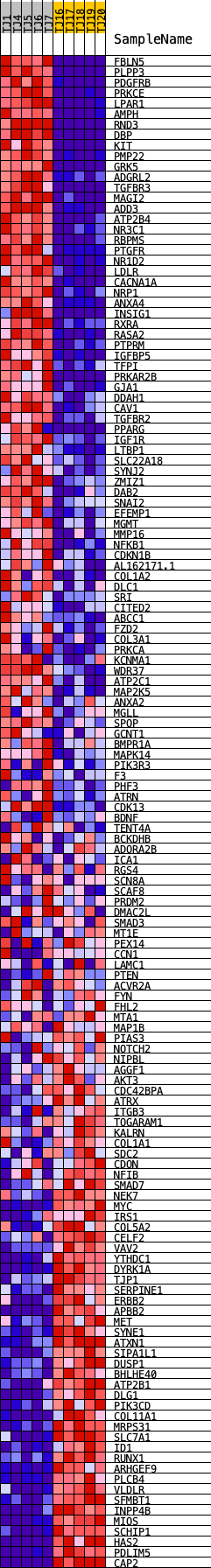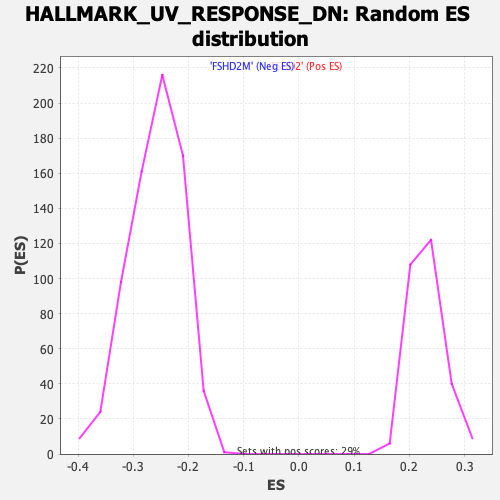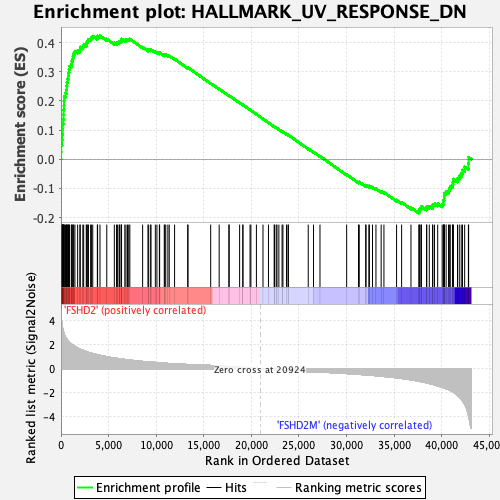
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_UV_RESPONSE_DN |
| Enrichment Score (ES) | 0.42318705 |
| Normalized Enrichment Score (NES) | 1.8281183 |
| Nominal p-value | 0.0 |
| FDR q-value | 6.230769E-4 |
| FWER p-Value | 0.002 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FBLN5 | ENSG00000140092.14 | 11 | 4.579 | 0.0247 | Yes | ||
| 2 | PLPP3 | ENSG00000162407.9 | 18 | 4.425 | 0.0486 | Yes | ||
| 3 | PDGFRB | ENSG00000113721.14 | 106 | 3.655 | 0.0665 | Yes | ||
| 4 | PRKCE | ENSG00000171132.14 | 128 | 3.568 | 0.0854 | Yes | ||
| 5 | LPAR1 | ENSG00000198121.13 | 173 | 3.367 | 0.1027 | Yes | ||
| 6 | AMPH | ENSG00000078053.17 | 178 | 3.355 | 0.1209 | Yes | ||
| 7 | RND3 | ENSG00000115963.13 | 242 | 3.154 | 0.1366 | Yes | ||
| 8 | DBP | ENSG00000105516.11 | 266 | 3.086 | 0.1529 | Yes | ||
| 9 | KIT | ENSG00000157404.15 | 270 | 3.076 | 0.1695 | Yes | ||
| 10 | PMP22 | ENSG00000109099.15 | 310 | 2.946 | 0.1847 | Yes | ||
| 11 | GRK5 | ENSG00000198873.12 | 321 | 2.912 | 0.2003 | Yes | ||
| 12 | ADGRL2 | ENSG00000117114.19 | 339 | 2.881 | 0.2156 | Yes | ||
| 13 | TGFBR3 | ENSG00000069702.11 | 474 | 2.641 | 0.2268 | Yes | ||
| 14 | MAGI2 | ENSG00000187391.21 | 570 | 2.509 | 0.2383 | Yes | ||
| 15 | ADD3 | ENSG00000148700.15 | 598 | 2.472 | 0.2511 | Yes | ||
| 16 | ATP2B4 | ENSG00000058668.14 | 627 | 2.432 | 0.2637 | Yes | ||
| 17 | NR3C1 | ENSG00000113580.15 | 693 | 2.369 | 0.2751 | Yes | ||
| 18 | RBPMS | ENSG00000157110.16 | 766 | 2.294 | 0.2859 | Yes | ||
| 19 | PTGFR | ENSG00000122420.10 | 801 | 2.260 | 0.2974 | Yes | ||
| 20 | NR1D2 | ENSG00000174738.13 | 848 | 2.218 | 0.3084 | Yes | ||
| 21 | LDLR | ENSG00000130164.13 | 900 | 2.174 | 0.3190 | Yes | ||
| 22 | CACNA1A | ENSG00000141837.20 | 1085 | 2.059 | 0.3260 | Yes | ||
| 23 | NRP1 | ENSG00000099250.18 | 1128 | 2.034 | 0.3361 | Yes | ||
| 24 | ANXA4 | ENSG00000196975.16 | 1230 | 1.975 | 0.3445 | Yes | ||
| 25 | INSIG1 | ENSG00000186480.13 | 1265 | 1.957 | 0.3543 | Yes | ||
