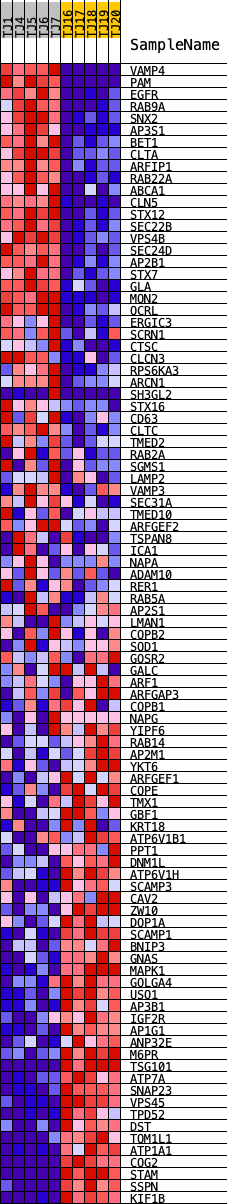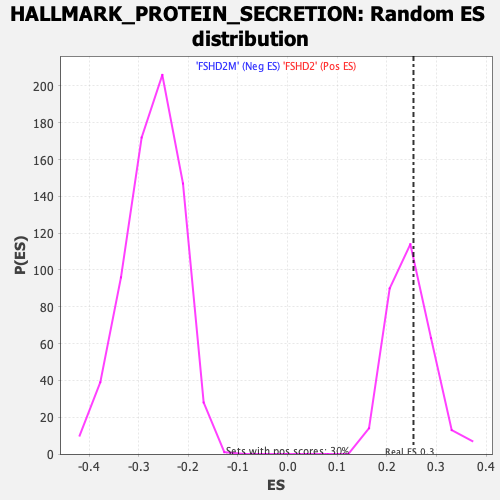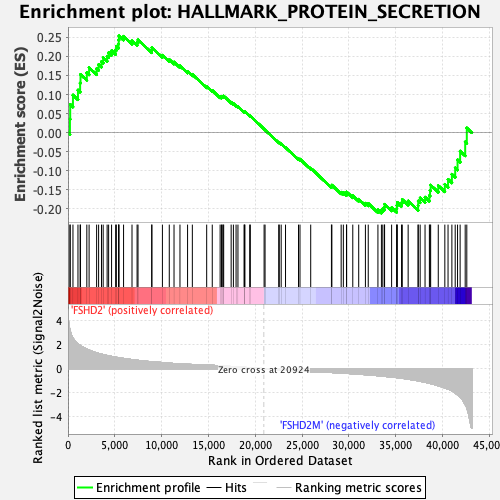
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_PROTEIN_SECRETION |
| Enrichment Score (ES) | 0.25376824 |
| Normalized Enrichment Score (NES) | 1.0293187 |
| Nominal p-value | 0.39202657 |
| FDR q-value | 0.39828065 |
| FWER p-Value | 0.995 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VAMP4 | ENSG00000117533.15 | 205 | 3.285 | 0.0355 | Yes | ||
| 2 | PAM | ENSG00000145730.20 | 235 | 3.193 | 0.0740 | Yes | ||
| 3 | EGFR | ENSG00000146648.18 | 529 | 2.570 | 0.0987 | Yes | ||
| 4 | RAB9A | ENSG00000123595.8 | 1064 | 2.074 | 0.1118 | Yes | ||
| 5 | SNX2 | ENSG00000205302.7 | 1294 | 1.944 | 0.1303 | Yes | ||
| 6 | AP3S1 | ENSG00000177879.16 | 1336 | 1.916 | 0.1528 | Yes | ||
| 7 | BET1 | ENSG00000105829.13 | 2010 | 1.627 | 0.1572 | Yes | ||
| 8 | CLTA | ENSG00000122705.17 | 2253 | 1.549 | 0.1705 | Yes | ||
| 9 | ARFIP1 | ENSG00000164144.16 | 3056 | 1.320 | 0.1681 | Yes | ||
| 10 | RAB22A | ENSG00000124209.4 | 3269 | 1.274 | 0.1788 | Yes | ||
| 11 | ABCA1 | ENSG00000165029.16 | 3576 | 1.212 | 0.1865 | Yes | ||
| 12 | CLN5 | ENSG00000102805.15 | 3756 | 1.173 | 0.1968 | Yes | ||
| 13 | STX12 | ENSG00000117758.14 | 4178 | 1.099 | 0.2005 | Yes | ||
| 14 | SEC22B | ENSG00000265808.4 | 4338 | 1.071 | 0.2099 | Yes | ||
| 15 | VPS4B | ENSG00000119541.10 | 4653 | 1.022 | 0.2151 | Yes | ||
| 16 | SEC24D | ENSG00000150961.15 | 5090 | 0.956 | 0.2167 | Yes | ||
| 17 | AP2B1 | ENSG00000006125.18 | 5163 | 0.946 | 0.2266 | Yes | ||
| 18 | STX7 | ENSG00000079950.14 | 5400 | 0.914 | 0.2324 | Yes | ||
| 19 | GLA | ENSG00000102393.10 | 5414 | 0.912 | 0.2433 | Yes | ||
| 20 | MON2 | ENSG00000061987.16 | 5442 | 0.907 | 0.2538 | Yes | ||
| 21 | OCRL | ENSG00000122126.17 | 5933 | 0.849 | 0.2528 | No | ||
| 22 | ERGIC3 | ENSG00000125991.19 | 6834 | 0.754 | 0.2411 | No | ||
| 23 | SCRN1 | ENSG00000136193.17 | 7377 | 0.703 | 0.2372 | No | ||
| 24 | CTSC | ENSG00000109861.16 | 7476 | 0.693 | 0.2434 | No | ||
| 25 | CLCN3 | ENSG00000109572.14 | 8935 | 0.573 | 0.2165 | No | ||
| 26 | RPS6KA3 | ENSG00000177189.14 | 8961 | 0.570 | 0.2229 | No | ||
| 27 | ARCN1 | ENSG00000095139.14 | 10081 | 0.505 | 0.2031 | No | ||
| 28 | SH3GL2 | ENSG00000107295.10 | 10814 | 0.458 | 0.1917 | No | ||
