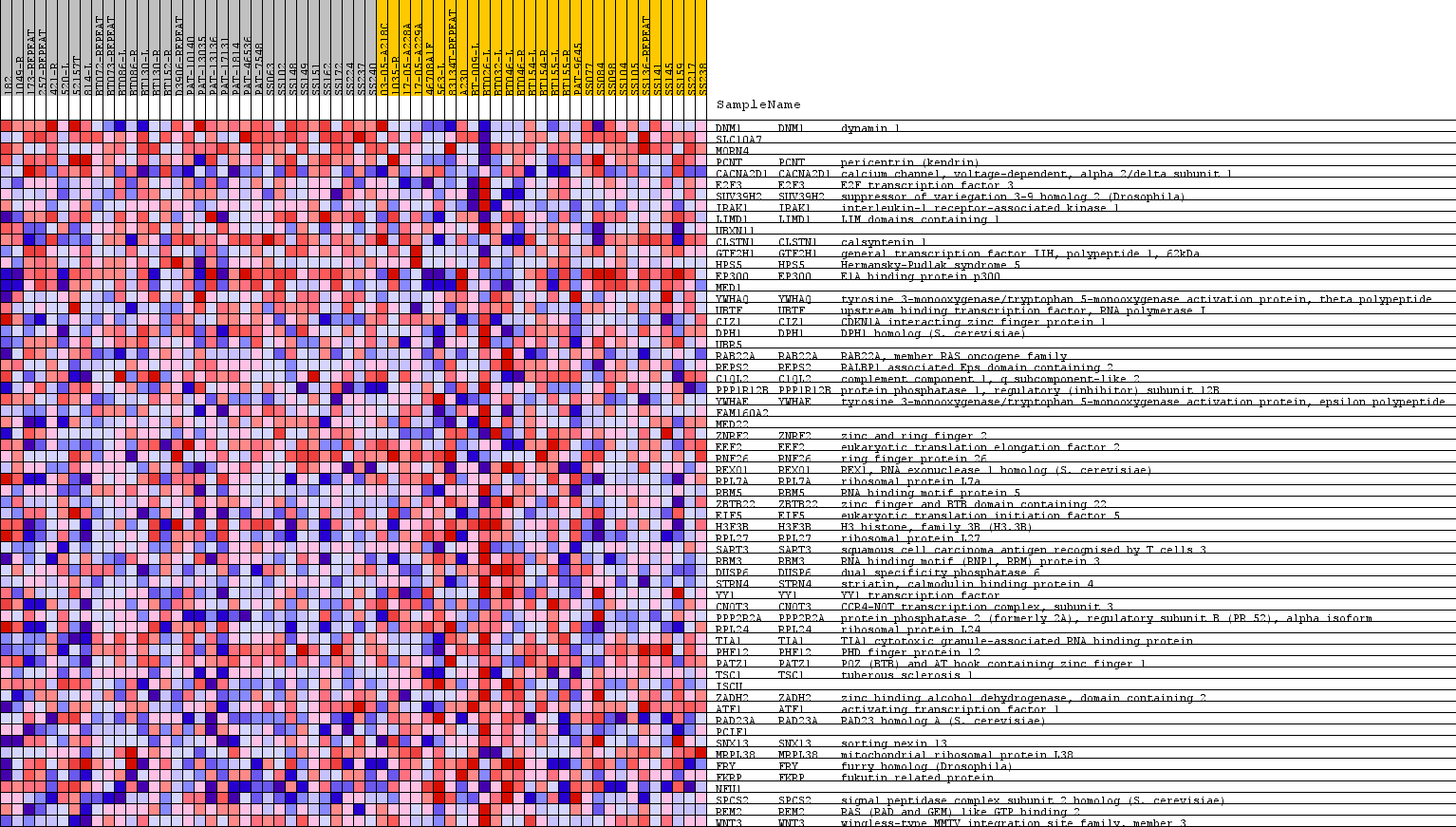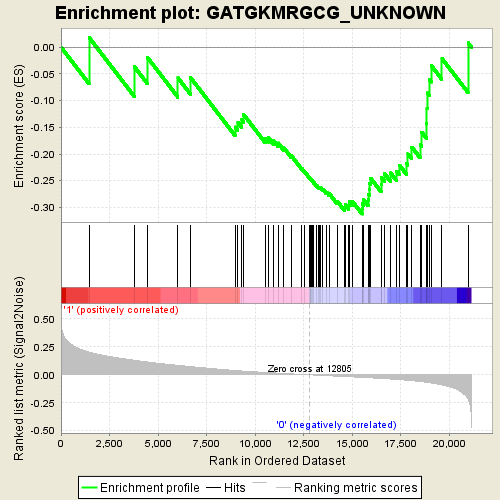
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NOV0093_collapsed_to_symbols.NOV0093.cls #obese_versus_normal.NOV0093.cls #obese_versus_normal_repos |
| Phenotype | NOV0093.cls#obese_versus_normal_repos |
| Upregulated in class | 0 |
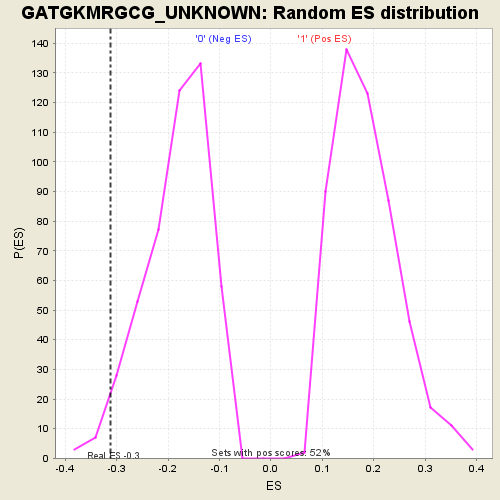
| GeneSet | GATGKMRGCG_UNKNOWN |
| Enrichment Score (ES) | -0.31234795 |
| Normalized Enrichment Score (NES) | -1.7098455 |
| Nominal p-value | 0.026915114 |
| FDR q-value | 0.99801654 |
| FWER p-Value | 0.996 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | DNM1 | DNM1 Entrez, Source | dynamin 1 | 1443 | 0.202 | 0.0184 | No |
| 2 | SLC10A7 | 3779 | 0.130 | -0.0365 | No | ||
| 3 | MORN4 | 4440 | 0.115 | -0.0186 | No | ||
| 4 | PCNT | PCNT Entrez, Source | pericentrin (kendrin) | 6023 | 0.084 | -0.0574 | No |
| 5 | CACNA2D1 | CACNA2D1 Entrez, Source | calcium channel, voltage-dependent, alpha 2/delta subunit 1 | 6677 | 0.073 | -0.0571 | No |
| 6 | E2F3 | E2F3 Entrez, Source | E2F transcription factor 3 | 8966 | 0.038 | -0.1493 | No |
| 7 | SUV39H2 | SUV39H2 Entrez, Source | suppressor of variegation 3-9 homolog 2 (Drosophila) | 9113 | 0.036 | -0.1407 | No |
| 8 | IRAK1 | IRAK1 Entrez, Source | interleukin-1 receptor-associated kinase 1 | 9314 | 0.034 | -0.1358 | No |
| 9 | LIMD1 | LIMD1 Entrez, Source | LIM domains containing 1 | 9411 | 0.033 | -0.1264 | No |
| 10 | UBXN11 | 10521 | 0.021 | -0.1701 | No | ||
| 11 | CLSTN1 | CLSTN1 Entrez, Source | calsyntenin 1 | 10682 | 0.019 | -0.1694 | No |
| 12 | GTF2H1 | GTF2H1 Entrez, Source | general transcription factor IIH, polypeptide 1, 62kDa | 10943 | 0.017 | -0.1745 | No |
| 13 | HPS5 | HPS5 Entrez, Source | Hermansky-Pudlak syndrome 5 | 11179 | 0.015 | -0.1793 | No |
| 14 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 11472 | 0.012 | -0.1881 | No |
| 15 | MED1 | 11860 | 0.008 | -0.2030 | No | ||
| 16 | YWHAQ | YWHAQ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide | 12408 | 0.003 | -0.2275 | No |
| 17 | UBTF | UBTF Entrez, Source | upstream binding transcription factor, RNA polymerase I | 12532 | 0.002 | -0.2324 | No |
