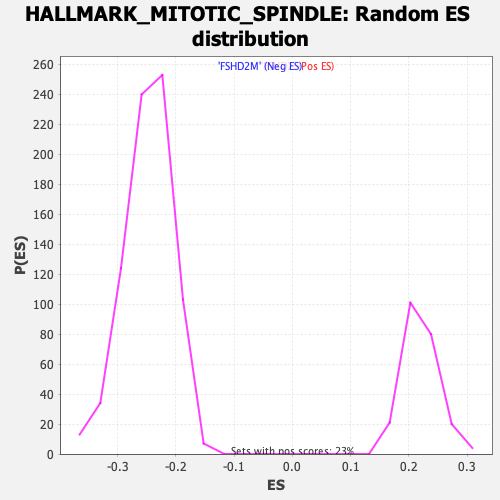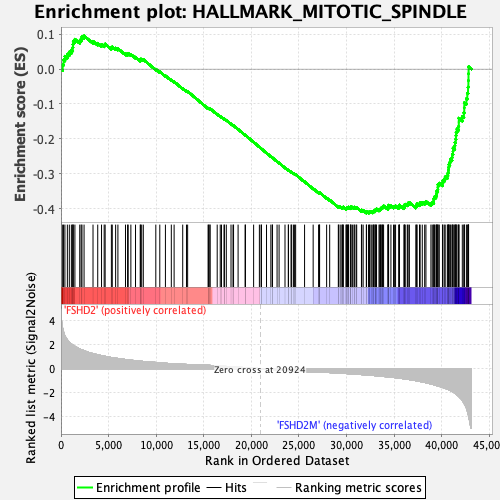
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_MITOTIC_SPINDLE |
| Enrichment Score (ES) | -0.4131707 |
| Normalized Enrichment Score (NES) | -1.6705418 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0030951472 |
| FWER p-Value | 0.04 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDC42EP4 | ENSG00000179604.10 | 177 | 3.355 | 0.0129 | No | ||
| 2 | EPB41 | ENSG00000159023.21 | 269 | 3.077 | 0.0264 | No | ||
| 3 | CLIP2 | ENSG00000106665.15 | 411 | 2.739 | 0.0371 | No | ||
| 4 | ARHGEF3 | ENSG00000163947.11 | 668 | 2.392 | 0.0432 | No | ||
| 5 | MID1 | ENSG00000101871.14 | 874 | 2.189 | 0.0496 | No | ||
| 6 | FSCN1 | ENSG00000075618.18 | 1089 | 2.055 | 0.0550 | No | ||
| 7 | PREX1 | ENSG00000124126.14 | 1235 | 1.973 | 0.0617 | No | ||
| 8 | RAPGEF6 | ENSG00000158987.21 | 1255 | 1.961 | 0.0712 | No | ||
| 9 | EPB41L2 | ENSG00000079819.19 | 1286 | 1.948 | 0.0804 | No | ||
| 10 | RASA2 | ENSG00000155903.12 | 1460 | 1.860 | 0.0858 | No | ||
| 11 | ARAP3 | ENSG00000120318.16 | 1970 | 1.642 | 0.0823 | No | ||
| 12 | MYH10 | ENSG00000133026.12 | 2114 | 1.594 | 0.0871 | No | ||
| 13 | ARHGAP29 | ENSG00000137962.13 | 2202 | 1.565 | 0.0930 | No | ||
| 14 | NCK1 | ENSG00000158092.7 | 2423 | 1.498 | 0.0955 | No | ||
| 15 | DOCK4 | ENSG00000128512.22 | 3363 | 1.252 | 0.0800 | No | ||
| 16 | WASL | ENSG00000106299.8 | 3858 | 1.154 | 0.0743 | No | ||
| 17 | SHROOM1 | ENSG00000164403.14 | 4257 | 1.082 | 0.0705 | No | ||
| 18 | MARCKS | ENSG00000277443.3 | 4535 | 1.038 | 0.0694 | No | ||
| 19 | CTTN | ENSG00000085733.16 | 4636 | 1.024 | 0.0722 | No | ||
| 20 | ARHGEF11 | ENSG00000132694.18 | 5292 | 0.928 | 0.0617 | No | ||
| 21 | ARFIP2 | ENSG00000132254.12 | 5385 | 0.916 | 0.0642 | No | ||
| 22 | TLK1 | ENSG00000198586.14 | 5738 | 0.871 | 0.0604 | No | ||
| 23 | NUMA1 | ENSG00000137497.17 | 5978 | 0.843 | 0.0591 | No | ||
| 24 | ABL1 | ENSG00000097007.19 | 6774 | 0.761 | 0.0445 | No | ||
| 25 | MYO9B | ENSG00000099331.13 | 6995 | 0.739 | 0.0431 | No | ||
| 26 | DOCK2 | ENSG00000134516.17 | 7056 | 0.733 | 0.0454 | No | ||
| 27 | SOS1 | ENSG00000115904.12 | 7339 | 0.706 | 0.0424 | No | ||
| 28 | FARP1 | ENSG00000152767.17 | 7828 | 0.661 | 0.0344 | No | ||
| 29 | ABR | ENSG00000159842.15 | 8332 | 0.617 | 0.0259 | No | ||
| 30 | LATS1 | ENSG00000131023.13 | 8337 | 0.617 | 0.0289 | No | ||
