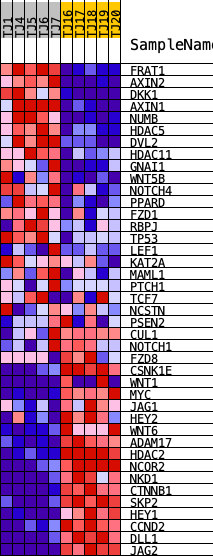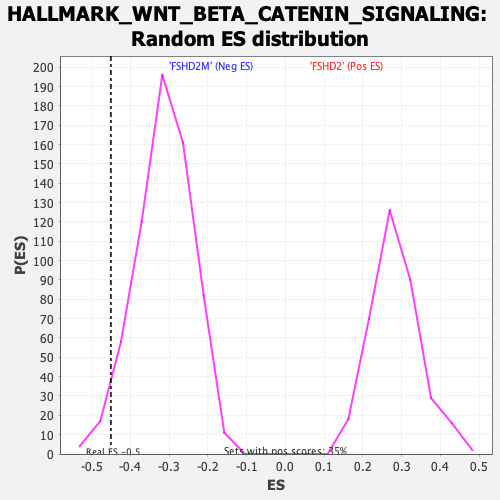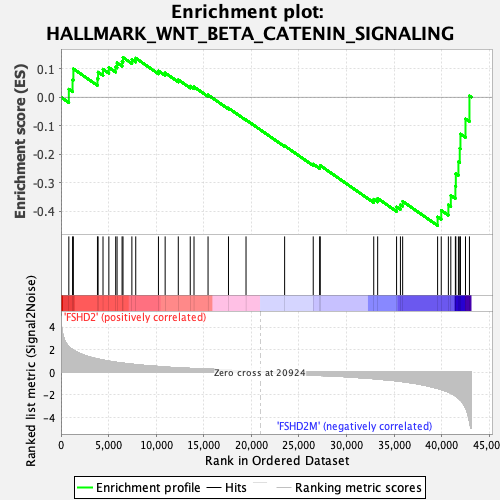
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | HALLMARK_WNT_BETA_CATENIN_SIGNALING |
| Enrichment Score (ES) | -0.45054495 |
| Normalized Enrichment Score (NES) | -1.4324806 |
| Nominal p-value | 0.033898305 |
| FDR q-value | 0.032207996 |
| FWER p-Value | 0.438 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FRAT1 | ENSG00000165879.9 | 822 | 2.241 | 0.0281 | No | ||
| 2 | AXIN2 | ENSG00000168646.13 | 1214 | 1.983 | 0.0608 | No | ||
| 3 | DKK1 | ENSG00000107984.10 | 1299 | 1.941 | 0.0997 | No | ||
| 4 | AXIN1 | ENSG00000103126.15 | 3830 | 1.159 | 0.0653 | No | ||
| 5 | NUMB | ENSG00000133961.20 | 3896 | 1.146 | 0.0880 | No | ||
| 6 | HDAC5 | ENSG00000108840.15 | 4413 | 1.060 | 0.0983 | No | ||
| 7 | DVL2 | ENSG00000004975.12 | 5027 | 0.964 | 0.1043 | No | ||
| 8 | HDAC11 | ENSG00000163517.15 | 5745 | 0.870 | 0.1060 | No | ||
| 9 | GNAI1 | ENSG00000127955.17 | 5889 | 0.854 | 0.1207 | No | ||
| 10 | WNT5B | ENSG00000111186.13 | 6416 | 0.796 | 0.1252 | No | ||
| 11 | NOTCH4 | ENSG00000204301.6 | 6513 | 0.787 | 0.1396 | No | ||
| 12 | PPARD | ENSG00000112033.14 | 7445 | 0.695 | 0.1326 | No | ||
| 13 | FZD1 | ENSG00000157240.4 | 7851 | 0.659 | 0.1371 | No | ||
| 14 | RBPJ | ENSG00000168214.20 | 10236 | 0.495 | 0.0921 | No | ||
| 15 | TP53 | ENSG00000141510.17 | 10940 | 0.449 | 0.0853 | No | ||
| 16 | LEF1 | ENSG00000138795.10 | 12321 | 0.374 | 0.0611 | No | ||
| 17 | KAT2A | ENSG00000108773.11 | 13577 | 0.322 | 0.0388 | No | ||
| 18 | MAML1 | ENSG00000161021.13 | 13972 | 0.319 | 0.0363 | No | ||
| 19 | PTCH1 | ENSG00000185920.15 | 15447 | 0.278 | 0.0080 | No | ||
| 20 | TCF7 | ENSG00000081059.20 | 17593 | 0.169 | -0.0383 | No | ||
| 21 | NCSTN | ENSG00000162736.17 | 19436 | 0.074 | -0.0795 | No | ||
| 22 | PSEN2 | ENSG00000143801.17 | 23491 | -0.140 | -0.1706 | No | ||
| 23 | CUL1 | ENSG00000055130.17 | 26496 | -0.266 | -0.2348 | No | ||
| 24 | NOTCH1 | ENSG00000148400.12 | 27170 | -0.285 | -0.2444 | No | ||
| 25 | FZD8 | ENSG00000177283.7 | 27233 | -0.290 | -0.2397 | No | ||
| 26 | CSNK1E | ENSG00000213923.12 | 32851 | -0.572 | -0.3580 | No | ||
| 27 | WNT1 | ENSG00000125084.12 | 33264 | -0.602 | -0.3549 | No | ||
| 28 | MYC | ENSG00000136997.19 | 35251 | -0.763 | -0.3850 | No | ||
| 29 | JAG1 | ENSG00000101384.12 | 35655 | -0.804 | -0.3774 | No | ||
| 30 | HEY2 | ENSG00000135547.9 | 35889 | -0.830 | -0.3653 | No | ||
| 31 | WNT6 | ENSG00000115596.4 | 39561 | -1.434 | -0.4204 | Yes | ||
| 32 | ADAM17 | ENSG00000151694.14 | 39944 | -1.533 | -0.3970 | Yes | ||
| 33 | HDAC2 | ENSG00000196591.12 | 40692 | -1.746 | -0.3775 | Yes | ||
| 34 | NCOR2 | ENSG00000196498.13 | 40948 | -1.842 | -0.3447 | Yes | ||
| 35 | NKD1 | ENSG00000140807.7 | 41428 | -2.086 | -0.3119 | Yes | ||
| 36 | CTNNB1 | ENSG00000168036.18 | 41475 | -2.117 | -0.2684 | Yes | ||
| 37 | SKP2 | ENSG00000145604.16 | 41748 | -2.288 | -0.2266 | Yes | ||
| 38 | HEY1 | ENSG00000164683.17 | 41887 | -2.391 | -0.1794 | Yes | ||
| 39 | CCND2 | ENSG00000118971.8 | 41960 | -2.462 | -0.1293 | Yes | ||
| 40 | DLL1 | ENSG00000198719.9 | 42490 | -3.113 | -0.0760 | Yes | ||
| 41 | JAG2 | ENSG00000184916.9 | 42906 | -4.304 | 0.0049 | Yes |