| 18 | CIZ1 | CIZ1 Entrez, Source | CDKN1A interacting zinc finger protein 1 | 12807 | -0.000 | -0.2454 | No |
| 19 | DPH1 | DPH1 Entrez, Source | DPH1 homolog (S. cerevisiae) | 12828 | -0.000 | -0.2462 | No |
| 20 | UBR5 | 12843 | -0.000 | -0.2467 | No | ||
| 21 | RAB22A | RAB22A Entrez, Source | RAB22A, member RAS oncogene family | 12919 | -0.001 | -0.2499 | No |
| 22 | REPS2 | REPS2 Entrez, Source | RALBP1 associated Eps domain containing 2 | 12931 | -0.001 | -0.2499 | No |
| 23 | C1QL2 | C1QL2 Entrez, Source | complement component 1, q subcomponent-like 2 | 13030 | -0.002 | -0.2538 | No |
| 24 | PPP1R12B | PPP1R12B Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12B | 13179 | -0.003 | -0.2594 | No |
| 25 | YWHAE | YWHAE Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon polypeptide | 13271 | -0.004 | -0.2620 | No |
| 26 | FAM160A2 | 13331 | -0.004 | -0.2629 | No | ||
| 27 | MED22 | 13381 | -0.005 | -0.2631 | No | ||
| 28 | ZNRF2 | ZNRF2 Entrez, Source | zinc and ring finger 2 | 13490 | -0.006 | -0.2657 | No |
| 29 | EEF2 | EEF2 Entrez, Source | eukaryotic translation elongation factor 2 | 13673 | -0.008 | -0.2711 | No |
| 30 | RNF26 | RNF26 Entrez, Source | ring finger protein 26 | 13816 | -0.009 | -0.2740 | No |
| 31 | REXO1 | REXO1 Entrez, Source | REX1, RNA exonuclease 1 homolog (S. cerevisiae) | 14224 | -0.012 | -0.2880 | No |
| 32 | RPL7A | RPL7A Entrez, Source | ribosomal protein L7a | 14613 | -0.015 | -0.2998 | No |
| 33 | RBM5 | RBM5 Entrez, Source | RNA binding motif protein 5 | 14644 | -0.016 | -0.2945 | No |
| 34 | ZBTB22 | ZBTB22 Entrez, Source | zinc finger and BTB domain containing 22 | 14833 | -0.017 | -0.2959 | No |
| 35 | EIF5 | EIF5 Entrez, Source | eukaryotic translation initiation factor 5 | 14849 | -0.018 | -0.2891 | No |
| 36 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 15014 | -0.019 | -0.2888 | No |
| 37 | RPL27 | RPL27 Entrez, Source | ribosomal protein L27 | 15511 | -0.023 | -0.3024 | Yes |
| 38 | SART3 | SART3 Entrez, Source | squamous cell carcinoma antigen recognised by T cells 3 | 15528 | -0.023 | -0.2931 | Yes |
| 39 | RBM3 | RBM3 Entrez, Source | RNA binding motif (RNP1, RRM) protein 3 | 15591 | -0.024 | -0.2857 | Yes |
| 40 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 15822 | -0.026 | -0.2854 | Yes |