| 31 | RHOF | ENSG00000139725.8 | 8475 | 0.608 | 0.0288 | No | ||
| 32 | WASF1 | ENSG00000112290.13 | 8645 | 0.596 | 0.0279 | No | ||
| 33 | BCR | ENSG00000186716.21 | 9959 | 0.515 | -0.0001 | No | ||
| 34 | UXT | ENSG00000126756.12 | 10378 | 0.486 | -0.0074 | No | ||
| 35 | MYO1E | ENSG00000157483.9 | 10969 | 0.447 | -0.0188 | No | ||
| 36 | KPTN | ENSG00000118162.14 | 11588 | 0.408 | -0.0312 | No | ||
| 37 | AC027237.1 | ENSG00000259191.2 | 11882 | 0.394 | -0.0360 | No | ||
| 38 | LLGL1 | ENSG00000131899.11 | 12786 | 0.356 | -0.0552 | No | ||
| 39 | SYNPO | ENSG00000171992.13 | 13181 | 0.347 | -0.0626 | No | ||
| 40 | CDC42EP2 | ENSG00000149798.5 | 13303 | 0.339 | -0.0637 | No | ||
| 41 | TSC1 | ENSG00000165699.14 | 15451 | 0.278 | -0.1123 | No | ||
| 42 | KIF3B | ENSG00000101350.8 | 15502 | 0.275 | -0.1121 | No | ||
| 43 | PLEKHG2 | ENSG00000090924.15 | 15640 | 0.267 | -0.1139 | No | ||
| 44 | ARHGAP5 | ENSG00000100852.13 | 15655 | 0.266 | -0.1129 | No | ||
| 45 | WASF2 | ENSG00000158195.11 | 16408 | 0.225 | -0.1293 | No | ||
| 46 | TUBGCP3 | ENSG00000126216.15 | 16727 | 0.209 | -0.1356 | No | ||
| 47 | FGD6 | ENSG00000180263.14 | 16859 | 0.204 | -0.1377 | No | ||
| 48 | GEMIN4 | ENSG00000179409.11 | 17129 | 0.191 | -0.1430 | No | ||
| 49 | SMC4 | ENSG00000113810.16 | 17164 | 0.189 | -0.1428 | No | ||
| 50 | CNTRL | ENSG00000119397.16 | 17364 | 0.180 | -0.1465 | No | ||
| 51 | CEP192 | ENSG00000101639.18 | 17859 | 0.154 | -0.1572 | No | ||
| 52 | NF1 | ENSG00000196712.17 | 18068 | 0.143 | -0.1614 | No | ||
| 53 | NEDD9 | ENSG00000111859.17 | 18126 | 0.140 | -0.1620 | No | ||
| 54 | CEP57 | ENSG00000166037.11 | 18603 | 0.119 | -0.1725 | No | ||
| 55 | KATNA1 | ENSG00000186625.14 | 19338 | 0.081 | -0.1892 | No | ||
| 56 | CEP250 | ENSG00000126001.16 | 19378 | 0.078 | -0.1897 | No | ||
| 57 | KIFAP3 | ENSG00000075945.13 | 20234 | 0.036 | -0.2094 | No | ||
| 58 | MARK4 | ENSG00000007047.15 | 20839 | 0.005 | -0.2235 | No | ||
| 59 | TUBGCP6 | ENSG00000128159.12 | 21020 | -0.005 | -0.2276 | No | ||
| 60 | PKD2 | ENSG00000118762.8 | 21057 | -0.007 | -0.2284 | No | ||
| 61 | RANBP9 | ENSG00000010017.13 | 21589 | -0.037 | -0.2406 | No | ||
| 62 | RAB3GAP1 | ENSG00000115839.17 | 22044 | -0.059 | -0.2509 | No | ||
| 63 | NUSAP1 | ENSG00000137804.14 | 22190 | -0.067 | -0.2539 | No | ||
| 64 | KIF3C | ENSG00000084731.15 | 22219 | -0.069 | -0.2542 | No | ||
| 65 | HOOK3 | ENSG00000168172.9 | 22697 | -0.096 | -0.2648 | No | ||