| 26 | RXRA | ENSG00000186350.12 | 1332 | 1.918 | 0.3632 | Yes | ||
| 27 | RASA2 | ENSG00000155903.12 | 1460 | 1.860 | 0.3704 | Yes | ||
| 28 | PTPRM | ENSG00000173482.16 | 1747 | 1.738 | 0.3732 | Yes | ||
| 29 | IGFBP5 | ENSG00000115461.5 | 1987 | 1.637 | 0.3766 | Yes | ||
| 30 | TFPI | ENSG00000003436.16 | 2020 | 1.622 | 0.3846 | Yes | ||
| 31 | PRKAR2B | ENSG00000005249.13 | 2294 | 1.537 | 0.3867 | Yes | ||
| 32 | GJA1 | ENSG00000152661.9 | 2383 | 1.509 | 0.3928 | Yes | ||
| 33 | DDAH1 | ENSG00000153904.20 | 2653 | 1.424 | 0.3943 | Yes | ||
| 34 | CAV1 | ENSG00000105974.12 | 2696 | 1.410 | 0.4010 | Yes | ||
| 35 | TGFBR2 | ENSG00000163513.18 | 2797 | 1.379 | 0.4062 | Yes | ||
| 36 | PPARG | ENSG00000132170.21 | 2883 | 1.360 | 0.4116 | Yes | ||
| 37 | IGF1R | ENSG00000140443.15 | 3111 | 1.309 | 0.4135 | Yes | ||
| 38 | LTBP1 | ENSG00000049323.16 | 3213 | 1.284 | 0.4181 | Yes | ||
| 39 | SLC22A18 | ENSG00000110628.15 | 3336 | 1.257 | 0.4221 | Yes | ||
| 40 | SYNJ2 | ENSG00000078269.15 | 3818 | 1.161 | 0.4172 | Yes | ||
| 41 | ZMIZ1 | ENSG00000108175.17 | 3844 | 1.156 | 0.4229 | Yes | ||
| 42 | DAB2 | ENSG00000153071.15 | 4095 | 1.112 | 0.4232 | Yes | ||
| 43 | SNAI2 | ENSG00000019549.13 | 4808 | 0.996 | 0.4120 | No | ||
| 44 | EFEMP1 | ENSG00000115380.20 | 5601 | 0.888 | 0.3984 | No | ||
| 45 | MGMT | ENSG00000170430.10 | 5829 | 0.861 | 0.3978 | No | ||
| 46 | MMP16 | ENSG00000156103.16 | 5924 | 0.851 | 0.4003 | No | ||
| 47 | NFKB1 | ENSG00000109320.13 | 6111 | 0.828 | 0.4005 | No | ||
| 48 | CDKN1B | ENSG00000111276.11 | 6128 | 0.825 | 0.4046 | No | ||
| 49 | AL162171.1 | ENSG00000258789.1 | 6323 | 0.804 | 0.4045 | No | ||
| 50 | COL1A2 | ENSG00000164692.17 | 6324 | 0.804 | 0.4088 | No | ||
| 51 | DLC1 | ENSG00000164741.15 | 6359 | 0.801 | 0.4124 | No | ||
| 52 | SRI | ENSG00000075142.13 | 6706 | 0.767 | 0.4085 | No | ||
| 53 | CITED2 | ENSG00000164442.10 | 6826 | 0.755 | 0.4099 | No | ||
| 54 | ABCC1 | ENSG00000103222.19 | 6987 | 0.740 | 0.4102 | No | ||
| 55 | FZD2 | ENSG00000180340.7 | 7116 | 0.726 | 0.4112 | No | ||