| 41 | STRN4 | STRN4 Entrez, Source | striatin, calmodulin binding protein 4 | 15845 | -0.026 | -0.2752 | Yes |
| 42 | YY1 | YY1 Entrez, Source | YY1 transcription factor | 15894 | -0.027 | -0.2660 | Yes |
| 43 | CNOT3 | CNOT3 Entrez, Source | CCR4-NOT transcription complex, subunit 3 | 15904 | -0.027 | -0.2549 | Yes |
| 44 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 15955 | -0.027 | -0.2456 | Yes |
| 45 | RPL24 | RPL24 Entrez, Source | ribosomal protein L24 | 16490 | -0.032 | -0.2570 | Yes |
| 46 | TIA1 | TIA1 Entrez, Source | TIA1 cytotoxic granule-associated RNA binding protein | 16502 | -0.032 | -0.2436 | Yes |
| 47 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 16658 | -0.034 | -0.2364 | Yes |
| 48 | PATZ1 | PATZ1 Entrez, Source | POZ (BTB) and AT hook containing zinc finger 1 | 16975 | -0.037 | -0.2353 | Yes |
| 49 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 17294 | -0.041 | -0.2327 | Yes |
| 50 | ISCU | 17430 | -0.043 | -0.2208 | Yes | ||
| 51 | ZADH2 | ZADH2 Entrez, Source | zinc binding alcohol dehydrogenase, domain containing 2 | 17808 | -0.048 | -0.2179 | Yes |
| 52 | ATF1 | ATF1 Entrez, Source | activating transcription factor 1 | 17871 | -0.049 | -0.1998 | Yes |
| 53 | RAD23A | RAD23A Entrez, Source | RAD23 homolog A (S. cerevisiae) | 18069 | -0.052 | -0.1868 | Yes |
| 54 | PCIF1 | 18498 | -0.060 | -0.1815 | Yes | ||
| 55 | SNX13 | SNX13 Entrez, Source | sorting nexin 13 | 18584 | -0.061 | -0.1592 | Yes |
| 56 | MRPL38 | MRPL38 Entrez, Source | mitochondrial ribosomal protein L38 | 18825 | -0.067 | -0.1419 | Yes |
| 57 | FRY | FRY Entrez, Source | furry homolog (Drosophila) | 18852 | -0.067 | -0.1142 | Yes |
| 58 | FKRP | FKRP Entrez, Source | fukutin related protein | 18859 | -0.068 | -0.0855 | Yes |
| 59 | NFU1 | 18974 | -0.070 | -0.0607 | Yes | ||
| 60 | SPCS2 | SPCS2 Entrez, Source | signal peptidase complex subunit 2 homolog (S. cerevisiae) | 19067 | -0.073 | -0.0339 | Yes |
| 61 | REM2 | REM2 Entrez, Source | RAS (RAD and GEM) like GTP binding 2 | 19625 | -0.091 | -0.0212 | Yes |
| 62 | WNT3 | WNT3 Entrez, Source | wingless-type MMTV integration site family, member 3 | 20968 | -0.217 | 0.0083 | Yes |