| 66 | FBXO5 | ENSG00000112029.10 | 22909 | -0.108 | -0.2692 | No | ||
| 67 | TUBGCP5 | ENSG00000275835.5 | 23530 | -0.142 | -0.2829 | No | ||
| 68 | DYNLL2 | ENSG00000264364.3 | 23885 | -0.164 | -0.2904 | No | ||
| 69 | NOTCH2 | ENSG00000134250.20 | 23887 | -0.164 | -0.2895 | No | ||
| 70 | RICTOR | ENSG00000164327.13 | 24164 | -0.180 | -0.2951 | No | ||
| 71 | PIF1 | ENSG00000140451.13 | 24228 | -0.184 | -0.2956 | No | ||
| 72 | TUBD1 | ENSG00000108423.15 | 24415 | -0.194 | -0.2989 | No | ||
| 73 | TAOK2 | ENSG00000149930.18 | 24538 | -0.201 | -0.3008 | No | ||
| 74 | PCGF5 | ENSG00000180628.15 | 24648 | -0.208 | -0.3022 | No | ||
| 75 | RHOT2 | ENSG00000140983.14 | 25586 | -0.232 | -0.3229 | No | ||
| 76 | ARHGDIA | ENSG00000141522.12 | 26488 | -0.265 | -0.3425 | No | ||
| 77 | ARHGEF7 | ENSG00000102606.18 | 27057 | -0.277 | -0.3544 | No | ||
| 78 | RALBP1 | ENSG00000017797.13 | 27156 | -0.284 | -0.3552 | No | ||
| 79 | CEP131 | ENSG00000141577.14 | 27164 | -0.285 | -0.3539 | No | ||
| 80 | SMC3 | ENSG00000108055.9 | 27903 | -0.302 | -0.3696 | No | ||
| 81 | CSNK1D | ENSG00000141551.14 | 28224 | -0.325 | -0.3754 | No | ||
| 82 | CYTH2 | ENSG00000105443.15 | 29131 | -0.356 | -0.3947 | No | ||
| 83 | ARFGEF1 | ENSG00000066777.9 | 29168 | -0.358 | -0.3937 | No | ||
| 84 | RABGAP1 | ENSG00000011454.17 | 29328 | -0.364 | -0.3956 | No | ||
| 85 | ARF6 | ENSG00000165527.7 | 29502 | -0.374 | -0.3977 | No | ||
| 86 | AURKA | ENSG00000087586.17 | 29575 | -0.379 | -0.3974 | No | ||
| 87 | ESPL1 | ENSG00000135476.11 | 29621 | -0.383 | -0.3965 | No | ||
| 88 | KIF11 | ENSG00000138160.6 | 29654 | -0.385 | -0.3953 | No | ||
| 89 | NDC80 | ENSG00000080986.13 | 29934 | -0.401 | -0.3998 | No | ||
| 90 | CDC42BPA | ENSG00000143776.18 | 30007 | -0.408 | -0.3994 | No | ||
| 91 | TUBGCP2 | ENSG00000130640.13 | 30039 | -0.410 | -0.3980 | No | ||
| 92 | CCNB2 | ENSG00000157456.8 | 30092 | -0.412 | -0.3972 | No | ||
| 93 | CLIP1 | ENSG00000130779.20 | 30196 | -0.420 | -0.3974 | No | ||
| 94 | BIRC5 | ENSG00000089685.15 | 30216 | -0.422 | -0.3957 | No | ||
| 95 | OPHN1 | ENSG00000079482.13 | 30426 | -0.433 | -0.3984 | No | ||
| 96 | TBCD | ENSG00000141556.21 | 30442 | -0.434 | -0.3965 | No | ||
| 97 | RAPGEF5 | ENSG00000136237.19 | 30447 | -0.435 | -0.3944 | No | ||
| 98 | CAPZB | ENSG00000077549.18 | 30609 | -0.443 | -0.3959 | No | ||