| 56 | COL3A1 | ENSG00000168542.14 | 7229 | 0.715 | 0.4124 | No | ||
| 57 | PRKCA | ENSG00000154229.12 | 8575 | 0.601 | 0.3844 | No | ||
| 58 | KCNMA1 | ENSG00000156113.23 | 9150 | 0.560 | 0.3741 | No | ||
| 59 | WDR37 | ENSG00000047056.16 | 9166 | 0.558 | 0.3768 | No | ||
| 60 | ATP2C1 | ENSG00000017260.19 | 9381 | 0.545 | 0.3748 | No | ||
| 61 | MAP2K5 | ENSG00000137764.20 | 9457 | 0.542 | 0.3760 | No | ||
| 62 | ANXA2 | ENSG00000182718.16 | 9909 | 0.516 | 0.3683 | No | ||
| 63 | MGLL | ENSG00000074416.14 | 10076 | 0.505 | 0.3672 | No | ||
| 64 | SPOP | ENSG00000121067.19 | 10334 | 0.489 | 0.3639 | No | ||
| 65 | GCNT1 | ENSG00000187210.14 | 10361 | 0.487 | 0.3659 | No | ||
| 66 | BMPR1A | ENSG00000107779.13 | 10847 | 0.455 | 0.3571 | No | ||
| 67 | MAPK14 | ENSG00000112062.10 | 10900 | 0.451 | 0.3584 | No | ||
| 68 | PIK3R3 | ENSG00000117461.15 | 11004 | 0.444 | 0.3584 | No | ||
| 69 | F3 | ENSG00000117525.14 | 11227 | 0.431 | 0.3556 | No | ||
| 70 | PHF3 | ENSG00000118482.11 | 11370 | 0.423 | 0.3546 | No | ||
| 71 | ATRN | ENSG00000088812.18 | 11931 | 0.391 | 0.3437 | No | ||
| 72 | CDK13 | ENSG00000065883.16 | 13312 | 0.339 | 0.3134 | No | ||
| 73 | BDNF | ENSG00000176697.19 | 13340 | 0.337 | 0.3146 | No | ||
| 74 | TENT4A | ENSG00000112941.14 | 15724 | 0.264 | 0.2606 | No | ||
| 75 | BCKDHB | ENSG00000083123.15 | 16615 | 0.213 | 0.2410 | No | ||
| 76 | ADORA2B | ENSG00000170425.3 | 17637 | 0.167 | 0.2182 | No | ||
| 77 | ICA1 | ENSG00000003147.19 | 17661 | 0.166 | 0.2186 | No | ||
| 78 | RGS4 | ENSG00000117152.13 | 18763 | 0.109 | 0.1935 | No | ||
| 79 | SCN8A | ENSG00000196876.17 | 19100 | 0.095 | 0.1862 | No | ||
| 80 | SCAF8 | ENSG00000213079.9 | 19111 | 0.094 | 0.1865 | No | ||
| 81 | PRDM2 | ENSG00000116731.22 | 19858 | 0.053 | 0.1694 | No | ||
| 82 | DMAC2L | ENSG00000125375.15 | 19947 | 0.050 | 0.1677 | No | ||
| 83 | SMAD3 | ENSG00000166949.16 | 20527 | 0.020 | 0.1543 | No | ||
| 84 | MT1E | ENSG00000169715.15 | 21218 | -0.017 | 0.1383 | No | ||
| 85 | PEX14 | ENSG00000142655.13 | 21793 | -0.047 | 0.1252 | No | ||
| 86 | CCN1 | ENSG00000142871.17 | 22390 | -0.079 | 0.1118 | No | ||
