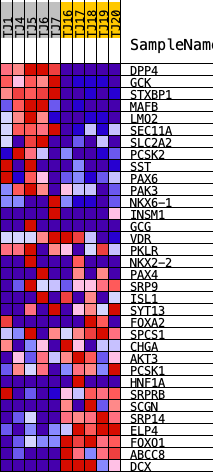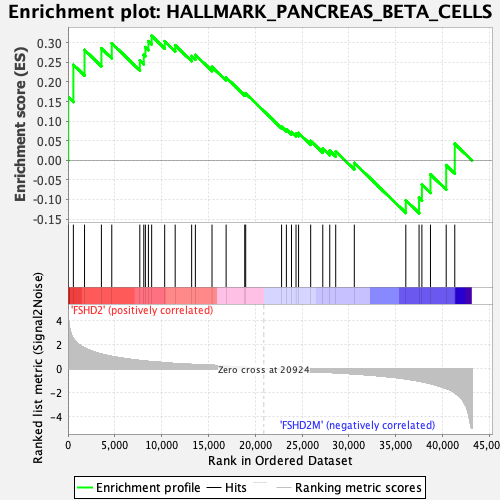
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_PANCREAS_BETA_CELLS |
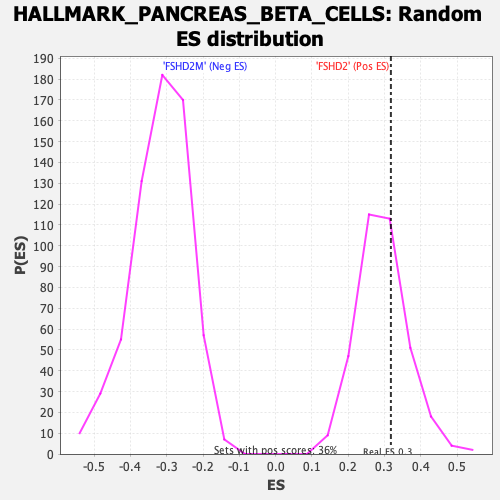
| Enrichment Score (ES) | 0.31778163 |
| Normalized Enrichment Score (NES) | 1.0774586 |
| Nominal p-value | 0.3454039 |
| FDR q-value | 0.32148707 |
| FWER p-Value | 0.981 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DPP4 | ENSG00000197635.10 | 30 | 4.288 | 0.1617 | Yes | ||
| 2 | GCK | ENSG00000106633.17 | 573 | 2.506 | 0.2440 | Yes | ||
| 3 | STXBP1 | ENSG00000136854.21 | 1765 | 1.730 | 0.2818 | Yes | ||
| 4 | MAFB | ENSG00000204103.4 | 3567 | 1.214 | 0.2860 | Yes | ||
| 5 | LMO2 | ENSG00000135363.12 | 4673 | 1.019 | 0.2989 | Yes | ||
| 6 | SEC11A | ENSG00000140612.14 | 7669 | 0.674 | 0.2549 | Yes | ||
| 7 | SLC2A2 | ENSG00000163581.14 | 8091 | 0.639 | 0.2693 | Yes | ||
| 8 | PCSK2 | ENSG00000125851.10 | 8263 | 0.622 | 0.2889 | Yes | ||
| 9 | SST | ENSG00000157005.4 | 8587 | 0.600 | 0.3041 | Yes | ||
| 10 | PAX6 | ENSG00000007372.23 | 8933 | 0.573 | 0.3178 | Yes | ||
| 11 | PAK3 | ENSG00000077264.15 | 10323 | 0.489 | 0.3041 | No | ||
| 12 | NKX6-1 | ENSG00000163623.9 | 11445 | 0.417 | 0.2938 | No | ||
| 13 | INSM1 | ENSG00000173404.5 | 13203 | 0.345 | 0.2661 | No | ||
| 14 | GCG | ENSG00000115263.15 | 13600 | 0.322 | 0.2692 | No | ||
| 15 | VDR | ENSG00000111424.11 | 15378 | 0.281 | 0.2385 | No | ||
| 16 | PKLR | ENSG00000143627.19 | 16884 | 0.203 | 0.2113 | No | ||
| 17 | NKX2-2 | ENSG00000125820.6 | 18871 | 0.105 | 0.1692 | No | ||
| 18 | PAX4 | ENSG00000106331.17 | 18975 | 0.100 | 0.1705 | No | ||
| 19 | SRP9 | ENSG00000143742.13 | 22807 | -0.102 | 0.0855 | No | ||
| 20 | ISL1 | ENSG00000016082.15 | 23319 | -0.131 | 0.0786 | No | ||
| 21 | SYT13 | ENSG00000019505.8 | 23864 | -0.162 | 0.0721 | No | ||
| 22 | FOXA2 | ENSG00000125798.14 | 24349 | -0.191 | 0.0681 | No | ||
| 23 | SPCS1 | ENSG00000114902.14 | 24620 | -0.206 | 0.0696 | No | ||
| 24 | CHGA | ENSG00000100604.13 | 25913 | -0.255 | 0.0493 | No | ||
| 25 | AKT3 | ENSG00000117020.18 | 27205 | -0.288 | 0.0302 | No | ||
| 26 | PCSK1 | ENSG00000175426.11 | 27950 | -0.305 | 0.0245 | No | ||
| 27 | HNF1A | ENSG00000135100.17 | 28586 | -0.341 | 0.0227 | No | ||
| 28 | SRPRB | ENSG00000144867.12 | 30572 | -0.439 | -0.0068 | No | ||
| 29 | SCGN | ENSG00000079689.14 | 36071 | -0.852 | -0.1021 | No | ||
| 30 | SRP14 | ENSG00000140319.11 | 37487 | -1.032 | -0.0959 | No | ||
| 31 | ELP4 | ENSG00000109911.19 | 37788 | -1.081 | -0.0619 | No | ||
| 32 | FOXO1 | ENSG00000150907.9 | 38705 | -1.244 | -0.0361 | No | ||
| 33 | ABCC8 | ENSG00000006071.14 | 40385 | -1.653 | -0.0125 | No | ||
| 34 | DCX | ENSG00000077279.18 | 41299 | -2.006 | 0.0422 | No |