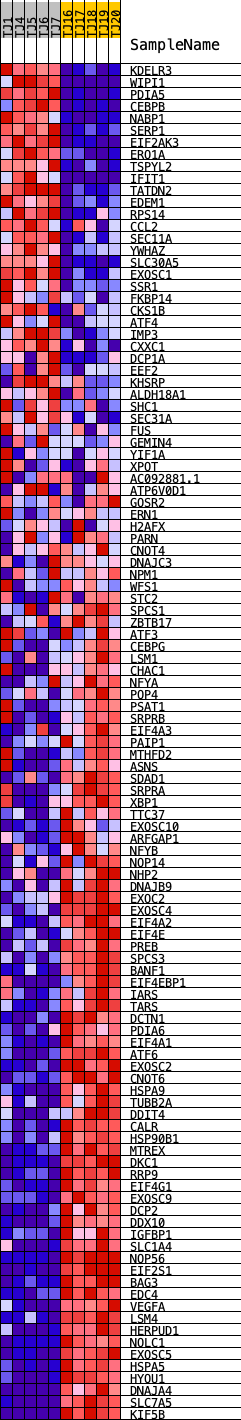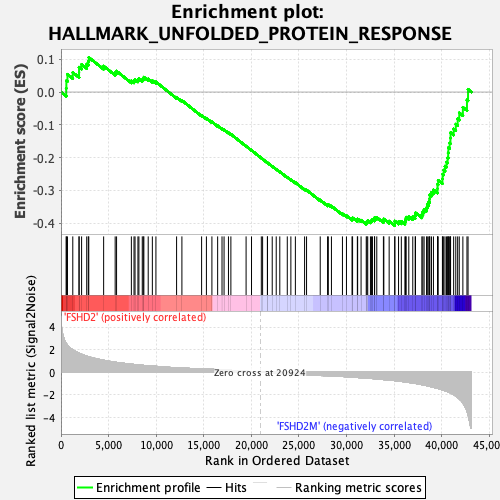
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
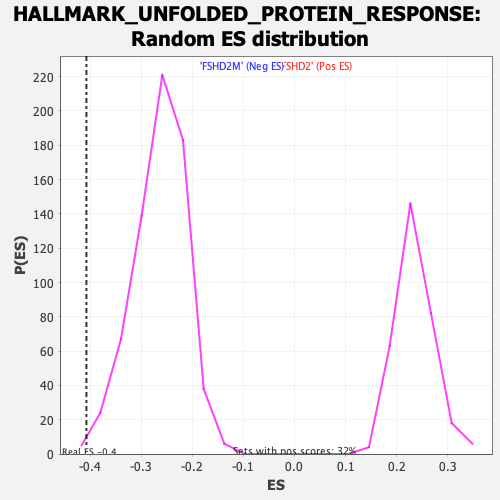
| GeneSet | HALLMARK_UNFOLDED_PROTEIN_RESPONSE |
| Enrichment Score (ES) | -0.4072238 |
| Normalized Enrichment Score (NES) | -1.543736 |
| Nominal p-value | 0.0014684288 |
| FDR q-value | 0.010283726 |
| FWER p-Value | 0.154 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KDELR3 | ENSG00000100196.11 | 547 | 2.546 | 0.0112 | No | ||
| 2 | WIPI1 | ENSG00000070540.13 | 563 | 2.524 | 0.0345 | No | ||
| 3 | PDIA5 | ENSG00000065485.20 | 679 | 2.379 | 0.0541 | No | ||
| 4 | CEBPB | ENSG00000172216.6 | 1238 | 1.972 | 0.0597 | No | ||
| 5 | NABP1 | ENSG00000173559.13 | 1896 | 1.670 | 0.0601 | No | ||
| 6 | SERP1 | ENSG00000120742.11 | 1898 | 1.669 | 0.0757 | No | ||
| 7 | EIF2AK3 | ENSG00000172071.14 | 2164 | 1.578 | 0.0843 | No | ||
| 8 | ERO1A | ENSG00000197930.13 | 2702 | 1.409 | 0.0851 | No | ||
| 9 | TSPYL2 | ENSG00000184205.14 | 2867 | 1.362 | 0.0940 | No | ||
| 10 | IFIT1 | ENSG00000185745.10 | 2918 | 1.351 | 0.1055 | No | ||
| 11 | TATDN2 | ENSG00000157014.11 | 4479 | 1.048 | 0.0791 | No | ||
| 12 | EDEM1 | ENSG00000134109.11 | 5690 | 0.878 | 0.0592 | No | ||
| 13 | RPS14 | ENSG00000164587.13 | 5837 | 0.860 | 0.0639 | No | ||
| 14 | CCL2 | ENSG00000108691.9 | 7391 | 0.701 | 0.0343 | No | ||
| 15 | SEC11A | ENSG00000140612.14 | 7669 | 0.674 | 0.0342 | No | ||
| 16 | YWHAZ | ENSG00000164924.18 | 7777 | 0.666 | 0.0380 | No | ||
| 17 | SLC30A5 | ENSG00000145740.19 | 8082 | 0.640 | 0.0369 | No | ||
| 18 | EXOSC1 | ENSG00000171311.13 | 8183 | 0.630 | 0.0405 | No | ||
| 19 | SSR1 | ENSG00000124783.14 | 8536 | 0.603 | 0.0380 | No | ||
| 20 | FKBP14 | ENSG00000106080.10 | 8640 | 0.596 | 0.0412 | No | ||
| 21 | CKS1B | ENSG00000173207.13 | 8704 | 0.590 | 0.0452 | No | ||
| 22 | ATF4 | ENSG00000128272.14 | 9161 | 0.559 | 0.0399 | No | ||
| 23 | IMP3 | ENSG00000177971.8 | 9596 | 0.530 | 0.0348 | No | ||
| 24 | CXXC1 | ENSG00000154832.14 | 9971 | 0.514 | 0.0309 | No | ||