| 87 | LAMC1 | ENSG00000135862.6 | 22432 | -0.081 | 0.1113 | No | ||
| 88 | PTEN | ENSG00000171862.11 | 22605 | -0.091 | 0.1078 | No | ||
| 89 | ACVR2A | ENSG00000121989.15 | 22689 | -0.096 | 0.1064 | No | ||
| 90 | FYN | ENSG00000010810.17 | 22887 | -0.107 | 0.1024 | No | ||
| 91 | FHL2 | ENSG00000115641.19 | 23221 | -0.125 | 0.0953 | No | ||
| 92 | MTA1 | ENSG00000182979.18 | 23313 | -0.130 | 0.0939 | No | ||
| 93 | MAP1B | ENSG00000131711.15 | 23686 | -0.151 | 0.0860 | No | ||
| 94 | PIAS3 | ENSG00000131788.16 | 23847 | -0.161 | 0.0832 | No | ||
| 95 | NOTCH2 | ENSG00000134250.20 | 23887 | -0.164 | 0.0832 | No | ||
| 96 | NIPBL | ENSG00000164190.19 | 25971 | -0.258 | 0.0361 | No | ||
| 97 | AGGF1 | ENSG00000164252.13 | 26526 | -0.268 | 0.0247 | No | ||
| 98 | AKT3 | ENSG00000117020.18 | 27205 | -0.288 | 0.0105 | No | ||
| 99 | CDC42BPA | ENSG00000143776.18 | 30007 | -0.408 | -0.0525 | No | ||
| 100 | ATRX | ENSG00000085224.22 | 31252 | -0.474 | -0.0788 | No | ||
| 101 | ITGB3 | ENSG00000259207.7 | 31330 | -0.481 | -0.0780 | No | ||
| 102 | TOGARAM1 | ENSG00000198718.13 | 32006 | -0.514 | -0.0909 | No | ||
| 103 | KALRN | ENSG00000160145.15 | 32042 | -0.517 | -0.0889 | No | ||
| 104 | COL1A1 | ENSG00000108821.13 | 32343 | -0.536 | -0.0930 | No | ||
| 105 | SDC2 | ENSG00000169439.12 | 32389 | -0.538 | -0.0911 | No | ||
| 106 | CDON | ENSG00000064309.14 | 32722 | -0.562 | -0.0958 | No | ||
| 107 | NFIB | ENSG00000147862.17 | 33084 | -0.589 | -0.1010 | No | ||
| 108 | SMAD7 | ENSG00000101665.9 | 33630 | -0.632 | -0.1102 | No | ||
| 109 | NEK7 | ENSG00000151414.15 | 33923 | -0.656 | -0.1134 | No | ||
| 110 | MYC | ENSG00000136997.19 | 35251 | -0.763 | -0.1402 | No | ||
| 111 | IRS1 | ENSG00000169047.5 | 35781 | -0.816 | -0.1480 | No | ||
| 112 | COL5A2 | ENSG00000204262.14 | 36764 | -0.936 | -0.1658 | No | ||
| 113 | CELF2 | ENSG00000048740.18 | 37582 | -1.047 | -0.1791 | No | ||
| 114 | VAV2 | ENSG00000160293.17 | 37686 | -1.065 | -0.1757 | No | ||
| 115 | YTHDC1 | ENSG00000083896.12 | 37689 | -1.065 | -0.1699 | No | ||
