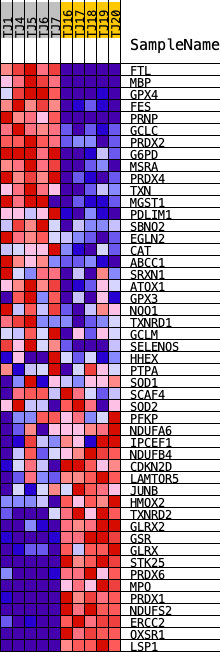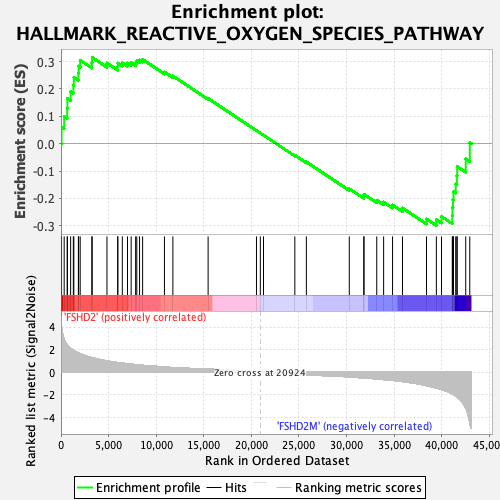
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
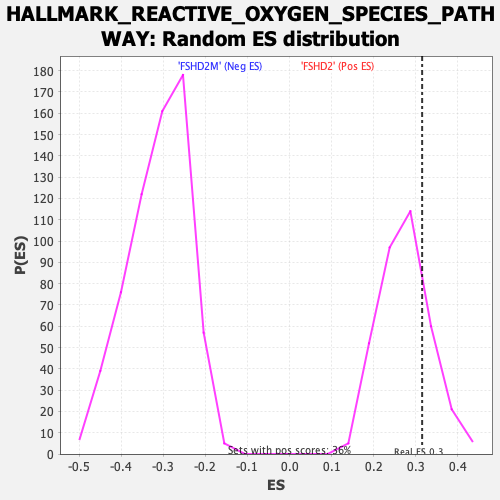
| GeneSet | HALLMARK_REACTIVE_OXYGEN_SPECIES_PATHWAY |
| Enrichment Score (ES) | 0.31496784 |
| Normalized Enrichment Score (NES) | 1.1498357 |
| Nominal p-value | 0.23380281 |
| FDR q-value | 0.196922 |
| FWER p-Value | 0.909 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FTL | ENSG00000087086.15 | 59 | 4.011 | 0.0608 | Yes | ||
| 2 | MBP | ENSG00000197971.15 | 334 | 2.886 | 0.0992 | Yes | ||
| 3 | GPX4 | ENSG00000167468.17 | 652 | 2.411 | 0.1292 | Yes | ||
| 4 | FES | ENSG00000182511.12 | 672 | 2.388 | 0.1657 | Yes | ||
| 5 | PRNP | ENSG00000171867.17 | 1012 | 2.101 | 0.1904 | Yes | ||
| 6 | GCLC | ENSG00000001084.13 | 1297 | 1.942 | 0.2139 | Yes | ||
| 7 | PRDX2 | ENSG00000167815.12 | 1348 | 1.906 | 0.2423 | Yes | ||
| 8 | G6PD | ENSG00000160211.19 | 1825 | 1.702 | 0.2577 | Yes | ||
| 9 | MSRA | ENSG00000175806.15 | 1853 | 1.691 | 0.2833 | Yes | ||
| 10 | PRDX4 | ENSG00000123131.13 | 2016 | 1.623 | 0.3047 | Yes | ||
| 11 | TXN | ENSG00000136810.13 | 3238 | 1.278 | 0.2961 | Yes | ||
| 12 | MGST1 | ENSG00000008394.13 | 3277 | 1.273 | 0.3150 | Yes | ||
| 13 | PDLIM1 | ENSG00000107438.9 | 4829 | 0.993 | 0.2943 | No | ||
| 14 | SBNO2 | ENSG00000064932.16 | 5957 | 0.846 | 0.2813 | No | ||
| 15 | EGLN2 | ENSG00000269858.6 | 5969 | 0.844 | 0.2941 | No | ||
| 16 | CAT | ENSG00000121691.7 | 6441 | 0.793 | 0.2955 | No | ||
| 17 | ABCC1 | ENSG00000103222.19 | 6987 | 0.740 | 0.2943 | No | ||
| 18 | SRXN1 | ENSG00000271303.1 | 7369 | 0.704 | 0.2964 | No | ||
| 19 | ATOX1 | ENSG00000177556.12 | 7840 | 0.661 | 0.2957 | No | ||
| 20 | GPX3 | ENSG00000211445.12 | 7962 | 0.650 | 0.3029 | No | ||
| 21 | NQO1 | ENSG00000181019.13 | 8253 | 0.622 | 0.3059 | No | ||
| 22 | TXNRD1 | ENSG00000198431.16 | 8574 | 0.601 | 0.3077 | No | ||
