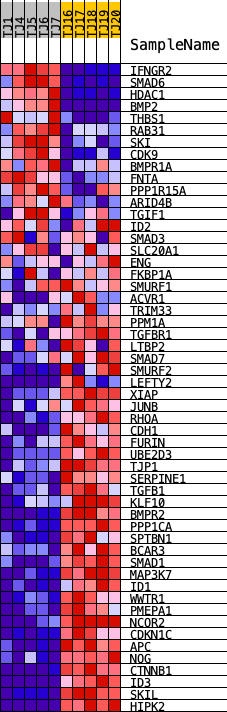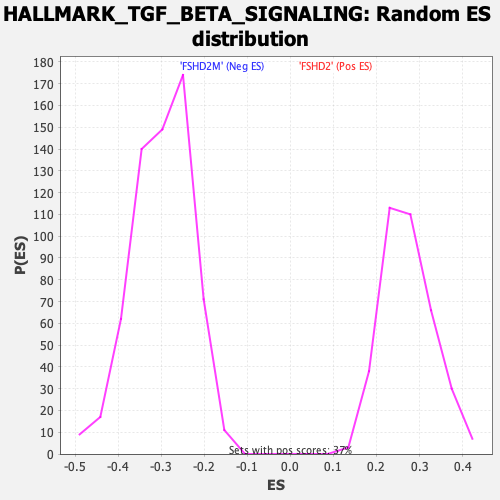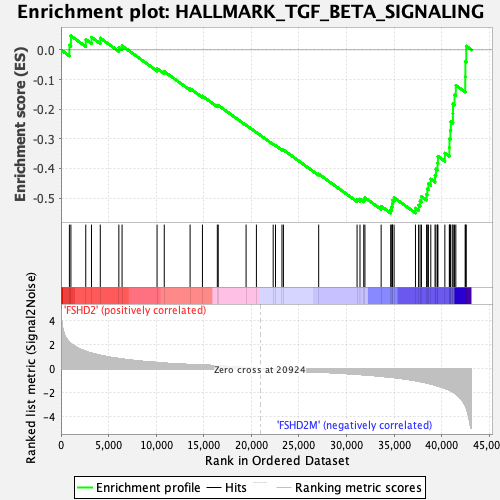
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_TGF_BETA_SIGNALING |
| Enrichment Score (ES) | -0.55062485 |
| Normalized Enrichment Score (NES) | -1.846143 |
| Nominal p-value | 0.0 |
| FDR q-value | 2.1865079E-4 |
| FWER p-Value | 0.002 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IFNGR2 | ENSG00000159128.14 | 882 | 2.184 | 0.0160 | No | ||
| 2 | SMAD6 | ENSG00000137834.15 | 1040 | 2.086 | 0.0472 | No | ||
| 3 | HDAC1 | ENSG00000116478.12 | 2600 | 1.442 | 0.0351 | No | ||
| 4 | BMP2 | ENSG00000125845.7 | 3203 | 1.288 | 0.0427 | No | ||
| 5 | THBS1 | ENSG00000137801.11 | 4127 | 1.107 | 0.0397 | No | ||
| 6 | RAB31 | ENSG00000168461.13 | 6077 | 0.831 | 0.0084 | No | ||
| 7 | SKI | ENSG00000157933.10 | 6415 | 0.796 | 0.0139 | No | ||
| 8 | CDK9 | ENSG00000136807.14 | 10090 | 0.504 | -0.0630 | No | ||
| 9 | BMPR1A | ENSG00000107779.13 | 10847 | 0.455 | -0.0730 | No | ||
| 10 | FNTA | ENSG00000168522.13 | 13560 | 0.323 | -0.1305 | No | ||
| 11 | PPP1R15A | ENSG00000087074.8 | 14858 | 0.289 | -0.1558 | No | ||
| 12 | ARID4B | ENSG00000054267.22 | 16421 | 0.224 | -0.1884 | No | ||
| 13 | TGIF1 | ENSG00000177426.20 | 16517 | 0.219 | -0.1869 | No | ||
| 14 | ID2 | ENSG00000115738.10 | 19442 | 0.074 | -0.2536 | No | ||
| 15 | SMAD3 | ENSG00000166949.16 | 20527 | 0.020 | -0.2784 | No | ||
| 16 | SLC20A1 | ENSG00000144136.11 | 22289 | -0.074 | -0.3181 | No | ||
| 17 | ENG | ENSG00000106991.13 | 22533 | -0.086 | -0.3223 | No | ||
| 18 | FKBP1A | ENSG00000088832.17 | 23211 | -0.124 | -0.3359 | No | ||
| 19 | SMURF1 | ENSG00000198742.9 | 23366 | -0.134 | -0.3373 | No | ||
| 20 | ACVR1 | ENSG00000115170.14 | 27068 | -0.278 | -0.4185 | No | ||
| 21 | TRIM33 | ENSG00000197323.12 | 31101 | -0.468 | -0.5044 | No | ||
| 22 | PPM1A | ENSG00000100614.18 | 31408 | -0.486 | -0.5033 | No | ||
| 23 | TGFBR1 | ENSG00000106799.13 | 31802 | -0.501 | -0.5041 | No | ||
| 24 | LTBP2 | ENSG00000119681.12 | 31926 | -0.510 | -0.4984 | No | ||