| 99 | MID1IP1 | ENSG00000165175.15 | 30794 | -0.454 | -0.3979 | No | ||
| 100 | ARHGEF12 | ENSG00000196914.9 | 30804 | -0.455 | -0.3958 | No | ||
| 101 | NEK2 | ENSG00000117650.13 | 30994 | -0.459 | -0.3979 | No | ||
| 102 | CDK1 | ENSG00000170312.16 | 31062 | -0.464 | -0.3971 | No | ||
| 103 | HDAC6 | ENSG00000094631.21 | 31561 | -0.493 | -0.4062 | No | ||
| 104 | CNTROB | ENSG00000170037.14 | 31586 | -0.495 | -0.4042 | No | ||
| 105 | KIF20B | ENSG00000138182.14 | 31739 | -0.499 | -0.4052 | No | ||
| 106 | SAC3D1 | ENSG00000168061.15 | 32081 | -0.520 | -0.4105 | Yes | ||
| 107 | CENPE | ENSG00000138778.12 | 32083 | -0.520 | -0.4079 | Yes | ||
| 108 | STAU1 | ENSG00000124214.19 | 32297 | -0.534 | -0.4102 | Yes | ||
| 109 | KNTC1 | ENSG00000184445.12 | 32401 | -0.539 | -0.4098 | Yes | ||
| 110 | MAP3K11 | ENSG00000173327.8 | 32451 | -0.542 | -0.4082 | Yes | ||
| 111 | GSN | ENSG00000148180.19 | 32592 | -0.551 | -0.4087 | Yes | ||
| 112 | RACGAP1 | ENSG00000161800.13 | 32756 | -0.565 | -0.4096 | Yes | ||
| 113 | CDC27 | ENSG00000004897.12 | 32830 | -0.571 | -0.4084 | Yes | ||
| 114 | FLNB | ENSG00000136068.14 | 32870 | -0.574 | -0.4064 | Yes | ||
| 115 | KIF15 | ENSG00000163808.17 | 32992 | -0.583 | -0.4063 | Yes | ||
| 116 | PCM1 | ENSG00000078674.17 | 33008 | -0.584 | -0.4036 | Yes | ||
| 117 | CDC42 | ENSG00000070831.17 | 33086 | -0.589 | -0.4024 | Yes | ||
| 118 | CENPF | ENSG00000117724.13 | 33145 | -0.594 | -0.4008 | Yes | ||
| 119 | KIF2C | ENSG00000142945.13 | 33404 | -0.613 | -0.4037 | Yes | ||
| 120 | TTK | ENSG00000112742.10 | 33483 | -0.619 | -0.4023 | Yes | ||
| 121 | TOP2A | ENSG00000131747.15 | 33556 | -0.624 | -0.4008 | Yes | ||
| 122 | ITSN1 | ENSG00000205726.15 | 33605 | -0.630 | -0.3988 | Yes | ||
| 123 | PRC1 | ENSG00000198901.14 | 33700 | -0.638 | -0.3977 | Yes | ||
| 124 | PCNT | ENSG00000160299.17 | 33778 | -0.643 | -0.3962 | Yes | ||
| 125 | DLGAP5 | ENSG00000126787.13 | 33812 | -0.646 | -0.3937 | Yes | ||
| 126 | NCK2 | ENSG00000071051.14 | 33887 | -0.653 | -0.3921 | Yes | ||
| 127 | CENPJ | ENSG00000151849.15 | 34340 | -0.687 | -0.3992 | Yes | ||
| 128 | KIF22 | ENSG00000079616.12 | 34367 | -0.690 | -0.3963 | Yes | ||
| 129 | TPX2 | ENSG00000088325.16 | 34396 | -0.692 | -0.3934 | Yes | ||
| 130 | KIF4A | ENSG00000090889.12 | 34413 | -0.694 | -0.3903 | Yes | ||
| 131 | ROCK1 | ENSG00000067900.8 | 34657 | -0.715 | -0.3923 | Yes | ||
| 132 | ARL8A | ENSG00000143862.8 | 34931 | -0.736 | -0.3949 | Yes | ||