| 29 | STX16 | ENSG00000124222.22 | 11321 | 0.426 | 0.1852 | No | ||
| 30 | CD63 | ENSG00000135404.11 | 11956 | 0.390 | 0.1752 | No | ||
| 31 | CLTC | ENSG00000141367.12 | 12768 | 0.357 | 0.1608 | No | ||
| 32 | TMED2 | ENSG00000086598.11 | 13277 | 0.341 | 0.1531 | No | ||
| 33 | RAB2A | ENSG00000104388.15 | 14808 | 0.291 | 0.1211 | No | ||
| 34 | SGMS1 | ENSG00000198964.14 | 15404 | 0.280 | 0.1108 | No | ||
| 35 | LAMP2 | ENSG00000005893.15 | 16244 | 0.233 | 0.0941 | No | ||
| 36 | VAMP3 | ENSG00000049245.13 | 16395 | 0.225 | 0.0934 | No | ||
| 37 | SEC31A | ENSG00000138674.17 | 16455 | 0.222 | 0.0947 | No | ||
| 38 | TMED10 | ENSG00000170348.9 | 16564 | 0.216 | 0.0949 | No | ||
| 39 | ARFGEF2 | ENSG00000124198.9 | 16623 | 0.213 | 0.0962 | No | ||
| 40 | TSPAN8 | ENSG00000127324.9 | 17421 | 0.176 | 0.0798 | No | ||
| 41 | ICA1 | ENSG00000003147.19 | 17661 | 0.166 | 0.0763 | No | ||
| 42 | NAPA | ENSG00000105402.8 | 17946 | 0.149 | 0.0715 | No | ||
| 43 | ADAM10 | ENSG00000137845.15 | 18147 | 0.139 | 0.0685 | No | ||
| 44 | RER1 | ENSG00000157916.20 | 18804 | 0.107 | 0.0546 | No | ||
| 45 | RAB5A | ENSG00000144566.11 | 18840 | 0.105 | 0.0551 | No | ||
| 46 | AP2S1 | ENSG00000042753.11 | 18891 | 0.104 | 0.0552 | No | ||
| 47 | LMAN1 | ENSG00000074695.6 | 19403 | 0.077 | 0.0442 | No | ||
| 48 | COPB2 | ENSG00000184432.10 | 19469 | 0.072 | 0.0436 | No | ||
| 49 | SOD1 | ENSG00000142168.14 | 20939 | -0.001 | 0.0095 | No | ||
| 50 | GOSR2 | ENSG00000108433.16 | 21042 | -0.006 | 0.0072 | No | ||
| 51 | GALC | ENSG00000054983.17 | 22503 | -0.084 | -0.0257 | No | ||
| 52 | ARF1 | ENSG00000143761.16 | 22558 | -0.088 | -0.0259 | No | ||
| 53 | ARFGAP3 | ENSG00000242247.11 | 22764 | -0.100 | -0.0294 | No | ||
| 54 | COPB1 | ENSG00000129083.12 | 23227 | -0.126 | -0.0386 | No | ||
| 55 | NAPG | ENSG00000134265.13 | 24614 | -0.205 | -0.0683 | No | ||
| 56 | YIPF6 | ENSG00000181704.12 | 24772 | -0.217 | -0.0693 | No | ||
| 57 | RAB14 | ENSG00000119396.11 | 25916 | -0.255 | -0.0927 | No | ||
| 58 | AP2M1 | ENSG00000161203.13 | 28129 | -0.319 | -0.1402 | No | ||
| 59 | YKT6 | ENSG00000106636.8 | 28177 | -0.321 | -0.1374 | No | ||
| 60 | ARFGEF1 | ENSG00000066777.9 | 29168 | -0.358 | -0.1560 | No | ||
| 61 | COPE | ENSG00000105669.14 | 29401 | -0.370 | -0.1569 | No | ||
| 62 | TMX1 | ENSG00000139921.13 | 29739 | -0.390 | -0.1599 | No | ||