| 25 | SMAD7 | ENSG00000101665.9 | 33630 | -0.632 | -0.5274 | No | ||
| 26 | SMURF2 | ENSG00000108854.16 | 34631 | -0.713 | -0.5387 | Yes | ||
| 27 | LEFTY2 | ENSG00000143768.13 | 34745 | -0.720 | -0.5293 | Yes | ||
| 28 | XIAP | ENSG00000101966.12 | 34810 | -0.725 | -0.5187 | Yes | ||
| 29 | JUNB | ENSG00000171223.6 | 34828 | -0.728 | -0.5069 | Yes | ||
| 30 | RHOA | ENSG00000067560.12 | 34992 | -0.741 | -0.4983 | Yes | ||
| 31 | CDH1 | ENSG00000039068.19 | 37235 | -0.997 | -0.5337 | Yes | ||
| 32 | FURIN | ENSG00000140564.12 | 37563 | -1.044 | -0.5239 | Yes | ||
| 33 | UBE2D3 | ENSG00000109332.20 | 37718 | -1.070 | -0.5095 | Yes | ||
| 34 | TJP1 | ENSG00000104067.16 | 37865 | -1.093 | -0.4947 | Yes | ||
| 35 | SERPINE1 | ENSG00000106366.8 | 38403 | -1.189 | -0.4873 | Yes | ||
| 36 | TGFB1 | ENSG00000105329.10 | 38478 | -1.201 | -0.4689 | Yes | ||
| 37 | KLF10 | ENSG00000155090.15 | 38597 | -1.222 | -0.4512 | Yes | ||
| 38 | BMPR2 | ENSG00000204217.15 | 38845 | -1.277 | -0.4356 | Yes | ||
| 39 | PPP1CA | ENSG00000172531.15 | 39292 | -1.374 | -0.4230 | Yes | ||
| 40 | SPTBN1 | ENSG00000115306.16 | 39387 | -1.397 | -0.4019 | Yes | ||
| 41 | BCAR3 | ENSG00000137936.18 | 39551 | -1.431 | -0.3817 | Yes | ||
| 42 | SMAD1 | ENSG00000170365.10 | 39595 | -1.441 | -0.3586 | Yes | ||
| 43 | MAP3K7 | ENSG00000135341.18 | 40323 | -1.633 | -0.3482 | Yes | ||
| 44 | ID1 | ENSG00000125968.9 | 40796 | -1.785 | -0.3294 | Yes | ||
| 45 | WWTR1 | ENSG00000018408.14 | 40806 | -1.787 | -0.2997 | Yes | ||
| 46 | PMEPA1 | ENSG00000124225.16 | 40906 | -1.826 | -0.2715 | Yes | ||
| 47 | NCOR2 | ENSG00000196498.13 | 40948 | -1.842 | -0.2417 | Yes | ||
| 48 | CDKN1C | ENSG00000129757.13 | 41169 | -1.945 | -0.2143 | Yes | ||
| 49 | APC | ENSG00000134982.17 | 41177 | -1.948 | -0.1819 | Yes | ||
| 50 | NOG | ENSG00000183691.5 | 41341 | -2.028 | -0.1518 | Yes | ||
| 51 | CTNNB1 | ENSG00000168036.18 | 41475 | -2.117 | -0.1195 | Yes | ||
| 52 | ID3 | ENSG00000117318.9 | 42462 | -3.078 | -0.0909 | Yes | ||
| 53 | SKIL | ENSG00000136603.14 | 42481 | -3.103 | -0.0395 | Yes | ||
| 54 | HIPK2 | ENSG00000064393.16 | 42572 | -3.249 | 0.0127 | Yes |