 | No |