| 133 | PLK1 | ENSG00000166851.15 | 35046 | -0.745 | -0.3938 | Yes | ||
| 134 | BUB1 | ENSG00000169679.15 | 35145 | -0.755 | -0.3922 | Yes | ||
| 135 | BRCA2 | ENSG00000139618.15 | 35470 | -0.784 | -0.3958 | Yes | ||
| 136 | PAFAH1B1 | ENSG00000007168.13 | 35553 | -0.792 | -0.3937 | Yes | ||
| 137 | SHROOM2 | ENSG00000146950.13 | 35559 | -0.793 | -0.3898 | Yes | ||
| 138 | ATG4B | ENSG00000168397.17 | 36023 | -0.846 | -0.3963 | Yes | ||
| 139 | RFC1 | ENSG00000035928.16 | 36076 | -0.852 | -0.3931 | Yes | ||
| 140 | MAP1S | ENSG00000130479.11 | 36081 | -0.853 | -0.3889 | Yes | ||
| 141 | CKAP5 | ENSG00000175216.15 | 36214 | -0.869 | -0.3876 | Yes | ||
| 142 | LMNB1 | ENSG00000113368.12 | 36411 | -0.890 | -0.3876 | Yes | ||
| 143 | FLNA | ENSG00000196924.17 | 36458 | -0.895 | -0.3841 | Yes | ||
| 144 | LRPPRC | ENSG00000138095.19 | 36597 | -0.914 | -0.3827 | Yes | ||
| 145 | INCENP | ENSG00000149503.13 | 37272 | -1.002 | -0.3933 | Yes | ||
| 146 | CDK5RAP2 | ENSG00000136861.18 | 37353 | -1.014 | -0.3900 | Yes | ||
| 147 | TUBA4A | ENSG00000127824.14 | 37405 | -1.021 | -0.3860 | Yes | ||
| 148 | VCL | ENSG00000035403.17 | 37662 | -1.061 | -0.3866 | Yes | ||
| 149 | ABI1 | ENSG00000136754.17 | 37724 | -1.071 | -0.3826 | Yes | ||
| 150 | ALMS1 | ENSG00000116127.18 | 37951 | -1.109 | -0.3822 | Yes | ||
| 151 | NET1 | ENSG00000173848.19 | 38203 | -1.154 | -0.3822 | Yes | ||
| 152 | KLC1 | ENSG00000126214.21 | 38334 | -1.177 | -0.3793 | Yes | ||
| 153 | CD2AP | ENSG00000198087.7 | 38859 | -1.280 | -0.3850 | Yes | ||
| 154 | NIN | ENSG00000100503.23 | 38999 | -1.310 | -0.3815 | Yes | ||
| 155 | SUN2 | ENSG00000100242.15 | 39161 | -1.344 | -0.3785 | Yes | ||
| 156 | CLASP1 | ENSG00000074054.18 | 39163 | -1.344 | -0.3717 | Yes | ||
| 157 | RASAL2 | ENSG00000075391.16 | 39225 | -1.357 | -0.3662 | Yes | ||
| 158 | SPTBN1 | ENSG00000115306.16 | 39387 | -1.397 | -0.3628 | Yes | ||
| 159 | SSH2 | ENSG00000141298.19 | 39439 | -1.407 | -0.3569 | Yes | ||
| 160 | DYNC1H1 | ENSG00000197102.12 | 39457 | -1.412 | -0.3501 | Yes | ||
| 161 | CEP72 | ENSG00000112877.8 | 39576 | -1.436 | -0.3456 | Yes | ||
| 162 | ANLN | ENSG00000011426.11 | 39589 | -1.439 | -0.3385 | Yes | ||
| 163 | SASS6 | ENSG00000156876.10 | 39594 | -1.441 | -0.3313 | Yes | ||
| 164 | PPP4R2 | ENSG00000163605.15 | 39736 | -1.475 | -0.3271 | Yes | ||
| 165 | SPTAN1 | ENSG00000197694.16 | 40071 | -1.565 | -0.3269 | Yes | ||