| 25 | DCP1A | ENSG00000272886.6 | 12142 | 0.380 | -0.0160 | No | ||
| 26 | EEF2 | ENSG00000167658.16 | 12700 | 0.361 | -0.0256 | No | ||
| 27 | KHSRP | ENSG00000088247.17 | 14770 | 0.294 | -0.0709 | No | ||
| 28 | ALDH18A1 | ENSG00000059573.9 | 15274 | 0.285 | -0.0799 | No | ||
| 29 | SHC1 | ENSG00000160691.18 | 15852 | 0.257 | -0.0910 | No | ||
| 30 | SEC31A | ENSG00000138674.17 | 16455 | 0.222 | -0.1029 | No | ||
| 31 | FUS | ENSG00000089280.18 | 16914 | 0.201 | -0.1116 | No | ||
| 32 | GEMIN4 | ENSG00000179409.11 | 17129 | 0.191 | -0.1148 | No | ||
| 33 | YIF1A | ENSG00000174851.16 | 17591 | 0.169 | -0.1239 | No | ||
| 34 | XPOT | ENSG00000184575.12 | 17845 | 0.155 | -0.1284 | No | ||
| 35 | AC092881.1 | ENSG00000279530.2 | 19443 | 0.074 | -0.1648 | No | ||
| 36 | ATP6V0D1 | ENSG00000159720.12 | 20013 | 0.048 | -0.1776 | No | ||
| 37 | GOSR2 | ENSG00000108433.16 | 21042 | -0.006 | -0.2015 | No | ||
| 38 | ERN1 | ENSG00000178607.16 | 21178 | -0.015 | -0.2044 | No | ||
| 39 | H2AFX | ENSG00000188486.3 | 21679 | -0.041 | -0.2157 | No | ||
| 40 | PARN | ENSG00000140694.17 | 21693 | -0.042 | -0.2156 | No | ||
| 41 | CNOT4 | ENSG00000080802.18 | 22188 | -0.067 | -0.2265 | No | ||
| 42 | DNAJC3 | ENSG00000102580.15 | 22608 | -0.091 | -0.2353 | No | ||
| 43 | NPM1 | ENSG00000181163.13 | 22984 | -0.111 | -0.2430 | No | ||
| 44 | WFS1 | ENSG00000109501.13 | 23770 | -0.155 | -0.2598 | No | ||
| 45 | STC2 | ENSG00000113739.10 | 24147 | -0.179 | -0.2669 | No | ||
| 46 | SPCS1 | ENSG00000114902.14 | 24620 | -0.206 | -0.2759 | No | ||
| 47 | ZBTB17 | ENSG00000116809.11 | 25589 | -0.233 | -0.2963 | No | ||
| 48 | ATF3 | ENSG00000162772.17 | 25795 | -0.246 | -0.2987 | No | ||
| 49 | CEBPG | ENSG00000153879.9 | 27230 | -0.290 | -0.3293 | No | ||
| 50 | LSM1 | ENSG00000175324.10 | 28001 | -0.309 | -0.3443 | No | ||
| 51 | CHAC1 | ENSG00000128965.13 | 28080 | -0.316 | -0.3432 | No | ||
| 52 | NFYA | ENSG00000001167.14 | 28408 | -0.331 | -0.3477 | No | ||
| 53 | POP4 | ENSG00000105171.10 | 29564 | -0.379 | -0.3710 | No | ||
| 54 | PSAT1 | ENSG00000135069.14 | 29985 | -0.406 | -0.3769 | No | ||