| 63 | GBF1 | ENSG00000107862.4 | 29755 | -0.391 | -0.1555 | No | ||
| 64 | KRT18 | ENSG00000111057.11 | 30409 | -0.432 | -0.1653 | No | ||
| 65 | ATP6V1B1 | ENSG00000116039.13 | 31042 | -0.463 | -0.1743 | No | ||
| 66 | PPT1 | ENSG00000131238.17 | 31757 | -0.501 | -0.1848 | No | ||
| 67 | DNM1L | ENSG00000087470.18 | 32060 | -0.519 | -0.1855 | No | ||
| 68 | ATP6V1H | ENSG00000047249.18 | 33094 | -0.590 | -0.2022 | No | ||
| 69 | SCAMP3 | ENSG00000116521.11 | 33466 | -0.618 | -0.2033 | No | ||
| 70 | CAV2 | ENSG00000105971.15 | 33613 | -0.630 | -0.1989 | No | ||
| 71 | ZW10 | ENSG00000086827.9 | 33786 | -0.643 | -0.1950 | No | ||
| 72 | DOP1A | ENSG00000083097.14 | 33806 | -0.646 | -0.1875 | No | ||
| 73 | SCAMP1 | ENSG00000085365.18 | 34556 | -0.707 | -0.1963 | No | ||
| 74 | BNIP3 | ENSG00000176171.11 | 35097 | -0.750 | -0.1996 | No | ||
| 75 | GNAS | ENSG00000087460.25 | 35113 | -0.751 | -0.1908 | No | ||
| 76 | MAPK1 | ENSG00000100030.14 | 35160 | -0.756 | -0.1825 | No | ||
| 77 | GOLGA4 | ENSG00000144674.16 | 35615 | -0.800 | -0.1833 | No | ||
| 78 | USO1 | ENSG00000138768.14 | 35693 | -0.808 | -0.1752 | No | ||
| 79 | AP3B1 | ENSG00000132842.13 | 36321 | -0.880 | -0.1789 | No | ||
| 80 | IGF2R | ENSG00000197081.14 | 37384 | -1.018 | -0.1911 | No | ||
| 81 | AP1G1 | ENSG00000166747.12 | 37396 | -1.019 | -0.1789 | No | ||
| 82 | ANP32E | ENSG00000143401.15 | 37604 | -1.051 | -0.1708 | No | ||
| 83 | M6PR | ENSG00000003056.8 | 38125 | -1.138 | -0.1689 | No | ||
| 84 | TSG101 | ENSG00000074319.13 | 38589 | -1.221 | -0.1647 | No | ||
| 85 | ATP7A | ENSG00000165240.20 | 38659 | -1.235 | -0.1512 | No | ||
| 86 | SNAP23 | ENSG00000092531.10 | 38725 | -1.249 | -0.1374 | No | ||
| 87 | VPS45 | ENSG00000136631.15 | 39537 | -1.429 | -0.1387 | No | ||
| 88 | TPD52 | ENSG00000076554.15 | 40238 | -1.609 | -0.1352 | No | ||
| 89 | DST | ENSG00000151914.20 | 40581 | -1.710 | -0.1222 | No | ||
| 90 | TOM1L1 | ENSG00000141198.16 | 40991 | -1.863 | -0.1088 | No | ||
| 91 | ATP1A1 | ENSG00000163399.16 | 41345 | -2.032 | -0.0921 | No | ||
| 92 | COG2 | ENSG00000135775.14 | 41596 | -2.191 | -0.0710 | No | ||
| 93 | STAM | ENSG00000136738.15 | 41871 | -2.381 | -0.0482 | No | ||
| 94 | SSPN | ENSG00000123096.11 | 42412 | -2.994 | -0.0240 | No | ||
| 95 | KIF1B | ENSG00000054523.17 | 42584 | -3.294 | 0.0124 | No |