| 23 | GCLM | ENSG00000023909.10 | 10865 | 0.454 | 0.2616 | No | ||
| 24 | SELENOS | ENSG00000131871.15 | 11752 | 0.400 | 0.2472 | No | ||
| 25 | HHEX | ENSG00000152804.11 | 15454 | 0.278 | 0.1656 | No | ||
| 26 | PTPA | ENSG00000119383.19 | 20534 | 0.020 | 0.0480 | No | ||
| 27 | SOD1 | ENSG00000142168.14 | 20939 | -0.001 | 0.0386 | No | ||
| 28 | SCAF4 | ENSG00000156304.15 | 21273 | -0.020 | 0.0312 | No | ||
| 29 | SOD2 | ENSG00000112096.18 | 24563 | -0.202 | -0.0420 | No | ||
| 30 | PFKP | ENSG00000067057.17 | 25775 | -0.245 | -0.0663 | No | ||
| 31 | NDUFA6 | ENSG00000184983.11 | 30283 | -0.428 | -0.1643 | No | ||
| 32 | IPCEF1 | ENSG00000074706.13 | 31807 | -0.502 | -0.1919 | No | ||
| 33 | NDUFB4 | ENSG00000065518.8 | 31830 | -0.503 | -0.1846 | No | ||
| 34 | CDKN2D | ENSG00000129355.7 | 33168 | -0.595 | -0.2064 | No | ||
| 35 | LAMTOR5 | ENSG00000134248.13 | 33891 | -0.653 | -0.2131 | No | ||
| 36 | JUNB | ENSG00000171223.6 | 34828 | -0.728 | -0.2235 | No | ||
| 37 | HMOX2 | ENSG00000103415.12 | 35871 | -0.827 | -0.2349 | No | ||
| 38 | TXNRD2 | ENSG00000184470.21 | 38395 | -1.187 | -0.2751 | No | ||
| 39 | GLRX2 | ENSG00000023572.9 | 39429 | -1.405 | -0.2773 | No | ||
| 40 | GSR | ENSG00000104687.14 | 39972 | -1.542 | -0.2660 | No | ||
| 41 | GLRX | ENSG00000173221.14 | 41105 | -1.909 | -0.2627 | No | ||
| 42 | STK25 | ENSG00000115694.15 | 41121 | -1.918 | -0.2333 | No | ||
| 43 | PRDX6 | ENSG00000117592.9 | 41173 | -1.946 | -0.2043 | No | ||
| 44 | MPO | ENSG00000005381.8 | 41231 | -1.970 | -0.1751 | No | ||
| 45 | PRDX1 | ENSG00000117450.14 | 41463 | -2.108 | -0.1478 | No | ||
| 46 | NDUFS2 | ENSG00000158864.12 | 41582 | -2.182 | -0.1167 | No | ||
| 47 | ERCC2 | ENSG00000104884.15 | 41619 | -2.203 | -0.0834 | No | ||
| 48 | OXSR1 | ENSG00000172939.9 | 42515 | -3.148 | -0.0554 | No | ||
| 49 | LSP1 | ENSG00000130592.15 | 42941 | -4.475 | 0.0041 | No |