| 55 | SRPRB | ENSG00000144867.12 | 30572 | -0.439 | -0.3865 | No | ||
| 56 | EIF4A3 | ENSG00000141543.12 | 30630 | -0.445 | -0.3836 | No | ||
| 57 | PAIP1 | ENSG00000172239.14 | 31130 | -0.470 | -0.3908 | No | ||
| 58 | MTHFD2 | ENSG00000065911.12 | 31162 | -0.472 | -0.3871 | No | ||
| 59 | ASNS | ENSG00000070669.17 | 31521 | -0.490 | -0.3908 | No | ||
| 60 | SDAD1 | ENSG00000198301.12 | 32064 | -0.519 | -0.3985 | No | ||
| 61 | SRPRA | ENSG00000182934.12 | 32137 | -0.522 | -0.3953 | No | ||
| 62 | XBP1 | ENSG00000100219.16 | 32221 | -0.529 | -0.3923 | No | ||
| 63 | TTC37 | ENSG00000198677.11 | 32506 | -0.543 | -0.3938 | No | ||
| 64 | EXOSC10 | ENSG00000171824.14 | 32623 | -0.553 | -0.3913 | No | ||
| 65 | ARFGAP1 | ENSG00000101199.13 | 32716 | -0.562 | -0.3882 | No | ||
| 66 | NFYB | ENSG00000120837.8 | 32930 | -0.578 | -0.3877 | No | ||
| 67 | NOP14 | ENSG00000087269.16 | 32946 | -0.579 | -0.3826 | No | ||
| 68 | NHP2 | ENSG00000145912.8 | 33172 | -0.596 | -0.3823 | No | ||
| 69 | DNAJB9 | ENSG00000128590.5 | 33867 | -0.651 | -0.3923 | No | ||
| 70 | EXOC2 | ENSG00000112685.14 | 33913 | -0.655 | -0.3872 | No | ||
| 71 | EXOSC4 | ENSG00000178896.9 | 34471 | -0.700 | -0.3936 | No | ||
| 72 | EIF4A2 | ENSG00000156976.17 | 35058 | -0.747 | -0.4002 | Yes | ||
| 73 | EIF4E | ENSG00000151247.12 | 35069 | -0.748 | -0.3934 | Yes | ||
| 74 | PREB | ENSG00000138073.14 | 35458 | -0.783 | -0.3951 | Yes | ||
| 75 | SPCS3 | ENSG00000129128.13 | 35750 | -0.813 | -0.3942 | Yes | ||
| 76 | BANF1 | ENSG00000175334.8 | 36126 | -0.859 | -0.3949 | Yes | ||
| 77 | EIF4EBP1 | ENSG00000187840.5 | 36191 | -0.867 | -0.3883 | Yes | ||
| 78 | IARS | ENSG00000196305.17 | 36258 | -0.873 | -0.3816 | Yes | ||
| 79 | TARS | ENSG00000113407.13 | 36534 | -0.904 | -0.3795 | Yes | ||
| 80 | DCTN1 | ENSG00000204843.12 | 36952 | -0.960 | -0.3802 | Yes | ||
| 81 | PDIA6 | ENSG00000143870.12 | 37205 | -0.993 | -0.3767 | Yes | ||
| 82 | EIF4A1 | ENSG00000161960.15 | 37239 | -0.997 | -0.3682 | Yes | ||
| 83 | ATF6 | ENSG00000118217.6 | 37906 | -1.101 | -0.3733 | Yes | ||
| 84 | EXOSC2 | ENSG00000130713.16 | 38009 | -1.118 | -0.3652 | Yes | ||
