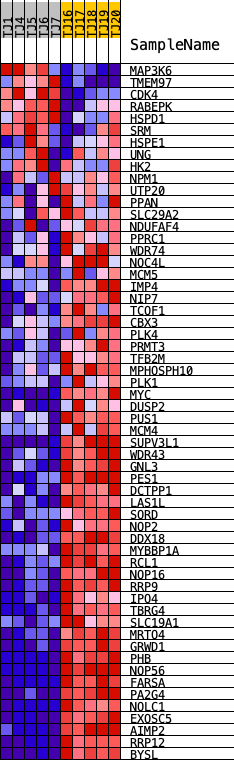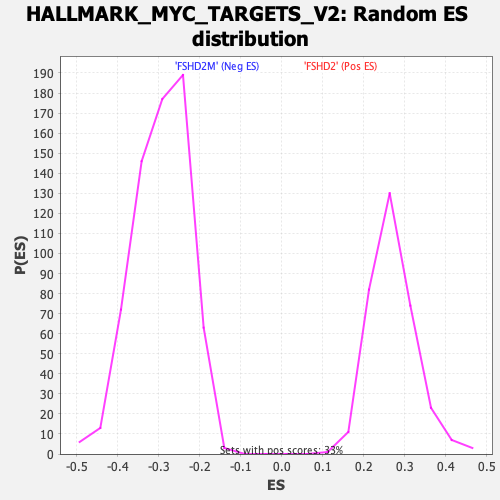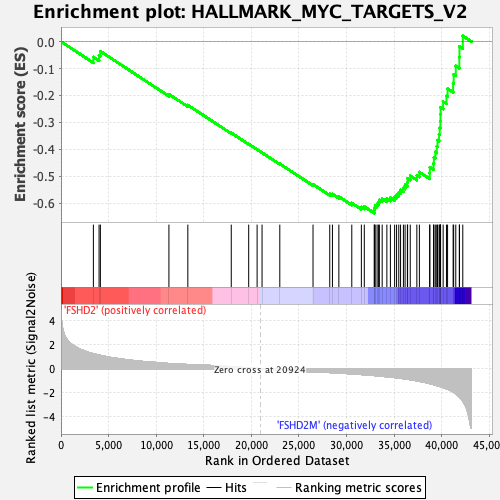
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_MYC_TARGETS_V2 |
| Enrichment Score (ES) | -0.6374056 |
| Normalized Enrichment Score (NES) | -2.1650703 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MAP3K6 | ENSG00000142733.17 | 3401 | 1.245 | -0.0575 | No | ||
| 2 | TMEM97 | ENSG00000109084.14 | 3999 | 1.128 | -0.0518 | No | ||
| 3 | CDK4 | ENSG00000135446.17 | 4137 | 1.105 | -0.0359 | No | ||
| 4 | RABEPK | ENSG00000136933.17 | 11336 | 0.425 | -0.1957 | No | ||
| 5 | HSPD1 | ENSG00000144381.17 | 13323 | 0.338 | -0.2360 | No | ||
| 6 | SRM | ENSG00000116649.10 | 17885 | 0.152 | -0.3393 | No | ||
| 7 | HSPE1 | ENSG00000115541.11 | 19716 | 0.059 | -0.3808 | No | ||
| 8 | UNG | ENSG00000076248.10 | 20600 | 0.017 | -0.4010 | No | ||
| 9 | HK2 | ENSG00000159399.9 | 21120 | -0.012 | -0.4128 | No | ||
| 10 | NPM1 | ENSG00000181163.13 | 22984 | -0.111 | -0.4542 | No | ||
| 11 | UTP20 | ENSG00000120800.5 | 26474 | -0.264 | -0.5306 | No | ||
| 12 | PPAN | ENSG00000130810.20 | 28231 | -0.325 | -0.5658 | No | ||
| 13 | SLC29A2 | ENSG00000174669.12 | 28514 | -0.337 | -0.5665 | No | ||
| 14 | NDUFAF4 | ENSG00000123545.6 | 29188 | -0.359 | -0.5759 | No | ||
| 15 | PPRC1 | ENSG00000148840.11 | 30546 | -0.437 | -0.5999 | No | ||
| 16 | WDR74 | ENSG00000133316.15 | 31552 | -0.493 | -0.6147 | No | ||
| 17 | NOC4L | ENSG00000184967.7 | 31861 | -0.506 | -0.6131 | No | ||
| 18 | MCM5 | ENSG00000100297.16 | 32907 | -0.577 | -0.6274 | Yes | ||
| 19 | IMP4 | ENSG00000136718.9 | 32922 | -0.577 | -0.6178 | Yes | ||
| 20 | NIP7 | ENSG00000132603.15 | 33007 | -0.584 | -0.6096 | Yes | ||
| 21 | TCOF1 | ENSG00000070814.20 | 33202 | -0.599 | -0.6038 | Yes | ||
| 22 | CBX3 | ENSG00000122565.19 | 33342 | -0.609 | -0.5965 | Yes | ||
| 23 | PLK4 | ENSG00000142731.11 | 33445 | -0.616 | -0.5882 | Yes | ||
| 24 | PRMT3 | ENSG00000185238.13 | 33738 | -0.640 | -0.5839 | Yes | ||
| 25 | TFB2M | ENSG00000162851.8 | 34231 | -0.677 | -0.5837 | Yes | ||
| 26 | MPHOSPH10 | ENSG00000124383.9 | 34601 | -0.710 | -0.5800 | Yes | ||
| 27 | PLK1 | ENSG00000166851.15 | 35046 | -0.745 | -0.5774 | Yes | ||