| 166 | ARHGAP27 | ENSG00000159314.11 | 40102 | -1.572 | -0.3196 | Yes | ||
| 167 | DLG1 | ENSG00000075711.21 | 40257 | -1.615 | -0.3150 | Yes | ||
| 168 | FGD4 | ENSG00000139132.14 | 40356 | -1.642 | -0.3090 | Yes | ||
| 169 | DST | ENSG00000151914.20 | 40581 | -1.710 | -0.3055 | Yes | ||
| 170 | ARHGAP10 | ENSG00000071205.12 | 40642 | -1.732 | -0.2981 | Yes | ||
| 171 | TRIO | ENSG00000038382.20 | 40687 | -1.744 | -0.2902 | Yes | ||
| 172 | BCAR1 | ENSG00000050820.17 | 40713 | -1.751 | -0.2819 | Yes | ||
| 173 | ARHGEF2 | ENSG00000116584.19 | 40744 | -1.763 | -0.2737 | Yes | ||
| 174 | KATNB1 | ENSG00000140854.13 | 40845 | -1.800 | -0.2669 | Yes | ||
| 175 | SMC1A | ENSG00000072501.17 | 40902 | -1.825 | -0.2589 | Yes | ||
| 176 | YWHAE | ENSG00000108953.17 | 41073 | -1.896 | -0.2532 | Yes | ||
| 177 | ALS2 | ENSG00000003393.15 | 41104 | -1.909 | -0.2442 | Yes | ||
| 178 | APC | ENSG00000134982.17 | 41177 | -1.948 | -0.2360 | Yes | ||
| 179 | ECT2 | ENSG00000114346.13 | 41183 | -1.951 | -0.2262 | Yes | ||
| 180 | PXN | ENSG00000089159.16 | 41365 | -2.042 | -0.2201 | Yes | ||
| 181 | MAPRE1 | ENSG00000101367.9 | 41383 | -2.053 | -0.2100 | Yes | ||
| 182 | AKAP13 | ENSG00000170776.21 | 41458 | -2.106 | -0.2010 | Yes | ||
| 183 | STK38L | ENSG00000211455.8 | 41490 | -2.127 | -0.1910 | Yes | ||
| 184 | BCL2L11 | ENSG00000153094.23 | 41506 | -2.133 | -0.1805 | Yes | ||
| 185 | MYH9 | ENSG00000100345.21 | 41599 | -2.193 | -0.1715 | Yes | ||
| 186 | CCDC88A | ENSG00000115355.17 | 41774 | -2.307 | -0.1638 | Yes | ||
| 187 | RASA1 | ENSG00000145715.15 | 41787 | -2.315 | -0.1523 | Yes | ||
| 188 | ACTN4 | ENSG00000130402.12 | 41788 | -2.315 | -0.1406 | Yes | ||
| 189 | TIAM1 | ENSG00000156299.13 | 42155 | -2.641 | -0.1357 | Yes | ||
| 190 | ARHGAP4 | ENSG00000089820.15 | 42321 | -2.875 | -0.1249 | Yes | ||
| 191 | CDC42EP1 | ENSG00000128283.7 | 42361 | -2.928 | -0.1110 | Yes | ||
| 192 | EZR | ENSG00000092820.18 | 42366 | -2.931 | -0.0962 | Yes | ||
| 193 | KIF1B | ENSG00000054523.17 | 42584 | -3.294 | -0.0845 | Yes | ||
| 194 | BIN1 | ENSG00000136717.15 | 42680 | -3.529 | -0.0688 | Yes | ||
| 195 | KIF5B | ENSG00000170759.11 | 42759 | -3.747 | -0.0515 | Yes | ||
| 196 | PDLIM5 | ENSG00000163110.15 | 42795 | -3.879 | -0.0327 | Yes | ||
| 197 | SORBS2 | ENSG00000154556.18 | 42806 | -3.930 | -0.0129 | Yes | ||
| 198 | PALLD | ENSG00000129116.19 | 42816 | -3.975 | 0.0071 | Yes |