| 85 | CNOT6 | ENSG00000113300.13 | 38142 | -1.143 | -0.3575 | Yes | ||
| 86 | HSPA9 | ENSG00000113013.15 | 38400 | -1.187 | -0.3524 | Yes | ||
| 87 | TUBB2A | ENSG00000137267.6 | 38477 | -1.201 | -0.3429 | Yes | ||
| 88 | DDIT4 | ENSG00000168209.5 | 38633 | -1.230 | -0.3349 | Yes | ||
| 89 | CALR | ENSG00000179218.13 | 38727 | -1.249 | -0.3254 | Yes | ||
| 90 | HSP90B1 | ENSG00000166598.15 | 38735 | -1.251 | -0.3138 | Yes | ||
| 91 | MTREX | ENSG00000039123.16 | 38916 | -1.292 | -0.3059 | Yes | ||
| 92 | DKC1 | ENSG00000130826.18 | 39123 | -1.336 | -0.2981 | Yes | ||
| 93 | RRP9 | ENSG00000114767.7 | 39550 | -1.431 | -0.2946 | Yes | ||
| 94 | EIF4G1 | ENSG00000114867.20 | 39554 | -1.432 | -0.2812 | Yes | ||
| 95 | EXOSC9 | ENSG00000123737.12 | 39622 | -1.447 | -0.2692 | Yes | ||
| 96 | DCP2 | ENSG00000172795.16 | 40065 | -1.562 | -0.2649 | Yes | ||
| 97 | DDX10 | ENSG00000178105.11 | 40086 | -1.568 | -0.2506 | Yes | ||
| 98 | IGFBP1 | ENSG00000146678.10 | 40197 | -1.599 | -0.2382 | Yes | ||
| 99 | SLC1A4 | ENSG00000115902.11 | 40339 | -1.639 | -0.2261 | Yes | ||
| 100 | NOP56 | ENSG00000101361.17 | 40492 | -1.685 | -0.2138 | Yes | ||
| 101 | EIF2S1 | ENSG00000134001.13 | 40613 | -1.722 | -0.2004 | Yes | ||
| 102 | BAG3 | ENSG00000151929.9 | 40672 | -1.741 | -0.1854 | Yes | ||
| 103 | EDC4 | ENSG00000038358.15 | 40693 | -1.747 | -0.1695 | Yes | ||
| 104 | VEGFA | ENSG00000112715.22 | 40824 | -1.792 | -0.1557 | Yes | ||
| 105 | LSM4 | ENSG00000130520.10 | 40907 | -1.826 | -0.1405 | Yes | ||
| 106 | HERPUD1 | ENSG00000051108.15 | 40923 | -1.832 | -0.1237 | Yes | ||
| 107 | NOLC1 | ENSG00000166197.16 | 41247 | -1.977 | -0.1126 | Yes | ||
| 108 | EXOSC5 | ENSG00000077348.9 | 41470 | -2.115 | -0.0979 | Yes | ||
| 109 | HSPA5 | ENSG00000044574.8 | 41668 | -2.232 | -0.0816 | Yes | ||
| 110 | HYOU1 | ENSG00000149428.19 | 41835 | -2.357 | -0.0633 | Yes | ||
| 111 | DNAJA4 | ENSG00000140403.12 | 42215 | -2.729 | -0.0465 | Yes | ||
| 112 | SLC7A5 | ENSG00000103257.9 | 42645 | -3.449 | -0.0242 | Yes | ||
| 113 | KIF5B | ENSG00000170759.11 | 42759 | -3.747 | 0.0084 | Yes |