| 28 | MYC | ENSG00000136997.19 | 35251 | -0.763 | -0.5689 | Yes | ||
| 29 | DUSP2 | ENSG00000158050.5 | 35482 | -0.785 | -0.5607 | Yes | ||
| 30 | PUS1 | ENSG00000177192.14 | 35670 | -0.806 | -0.5511 | Yes | ||
| 31 | MCM4 | ENSG00000104738.18 | 35969 | -0.840 | -0.5435 | Yes | ||
| 32 | SUPV3L1 | ENSG00000156502.14 | 36144 | -0.861 | -0.5327 | Yes | ||
| 33 | WDR43 | ENSG00000163811.12 | 36387 | -0.886 | -0.5230 | Yes | ||
| 34 | GNL3 | ENSG00000163938.17 | 36410 | -0.890 | -0.5081 | Yes | ||
| 35 | PES1 | ENSG00000100029.18 | 36687 | -0.928 | -0.4985 | Yes | ||
| 36 | DCTPP1 | ENSG00000179958.9 | 37380 | -1.017 | -0.4970 | Yes | ||
| 37 | LAS1L | ENSG00000001497.16 | 37647 | -1.058 | -0.4849 | Yes | ||
| 38 | SORD | ENSG00000140263.14 | 38729 | -1.249 | -0.4884 | Yes | ||
| 39 | NOP2 | ENSG00000111641.11 | 38743 | -1.252 | -0.4671 | Yes | ||
| 40 | DDX18 | ENSG00000088205.13 | 39130 | -1.337 | -0.4529 | Yes | ||
| 41 | MYBBP1A | ENSG00000132382.14 | 39171 | -1.346 | -0.4306 | Yes | ||
| 42 | RCL1 | ENSG00000120158.12 | 39318 | -1.381 | -0.4101 | Yes | ||
| 43 | NOP16 | ENSG00000048162.20 | 39472 | -1.417 | -0.3892 | Yes | ||
| 44 | RRP9 | ENSG00000114767.7 | 39550 | -1.431 | -0.3663 | Yes | ||
| 45 | IPO4 | ENSG00000196497.17 | 39715 | -1.470 | -0.3447 | Yes | ||
| 46 | TBRG4 | ENSG00000136270.14 | 39777 | -1.485 | -0.3204 | Yes | ||
| 47 | SLC19A1 | ENSG00000173638.19 | 39857 | -1.508 | -0.2962 | Yes | ||
| 48 | MRTO4 | ENSG00000053372.5 | 39866 | -1.511 | -0.2703 | Yes | ||
| 49 | GRWD1 | ENSG00000105447.13 | 39873 | -1.515 | -0.2442 | Yes | ||
| 50 | PHB | ENSG00000167085.11 | 40143 | -1.585 | -0.2231 | Yes | ||
| 51 | NOP56 | ENSG00000101361.17 | 40492 | -1.685 | -0.2020 | Yes | ||
| 52 | FARSA | ENSG00000179115.11 | 40608 | -1.720 | -0.1750 | Yes | ||
| 53 | PA2G4 | ENSG00000170515.14 | 41191 | -1.955 | -0.1547 | Yes | ||
| 54 | NOLC1 | ENSG00000166197.16 | 41247 | -1.977 | -0.1218 | Yes | ||
| 55 | EXOSC5 | ENSG00000077348.9 | 41470 | -2.115 | -0.0904 | Yes | ||
| 56 | AIMP2 | ENSG00000106305.10 | 41843 | -2.363 | -0.0583 | Yes | ||
| 57 | RRP12 | ENSG00000052749.13 | 41858 | -2.372 | -0.0176 | Yes | ||
| 58 | BYSL | ENSG00000112578.10 | 42206 | -2.711 | 0.0212 | Yes |