| 116 | DYRK1A | ENSG00000157540.21 | 37834 | -1.089 | -0.1674 | No | ||
| 117 | TJP1 | ENSG00000104067.16 | 37865 | -1.093 | -0.1621 | No | ||
| 118 | SERPINE1 | ENSG00000106366.8 | 38403 | -1.189 | -0.1681 | No | ||
| 119 | ERBB2 | ENSG00000141736.13 | 38420 | -1.191 | -0.1620 | No | ||
| 120 | APBB2 | ENSG00000163697.17 | 38687 | -1.240 | -0.1615 | No | ||
| 121 | MET | ENSG00000105976.15 | 39028 | -1.316 | -0.1622 | No | ||
| 122 | SYNE1 | ENSG00000131018.24 | 39062 | -1.323 | -0.1558 | No | ||
| 123 | ATXN1 | ENSG00000124788.18 | 39238 | -1.362 | -0.1524 | No | ||
| 124 | SIPA1L1 | ENSG00000197555.9 | 39570 | -1.435 | -0.1523 | No | ||
| 125 | DUSP1 | ENSG00000120129.6 | 40039 | -1.558 | -0.1547 | No | ||
| 126 | BHLHE40 | ENSG00000134107.5 | 40173 | -1.593 | -0.1491 | No | ||
| 127 | ATP2B1 | ENSG00000070961.15 | 40174 | -1.593 | -0.1405 | No | ||
| 128 | DLG1 | ENSG00000075711.21 | 40257 | -1.615 | -0.1336 | No | ||
| 129 | PIK3CD | ENSG00000171608.16 | 40260 | -1.616 | -0.1248 | No | ||
| 130 | COL11A1 | ENSG00000060718.22 | 40275 | -1.618 | -0.1164 | No | ||
| 131 | MRPS31 | ENSG00000102738.8 | 40457 | -1.673 | -0.1115 | No | ||
| 132 | SLC7A1 | ENSG00000139514.13 | 40707 | -1.750 | -0.1077 | No | ||
| 133 | ID1 | ENSG00000125968.9 | 40796 | -1.785 | -0.1001 | No | ||
| 134 | RUNX1 | ENSG00000159216.18 | 40891 | -1.819 | -0.0924 | No | ||
| 135 | ARHGEF9 | ENSG00000131089.16 | 41120 | -1.918 | -0.0872 | No | ||
| 136 | PLCB4 | ENSG00000101333.16 | 41175 | -1.947 | -0.0779 | No | ||
| 137 | VLDLR | ENSG00000147852.16 | 41219 | -1.967 | -0.0682 | No | ||
| 138 | SFMBT1 | ENSG00000163935.14 | 41651 | -2.222 | -0.0661 | No | ||
| 139 | INPP4B | ENSG00000109452.12 | 41836 | -2.357 | -0.0576 | No | ||
| 140 | MIOS | ENSG00000164654.16 | 42040 | -2.520 | -0.0486 | No | ||
| 141 | SCHIP1 | ENSG00000151967.18 | 42173 | -2.665 | -0.0371 | No | ||
| 142 | HAS2 | ENSG00000170961.7 | 42403 | -2.985 | -0.0262 | No | ||
| 143 | PDLIM5 | ENSG00000163110.15 | 42795 | -3.879 | -0.0142 | No | ||
| 144 | CAP2 | ENSG00000112186.12 | 42820 | -3.986 | 0.0070 | No |